External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G02860 199 / 3e-64
BAH1, NLA
nitrogen limitation adaptation, BENZOIC ACID HYPERSENSITIVE 1, SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
AT2G38920 129 / 4e-37
SPX (SYG1/Pho81/XPR1) domain-containing protein / zinc finger (C3HC4-type RING finger) protein-related
AT3G07200 42 / 5e-05
RING/U-box superfamily protein (.1.2)
AT3G27330 40 / 0.0003
zinc finger (C3HC4-type RING finger) family protein (.1)
AT4G28270 39 / 0.0003
ATRMA2
RING membrane-anchor 2 (.1)
AT1G18660 40 / 0.0004
zinc finger (C3HC4-type RING finger) family protein (.1), zinc finger (C3HC4-type RING finger) family protein (.2), zinc finger (C3HC4-type RING finger) family protein (.3), zinc finger (C3HC4-type RING finger) family protein (.4)
AT4G03510 39 / 0.0004
ATRMA1, RMA1
RING membrane-anchor 1 (.1.2)
AT4G27470 38 / 0.001
ATRMA3
RING membrane-anchor 3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030375
273 / 4e-93
AT1G02860 386 / 1e-134
nitrogen limitation adaptation, BENZOIC ACID HYPERSENSITIVE 1, SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
Lus10012290
117 / 4e-32
AT2G38920 392 / 5e-137
SPX (SYG1/Pho81/XPR1) domain-containing protein / zinc finger (C3HC4-type RING finger) protein-related
Lus10015986
116 / 8e-32
AT2G38920 389 / 1e-135
SPX (SYG1/Pho81/XPR1) domain-containing protein / zinc finger (C3HC4-type RING finger) protein-related
Lus10015722
42 / 7e-05
AT1G18660 667 / 0.0
zinc finger (C3HC4-type RING finger) family protein (.1), zinc finger (C3HC4-type RING finger) family protein (.2), zinc finger (C3HC4-type RING finger) family protein (.3), zinc finger (C3HC4-type RING finger) family protein (.4)
Lus10019075
42 / 8e-05
AT1G18660 583 / 0.0
zinc finger (C3HC4-type RING finger) family protein (.1), zinc finger (C3HC4-type RING finger) family protein (.2), zinc finger (C3HC4-type RING finger) family protein (.3), zinc finger (C3HC4-type RING finger) family protein (.4)
Lus10022324
39 / 0.0006
AT3G27330 251 / 4e-75
zinc finger (C3HC4-type RING finger) family protein (.1)
Lus10032119
39 / 0.0008
AT3G27330 251 / 1e-75
zinc finger (C3HC4-type RING finger) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G205400
221 / 1e-72
AT1G02860 437 / 5e-155
nitrogen limitation adaptation, BENZOIC ACID HYPERSENSITIVE 1, SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
Potri.014G130300
216 / 4e-71
AT1G02860 448 / 3e-159
nitrogen limitation adaptation, BENZOIC ACID HYPERSENSITIVE 1, SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
Potri.006G198600
129 / 5e-37
AT2G38920 390 / 3e-136
SPX (SYG1/Pho81/XPR1) domain-containing protein / zinc finger (C3HC4-type RING finger) protein-related
Potri.016G064600
128 / 1e-36
AT2G38920 414 / 2e-145
SPX (SYG1/Pho81/XPR1) domain-containing protein / zinc finger (C3HC4-type RING finger) protein-related
Potri.013G132300
42 / 4e-05
AT4G03510 191 / 2e-60
RING membrane-anchor 1 (.1.2)
Potri.001G336832
41 / 0.0001
AT3G27330 235 / 2e-69
zinc finger (C3HC4-type RING finger) family protein (.1)
Potri.012G068100
39 / 0.0007
AT1G18660 679 / 0.0
zinc finger (C3HC4-type RING finger) family protein (.1), zinc finger (C3HC4-type RING finger) family protein (.2), zinc finger (C3HC4-type RING finger) family protein (.3), zinc finger (C3HC4-type RING finger) family protein (.4)
PFAM info
Representative CDS sequence
>Lus10007867 pacid=23143182 polypeptide=Lus10007867 locus=Lus10007867.g ID=Lus10007867.BGIv1.0 annot-version=v1.0
ATGGCTTTCCATATCAACTTGAGAGCAACATCATCAAACAAACCACCTCCTGCTGCTGGTGCTGCTTCTTCGAATCTATTCCGGGGCTCCTCCCTTTCTT
TTGATGATGAACATAGTAAAAAGCCATCATTGTCTTGTGACCTCTTTGACTCAAAAAAGGTTGACATTGATCTCACTTGTTCCATATGCCTGACGGTATT
CGATCCAGTTTCACTGAGATGCGGCCATATCTTCTGCTATATCTGTGCTTGTTCAGCTGCTTCGGTAACAGTCGTTGATGGACTCCAGTCTGCTGATCCT
CAACGGAAATGCCCTATTTGCAGGGAGACTGGAGTTTTTGAAGGTGTTGTGCACTTGGAAGAGCTTAACATTCTATTAAGCCAGAGTTGCAAAGAGTATT
GGGAGGAGAGGCTTCAATCAGAAAGGCGAGAGAGGGTTAAGCAAGCAAAACAGCACTGGGAAAACTAG
AA sequence
>Lus10007867 pacid=23143182 polypeptide=Lus10007867 locus=Lus10007867.g ID=Lus10007867.BGIv1.0 annot-version=v1.0
MAFHINLRATSSNKPPPAAGAASSNLFRGSSLSFDDEHSKKPSLSCDLFDSKKVDIDLTCSICLTVFDPVSLRCGHIFCYICACSAASVTVVDGLQSADP
QRKCPICRETGVFEGVVHLEELNILLSQSCKEYWEERLQSERRERVKQAKQHWEN
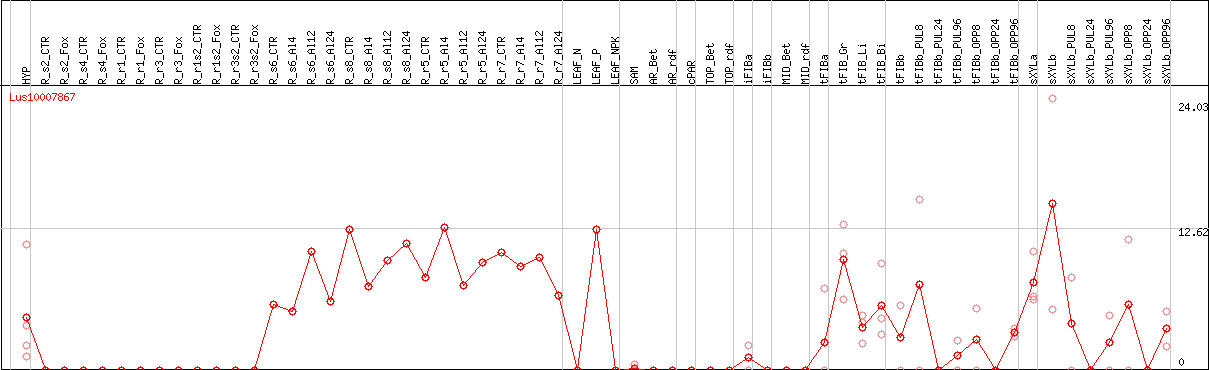
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007867 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.