Lus10007734 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10007734 pacid=23149268 polypeptide=Lus10007734 locus=Lus10007734.g ID=Lus10007734.BGIv1.0 annot-version=v1.0
ATGGGGAAACGTAGTAATAGAAGGAAGAATGCCGCCATGTTGGACAGTGATGATGAGGATAATAGTAGCGTGAGTTCGTCGTCTACTGTTCGTTCGGATC
GTATGTCTGTCAATGGGACTGAAGAGGTTGAGGTTGAGAAGGATAACTTGCTAGATCAGGCTTTGGATGCTTTGTACGAGAAGAGGAGTACAACAAGAGA
AAAAGCGTTGGCTTCAATTATCGAGGCTTTCAATACCAATTTGCAGCATGAGTTTGTGGAAAAGAAGTTTGCAACTTTGCTACATCAATGTATTAGCTGC
ATTAAAAAAGGGTCCACCACAGAGATAGCTCTTGCTTCTCATGCCATTGGATTATTGTCATTGACGGTTGGTTCTGGGGATAATGCCCGTGAAATATTGG
AAGAGTCGCTCGCTCCAATTTCCCAGGCACTCAAGTCTGGAACAGACTCGTTGAGGAGGGTAGCATTGTTGGAGTGTCTTGCTATCATAACTTTTGTTGG
TGGAGACGATCCTGAGCAAACAGAACAATCTATGCAGATCATGTGGCAGTTGGTTCGGCCTAAGCTTGGCCCAAATGTTGTTGCTGTCAAACCTTCTAGT
GCTGTAATTACAGCAGTGGCGTCTGCTTGGGCATTTCTTTTGACATCCATGGACCAGAGGACACTAAGCCCCAAGGATTGGCAAGAGTCCATTTCATACT
TCTCTACTCTCCTCGACAAGGAGGACCGAACTATTCGCATTGCAGCTGGTGAAGCACTGGCTTTTATTTTTGAGACGGGATGCCTAGAAAAGTTTGCTGC
TGAGGGTGTGGAAAGCTCAGTTCAAGATGGGGATAAATCGTATACACACCTACGAGGTCTAAAAGCGAAAATCTTAAATCAAGTGAGAGACCTCGCAGCA
GAGGCCGGAGGAAAAGGGTCAGCGAAGAAAGACCTGAACAGCCAGCGGAACTTGTTTAAGGACATCCTGGAGTTCATTGAGGACGGTTACAGTCCTGAAG
TCACAGTGAAGATAGGTGGAGATCCATTCCAGACGTCCACCTGGTATCAACTTACACAGCTTAACTTCTTGAAGCATTTTCTTGGTGGTGGCTTTGTCAA
ACACATGCAGGATAATGAATTCCTCCAAGATGTACTTGGATTCGCACCAAAGAAAAAGCTTTTTCAGGGTGCTGATTCTCAGCTATCAAGTGGTGATAAG
AGAATGTTCCGGTCACCAAATTCGGTTATGAACAAAGCCAGAACTCAATACCTAAACAAGCAACGAATGCTGTCTGAGGTGAAGATATTGTTGATGTGGA
TGGTGTCTTTGAGACTGATAAATGTTCTTCCACATGGTCAACAAGCTGCGATTGGCGGGGCACAATTCTCTGCCAAACCTGAGAACAGCCCTATGGATCT
TGTTTCACCCGCAATGAATTTGGAGAATGGTGGTGCTGCTGTGCATATTGGACATCCTTTTGGTTCTTTGAATCCTTCTCAAGGAGGAAACCATAGTGCT
