External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G09960 363 / 3e-123
ATSUC4, SUC4, ATSUT4, SUT4
sucrose transporter 4 (.1)
AT1G22710 232 / 3e-72
SUT1, ATSUC2, SUC2
SUCROSE TRANSPORTER 1, ARABIDOPSIS THALIANA SUCROSE-PROTON SYMPORTER 2, sucrose-proton symporter 2 (.1)
AT2G14670 225 / 1e-69
ATSUC8
sucrose-proton symporter 8 (.1)
AT5G43610 221 / 3e-68
ATSUC6
sucrose-proton symporter 6 (.1)
AT1G66570 221 / 3e-68
ATSUC7
sucrose-proton symporter 7 (.1.2.3)
AT5G06170 217 / 8e-67
ATSUC9
sucrose-proton symporter 9 (.1)
AT1G71880 214 / 2e-65
ATSUC1, SUC1
ARABIDOPSIS THALIANA SUCROSE-PROTON SYMPORTER 1, sucrose-proton symporter 1 (.1)
AT1G71890 207 / 1e-62
SUC5, ATSUC5
SUCROSE-PROTON SYMPORTER 5, Major facilitator superfamily protein (.1)
AT2G02860 200 / 2e-60
ATSUT2, ATSUC3, SUT2
ARABIDOPSIS THALIANA SUCROSE TRANSPORTER 3, sucrose transporter 2 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032815
514 / 0
AT1G09960 694 / 0.0
sucrose transporter 4 (.1)
Lus10018915
225 / 2e-69
AT1G71890 616 / 0.0
SUCROSE-PROTON SYMPORTER 5, Major facilitator superfamily protein (.1)
Lus10028616
223 / 1e-68
AT1G71890 617 / 0.0
SUCROSE-PROTON SYMPORTER 5, Major facilitator superfamily protein (.1)
Lus10017910
216 / 4e-66
AT1G71890 693 / 0.0
SUCROSE-PROTON SYMPORTER 5, Major facilitator superfamily protein (.1)
Lus10009428
208 / 2e-64
AT1G22710 471 / 1e-164
SUCROSE TRANSPORTER 1, ARABIDOPSIS THALIANA SUCROSE-PROTON SYMPORTER 2, sucrose-proton symporter 2 (.1)
Lus10007702
206 / 2e-62
AT1G22710 588 / 0.0
SUCROSE TRANSPORTER 1, ARABIDOPSIS THALIANA SUCROSE-PROTON SYMPORTER 2, sucrose-proton symporter 2 (.1)
Lus10030492
207 / 2e-61
AT2G02860 863 / 0.0
ARABIDOPSIS THALIANA SUCROSE TRANSPORTER 3, sucrose transporter 2 (.1.2)
Lus10029766
198 / 6e-59
AT1G22710 655 / 0.0
SUCROSE TRANSPORTER 1, ARABIDOPSIS THALIANA SUCROSE-PROTON SYMPORTER 2, sucrose-proton symporter 2 (.1)
Lus10028614
197 / 1e-58
AT1G71890 603 / 0.0
SUCROSE-PROTON SYMPORTER 5, Major facilitator superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G106900
402 / 9e-139
AT1G09960 674 / 0.0
sucrose transporter 4 (.1)
Potri.019G085800
219 / 6e-67
AT1G22710 645 / 0.0
