External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G20540 62 / 7e-13
MEF21
mitochondrial editing factor 21 (.1)
AT1G08070 56 / 6e-11
EMB3102, OTP82
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G59720 54 / 3e-10
CRR28
CHLORORESPIRATORY REDUCTION28, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G29760 54 / 3e-10
OTP81
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G49142 48 / 4e-08
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G22410 47 / 1e-07
SLO1
SLOW GROWTH 1 (.1)
AT1G11290 46 / 1e-07
CRR22
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G28660 46 / 2e-07
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G02750 46 / 2e-07
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G02980 45 / 4e-07
OTP85
ORGANELLE TRANSCRIPT PROCESSING 85, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022124
131 / 1e-39
AT2G20540 345 / 1e-116
mitochondrial editing factor 21 (.1)
Lus10030125
133 / 8e-39
AT2G20540 618 / 0.0
mitochondrial editing factor 21 (.1)
Lus10033026
50 / 8e-09
AT1G08070 921 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10014211
49 / 3e-08
AT3G11460 791 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10002288
48 / 4e-08
AT1G31430 689 / 0.0
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
Lus10000110
48 / 4e-08
AT4G37380 413 / 9e-142
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10017446
48 / 5e-08
AT1G11290 1152 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10018223
48 / 5e-08
AT2G29760 931 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10040680
48 / 5e-08
AT2G29760 926 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G035900
81 / 7e-20
AT2G20540 709 / 0.0
mitochondrial editing factor 21 (.1)
Potri.008G106500
57 / 2e-11
AT2G29760 422 / 4e-139
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.007G050200
57 / 2e-11
AT4G37380 817 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.003G031600
57 / 3e-11
AT3G15930 784 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.002G027800
51 / 4e-09
AT1G25360 1047 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.003G191000
51 / 4e-09
AT1G11290 1178 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G090600
50 / 6e-09
AT2G40720 912 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G239600
50 / 7e-09
AT5G59200 694 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 80, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.013G103600
50 / 8e-09
AT1G08070 477 / 7e-160
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.009G044700
50 / 1e-08
AT2G29760 1013 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10007685 pacid=23165685 polypeptide=Lus10007685 locus=Lus10007685.g ID=Lus10007685.BGIv1.0 annot-version=v1.0
ATGAGGACACTAATAAGAAGCAAGGGCATGAAGAAAACTCCAGGGTGTAGCTCGATTGAAATCAACAATGTAGCTACTGAGTTCTCTTCTGGCGATGATT
CGAAGCCCTTCTCAAGACAGATCTTCAAGTTGCTAGAGATGCTGACTTCTTGTGAGGAGATAAGTGATCAAATACTTGACATCATTCCAGAGGATGCAGG
CAGCTGA
AA sequence
>Lus10007685 pacid=23165685 polypeptide=Lus10007685 locus=Lus10007685.g ID=Lus10007685.BGIv1.0 annot-version=v1.0
MRTLIRSKGMKKTPGCSSIEINNVATEFSSGDDSKPFSRQIFKLLEMLTSCEEISDQILDIIPEDAGS
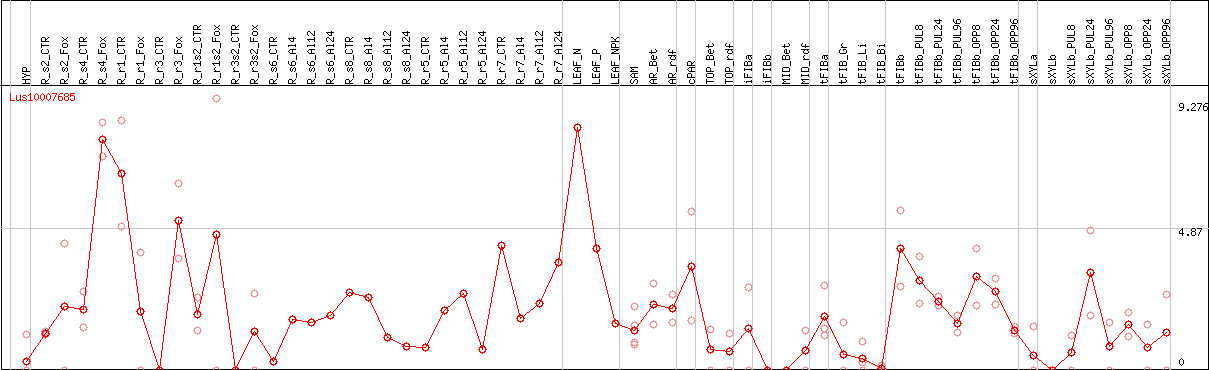
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007685 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.