GCTGCGGAACAAATGTGGATGTACAAGAACAATGAGACTGGTAGCTATCCTTGTGCGCCTGTGGGAACCCAGGGACAGGATATGCCTCCACCACCTCATT
TTCAGCAGGCCCCATATGCTCAACCTGTATACCAACCTCCCAATAGTTCCTGGGATCCTCGGGGTTCAAATCATCATGTTCCTTTCAATCAGGAGCCGTC
AGCAGTGATGAATACTAGCTTCCAGGGAAGCACTGTTGCACCTCCTTTTGTGGCCGCATCATCGACTCCGTTAGCACATGGACCAGATTGGTTACGTAGG
GCGAGAACTCCTTTCTGGAAATTGGTTCTGAAACAGTTTGATGATTTACTTGTTAAGATACTGATAGCTGCTGCAATTGTTTCTTTCGTTTTGGCTTTGA
TCAACGGGGAAACTGGCCTGACTGCCTTTGTGGAGCCTTTTGTTATTCTCTTGATTTTGGCTGCCAATGCTGCAGTTGGCGTTATTACAGAAACGAATGC
TGAAAAAGCTCTCGAGGAATTACGTGCCTATCAAGCTGAAGCTGCAACTGTGCTGCGGAATGGCTGTTTTTCAATACTTCCGGCAGCAGATCTTGTTCCC
GGTGACATAGTGGAAGTGACAGTGGGATGCAAGGTTCCAGCTGACATGAGAATGATTGAGATGTTAAGTGATCAGTTACGCGTGGACCAGGCAATCCTTA
CTGGTGAAAGCAGTTCGGCTGACAAGGAGCTCGAGTCAACTGTGACAACCAATGCTGTATATCAGGACAAAACAAACATCCTTTTCTCGGGTACTGTTGT
GGTCGCGGGAAGAGCAAGAGCTGTGGTTGTTGGAGTTGGTGCTAATACTGCTATGGGGAGCATACGCGATTCAATGTTACATACAGACGATGAAGCTACA
CCACTCAAGAAGAAGTTGGATGAGTTTGGCAGTTTTCTGGCTAAGGTTATTGCAGGGATTTGTGTCCTAGTATGGATTGTAAACATAGGCCACTTCCGTG
ATGCTGCTCATGGTGGTTTCTTCAGGGGGGCAATCCATTACTTCAAGATTGCAGTTGCTCTTGCAGTTGCTGCAATTCCCGAAGGACTTCCTGCTGTTGT
TACCACGTGTTTGGCATTGGGAACAAAGAGGATGGCTCGGTTGAACGCGATTGTGCGGTCTTTGCCATCAGTTGAGACTTTGGGCTGTACTACTGTCATT
TGCAGTGACAAGACAGGGACTTTGACCACAAATATGATGTCTGTCTCTAAGATATGTGTTTTGGACTCTTTTCATCATGGGCCTTTGGTCACCGAGTACA
GTATAAGTGGGACCACTTATGCTCCAGATGGAATTATATACGACAACAATGGATTGCAGCTTGAACTGCCAGCTAAGTTGCCCAGTCTTCTTCACCTGGC
TATGTGCTCAGCCCTTTGTAATGAATCCGTCCTGCAATACAACCCAGATAAAACATGTTATGAAAAAATTGGGGAGTCAACTGAAGTCGCACTTCGGGTT
CTAGCAGAGAAGGTTGGTCTTCCTGGTTTCGATTCTATTTCGTCTGCTCTCCACATGCTAAGCAAGCACGAGCGCGCCTCCTACTGTAATCATTATTGGG
AAAACCAATTCAGAAAGATTTCTGCATTGGAATTCTCACGTGATCGGAAGATGATGAGCGTCTTATGCAGTCAAAAGCACACACAAGTTATGTTTTCGAA