SUCROSE TRANSPORTER 1, ARABIDOPSIS THALIANA SUCROSE-PROTON SYMPORTER 2, sucrose-proton symporter 2 (.1)
Potri.013G115200
210 / 1e-63
AT1G22710 643 / 0.0
SUCROSE TRANSPORTER 1, ARABIDOPSIS THALIANA SUCROSE-PROTON SYMPORTER 2, sucrose-proton symporter 2 (.1)
Potri.010G093600
207 / 5e-62
AT2G02860 816 / 0.0
ARABIDOPSIS THALIANA SUCROSE TRANSPORTER 3, sucrose transporter 2 (.1.2)
Potri.008G148100
205 / 4e-61
AT2G02860 849 / 0.0
ARABIDOPSIS THALIANA SUCROSE TRANSPORTER 3, sucrose transporter 2 (.1.2)
PFAM info
Representative CDS sequence
>Lus10007691 pacid=23164947 polypeptide=Lus10007691 locus=Lus10007691.g ID=Lus10007691.BGIv1.0 annot-version=v1.0
ATGGCTGTTGGCAATATCCTTGGCTTTGCTACTGGAGCTTTCAGTGGTTGGTACAAAATTTTTCGGTTCACTCTTACTACTGCATGTGACGTGGACTGTG
CCAACCTTAAAGCTGCTTTCTATCTTGATGTTGTCTGTATTGTAATCACCGCATATATCAGTATTTCATCAGCTAAAGAACCATCTATAAGCTTGCTTCA
TGCATCTACTCCTGTTGTTGAAGAAGAAGCAGTTGAGTCCAGCCCTGTAGAAGAGGCTTTCCTCTGGGAGCTGTTTGGTACTTTCGGATATTTCCCAGCA
TCTGTGTGGACTATATTGCTTGTTACTGCTCTCAATTGGATCGGTTGGTTTCCGTTTCTACTCTTTGATACCGACTGGATGGGGCGAGAAATCTATGGCG
GGGACCCAGATGGAGGACCAAGTTATAGTTCTGGAGTGAGAATGGGAGCGTTTGCACTTATGTTAAATTCCGTGCTTTTAGGAATAACGTCAGTACTGAT
GGAGAAGATCTGCAGGAAGTGGGGAGCTGGCTTCATTTGGGGACTCTCTAACATCTTCATGGCTATCTGCTTCCTTTCAATGCTTATAATTGCCTATGTA
GCATACCATATGGGCTATCTGGGTCATGGTGAACCACCAGTTGGTATTCTTGTTGCTGCGCTGACTGTTTTTGCATTCCTTGGGATTCCATTGGCAATCA
CCTATAGTGTTCCATATGCCTTAGTGTCCTCACGAATTGAGCCTTTGGGGCTTGGCCAAGGGTTATCGATGGGTGTATTGAATTTGGCAATTGTCGTCCC
GCAGGTGCTGGTGTCGGTGGGATCTGGTCCGTGGGATCAGTTGTTCGGAGGTGGAAACTCGCCAGCCATAGCTGTGGGAGGTTTGGCAGCACTAATAGGT
GGAGTGATCGCTATATTGGGGATTCCTCGTTTCACTGCTCAAAAGCCAAGGCTTCTTCCATGA
AA sequence
>Lus10007691 pacid=23164947 polypeptide=Lus10007691 locus=Lus10007691.g ID=Lus10007691.BGIv1.0 annot-version=v1.0
MAVGNILGFATGAFSGWYKIFRFTLTTACDVDCANLKAAFYLDVVCIVITAYISISSAKEPSISLLHASTPVVEEEAVESSPVEEAFLWELFGTFGYFPA
SVWTILLVTALNWIGWFPFLLFDTDWMGREIYGGDPDGGPSYSSGVRMGAFALMLNSVLLGITSVLMEKICRKWGAGFIWGLSNIFMAICFLSMLIIAYV
AYHMGYLGHGEPPVGILVAALTVFAFLGIPLAITYSVPYALVSSRIEPLGLGQGLSMGVLNLAIVVPQVLVSVGSGPWDQLFGGGNSPAIAVGGLAALIG
GVIAILGIPRFTAQKPRLLP
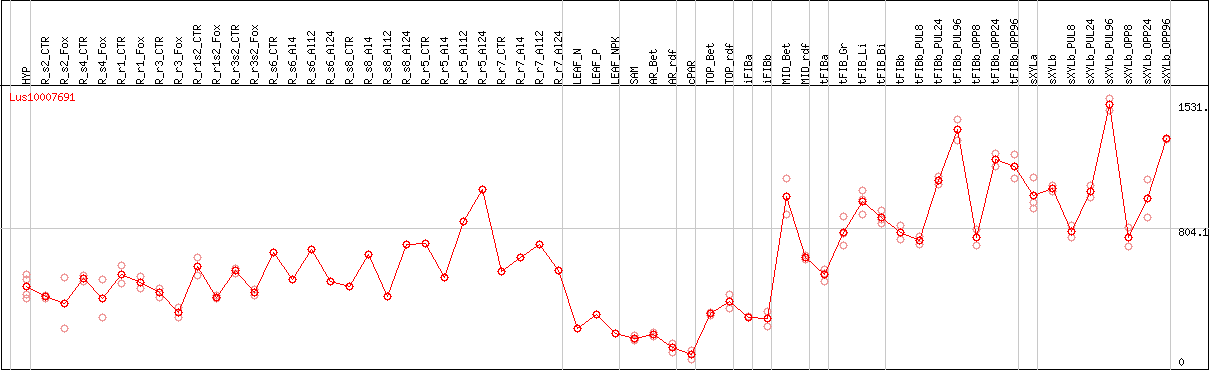
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007691 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.