AGGTGCCCCAGAGAGTATTTTGTCAAGATGCTCTCATATCCTCTGTAATGATAATGGGTCCACCATTCCGCTATCTGCTAACATTCGAGATGAAGTAGAA
TCAAGGTTTCACAGTTTTGCAGGGAAAGAAACATTGAGATGCCTGGCTCTGGCCATGAAACAGATGCCTGTAGAGCAGCAGAGCCTCTCCCTGGATGATG
AGAAGGGCCTGACTTTCATTGGGTTGGTTGGAATGCTTGATCCACCAAGAGAAGAAGTAAGAAATGCTATGCTTTCATGCATGACTGCTGGTATACGCGT
CATTGTTGTTACTGGTGACAATAAGTCAACAGCAGAGTCACTTTGCCGGAAGATAGGCGCTTTCGATCACTTGGAAGATTTTTCCGGGCATTCTTATACT
GCCTCGGAGTTTGAAGAACTTCCACCTTTACAGCAAACACTTGCTCTGCAACGCATGGCACTCTTTACTAGGGTGGAACCAACTCACAAAAAAATGCTGG
TGGAGGCCTTGCAACATCAAAATGAAGTGGTGGCAATGACTGGTGATGGTGTGAATGATGCACCTGCCCTAAAGAAGGCAAACATAGGAATCGCCATGGG
GTCTGGAACAGCAGTTGCAAAGAGTGCCTCTGATATGGTGTTGGCCGATGATAATTTTGCTTCAATTGTTGCGGCTGTTGCAGAAGGGAGGGCAATTTAT
AATAACACAAAGCAGTTCATCAGATACATGATCTCTTCAAATATTGGTGAAGTTGTTTGTATCTTCGTGGCTGCTGTACTTGGAATACCAGACACCTTGG
CTCCTGTCCAACTGCTATGGGTAAATCTGGTGACTGATGGCTTGCCAGCAACCGCAATTGGTTTTAACAAGCAAGATAGTGATGTTATGAAGGTCAAACC
TCGGAAGGTTAATGAAGCAGTAGTTAGTGGATGGGTGTTTTTCCGTTATTTGGTTATTGGAGCCTACGTCGGCCTGGCCACAGTTGCAGGATTTGTGTGG
TGGTTTTTGTACTCAGATAATGGTCCGAAATTACCATATCGAGAACTGATGAGTTTCGACACGTGTTCAACAAGGGATACTACTTACCCATGTAGCATAT
TTGAGGACCGGCATCCATCAACTGTATCAATGACAGTGCTTGTTGTTGTTGAGATGTTTAATGCTCTAAACAATCTTAGCGAAAATCAGTCTCTATTTGT
TATCCCTCCCTGGAGTAATATGTGGCTGGTTGGATCAATTATCCTGACTATGATCCTTCACGTACTAATCTTATATATCCCTCCTCTCTCTGTTCTTTTC
TCTGTGACACCTCTGTCTTGGTCTGATTGGAAAATCATATTGTATCTTTCATTCCCTGTTATAATTATTGACGAAATCTTGAAGTTCTTCTCGAGAAAAC
CAAACGGTTTGAGGTTGGGTCTTCGGTTCCGAAGGAATGATTTTATACCGAAGAGCGATCGCCATGATAAGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10007734 pacid=23149268 polypeptide=Lus10007734 locus=Lus10007734.g ID=Lus10007734.BGIv1.0 annot-version=v1.0
MGKRSNRRKNAAMLDSDDEDNSSVSSSSTVRSDRMSVNGTEEVEVEKDNLLDQALDALYEKRSTTREKALASIIEAFNTNLQHEFVEKKFATLLHQCISC
IKKGSTTEIALASHAIGLLSLTVGSGDNAREILEESLAPISQALKSGTDSLRRVALLECLAIITFVGGDDPEQTEQSMQIMWQLVRPKLGPNVVAVKPSS
AVITAVASAWAFLLTSMDQRTLSPKDWQESISYFSTLLDKEDRTIRIAAGEALAFIFETGCLEKFAAEGVESSVQDGDKSYTHLRGLKAKILNQVRDLAA
EAGGKGSAKKDLNSQRNLFKDILEFIEDGYSPEVTVKIGGDPFQTSTWYQLTQLNFLKHFLGGGFVKHMQDNEFLQDVLGFAPKKKLFQGADSQLSSGDK
RMFRSPNSVMNKARTQYLNKQRMLSEVKILLMWMVSLRLINVLPHGQQAAIGGAQFSAKPENSPMDLVSPAMNLENGGAAVHIGHPFGSLNPSQGGNHSA
AAEQMWMYKNNETGSYPCAPVGTQGQDMPPPPHFQQAPYAQPVYQPPNSSWDPRGSNHHVPFNQEPSAVMNTSFQGSTVAPPFVAASSTPLAHGPDWLRR
ARTPFWKLVLKQFDDLLVKILIAAAIVSFVLALINGETGLTAFVEPFVILLILAANAAVGVITETNAEKALEELRAYQAEAATVLRNGCFSILPAADLVP
GDIVEVTVGCKVPADMRMIEMLSDQLRVDQAILTGESSSADKELESTVTTNAVYQDKTNILFSGTVVVAGRARAVVVGVGANTAMGSIRDSMLHTDDEAT
PLKKKLDEFGSFLAKVIAGICVLVWIVNIGHFRDAAHGGFFRGAIHYFKIAVALAVAAIPEGLPAVVTTCLALGTKRMARLNAIVRSLPSVETLGCTTVI
CSDKTGTLTTNMMSVSKICVLDSFHHGPLVTEYSISGTTYAPDGIIYDNNGLQLELPAKLPSLLHLAMCSALCNESVLQYNPDKTCYEKIGESTEVALRV
LAEKVGLPGFDSISSALHMLSKHERASYCNHYWENQFRKISALEFSRDRKMMSVLCSQKHTQVMFSKGAPESILSRCSHILCNDNGSTIPLSANIRDEVE
SRFHSFAGKETLRCLALAMKQMPVEQQSLSLDDEKGLTFIGLVGMLDPPREEVRNAMLSCMTAGIRVIVVTGDNKSTAESLCRKIGAFDHLEDFSGHSYT
ASEFEELPPLQQTLALQRMALFTRVEPTHKKMLVEALQHQNEVVAMTGDGVNDAPALKKANIGIAMGSGTAVAKSASDMVLADDNFASIVAAVAEGRAIY
NNTKQFIRYMISSNIGEVVCIFVAAVLGIPDTLAPVQLLWVNLVTDGLPATAIGFNKQDSDVMKVKPRKVNEAVVSGWVFFRYLVIGAYVGLATVAGFVW
WFLYSDNGPKLPYRELMSFDTCSTRDTTYPCSIFEDRHPSTVSMTVLVVVEMFNALNNLSENQSLFVIPPWSNMWLVGSIILTMILHVLILYIPPLSVLF
SVTPLSWSDWKIILYLSFPVIIIDEILKFFSRKPNGLRLGLRFRRNDFIPKSDRHDK
|
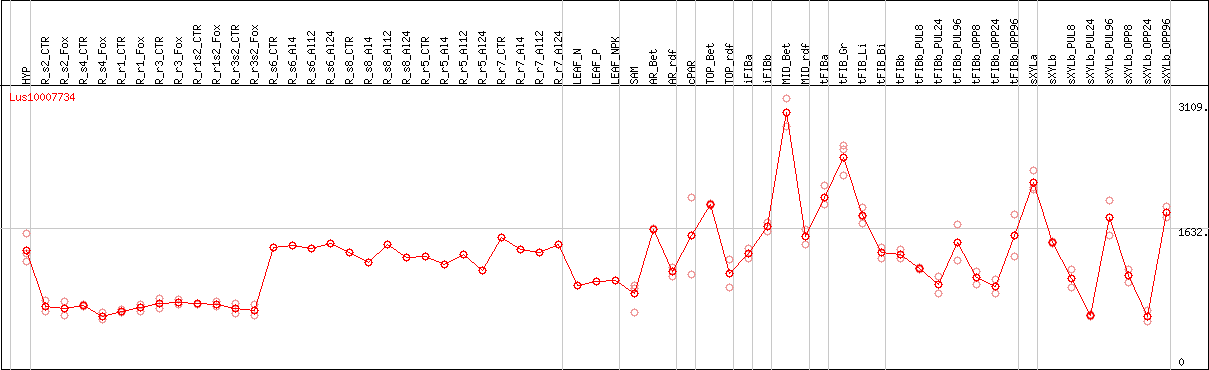
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007734 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
