Lus10007938 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10007938 pacid=23162297 polypeptide=Lus10007938 locus=Lus10007938.g ID=Lus10007938.BGIv1.0 annot-version=v1.0
ATGAAAGACAGGCCGCCGTCCCCTTACGGCGCCGTTTACGTTCCTCCGCACCAGCGTCTCCGCTCCGTGATCGTCTCCCCCAATTTCAACTCCTCCTCTG
CATCTTCTCATCTTCTGGAGTCCAGAACTACTGACAGTCGATGCGCTGCTCTCAGCCGCAGTAAGGCTAGCAGTCCTTCCCCTACAGAACATCAGCAGAG
AAATGGCGGTGCTGCTGAGGTCTGTAACAAGAATCCGATTCGGAAATTTGCCTCTGCCTGCGTCAATGCTGTCTCCGTGGACGGGTCCGATCACGATATG
GAGTCCTTGCCAGCCCACCCATCAGGAACTTATATGTCTGATAGCACTGAAGAGTGGAGATGGAAGTTCACGATGCTTTTGCGTGACAAAGAGAAGAAAG
AAGTGGTATCAAGGGAGAAAAAGGACAGACGTGATTTCGAGCAACTAGCCGCTTTAGCAAGCAAAATGGGTCTATATAGCCGTGTGTATGTGAAAGTTGT
TGTCTTTAGCAAAGTCCCACTGCCGAACTACAGATTTGATTTGGATGATAAGCGTCCTCAACGGGAGGTTAGCTTGCCTCTTGGCGTATTGAAAAGAGTT
GATACGTATCTTGGAGACTATGTCTTCCAAAAGTCCAAGACTAAGGAATGCTGTCTGGATATGTCTAGATCAAACAGTAGTGGTAGCCTTGGGGCTGATG
AAGGGCTGTTTGAGCTACCGGAATCCCTGGCTTCTAGCAATATTGTGTTGGAGAAAGCCTTGCGGCGGAGAAGTCAACATTTGTATGATAAGCAATTGTC
TTGGCAGGAGTCTCCAGAAGGAAGGAAAATGATGGAATTTCGAAAAACCCTTCCTGCATACAAGGAGAAGAATGCAATATTGACAGCTGTTGCACAGAAT
CAGGTGGTCATCATATCAGGTGAGACTGGTTGCGGTAAGACAACACAAATTCCCCAGTTTATCCTGGAATCTGTAATAGATGCTGCTCGTGGAGCTGCTT
GTAGTATTATATGCACACAACCAAGACGAATATCTGCGATCTCTGTTTCTGAAAGAGTTGCTACGGAGAGAGGAGAGAAACTTGGAGAATCTGTCGGTTA
TAAGGTTCGGCTTGAAGGTATTAAAGGCAGGGAAACACACCTCCTCTTCTGCACAACAGGAATCTTATTGAGAAGATTACTTGTGGATCGAAGCTTGAAG
GGAGTAAGCCATGTGATAGTGGACGAGATTCATGAGCGTGGAATGAATGAAGATTTTCTGCTTATTGTCCTTAAAGATCTGCTATCCCGTCGACCAGACC
TGAGGCTCGTTCTGATGAGTGCAACCCTTGATGCGGAGCTTTTCTCATCCTACTTTAACAGAGCTCCTATTCTCCGTATCCCAGGTTTTACATTCCCAGT
GAAAACTCATTTCCTGGAAAATATTCTGGAAATGACGGGATACCGTTTGACTCCATACAATCAAATTGACGATTATGGTCAAGAGAGGGCATGGAAACTA
AATAAGCAAGGTCCAAAAAGGAGAAAGAGCCAAATTGCGTCGGCAGTTGAGGATGCACTCAGAACAGCTGATTTCAGGAAATACAGAAAAGAGACTCGAG
ATTCTCTGTCTTTATGGAATCCTGATTGCATTGGTTTTAATCTGATCGAGTATCTACTGAGCTACATCTGTGAAAATGAGAGACCTGGTGCTATTCTGGT
TTTTTTGACTGGGCTTGACGACATAAGTTCTTTAAAGGACAAACTCCAAGCTCATACCATCTTGGGAGATCCTAGACGAGTCTTGTTGCTCACTTGCCAT
GGCTCAATGGCCAGCTCTGAGCAGAGGTTAATATTTGATGTGCCACCTTATGGCGTGAGAAAAATAGTGCTGGCCACGAATATTGCCGAGACAAGCCTCA
CAATAAATGATGTTGTTTATGTTATTGAGTGTGGGAAAGCAAAGGAGACTTCATACGATGCTCTCAACAACACTCCTTGTCTGCTCCCTTCTTGGATTTC
TAAAGTTTCTGCTCAACAAAGAAGAGGAAGAGCTGGCCGGGTTCAACCGGGAGAATGTTACCATCTATATCCAAGATGTGTATATGATGCTTTTGCGGAG
TATCAGTTGCCAGAAATCTTGAGAACGCCTTTACAGTCACTCTGTTTGCAAATCAAGAGTTTGAAACTTGGAAGTATCTCTGAGTTTCTGTCTAAGGCTT
TGCAATCACCCGAGCTACTGGCGGTTGAGAAAGCTATTGAGTACTTGAAAGTTATAGGCGCCTTGGATAAAAAAGAAGATTTGACTGTGCTGGGGAAGTA
CTTGACTAGGCTTCCCGTGGAGCCTAAACTCGGAAAGATGCTCGTTTTAGGTGCTATCTTCAATTGCCTGGATCCGATTTCAACCATTGTTGCAGGACTA
AGTTTTCGAGATCCTTTCTTAACACCCCTTGACAAGAAAGATCTTGCAGATGCCGCGAAAGCTCAGTTTTCACGTGACTATAGCGACCACATGGCCCTTG
TTCGGGCCTTTAATGGCTGGAAAGATGCTGAAAGGAATTTCGCTGGTCATGACTATTGCTGGAAGAATTTCCTCTCTGTTCAATCCATGAAGGCTATTGA
ATCTCTCAGGAAGGAGTTCTTCTCCTTGATAAGGGATATTGGCCTGATTGATAGTAATGCACACAGTAACGGTTTGAGCCACGAGGAGCACTTTGTTCGA
GCAGTTATTTGTTACGGCCTTTACCCCGGAGTGTGCTCTGTCGTGAACAATGAAAAGTCGTTTTCGCTCAAAACTATGGAGGATGGTCAGGTGCTGCTAT
ATTCTAACTCTGTCAATGCTCAGAATTCCAGCATCCCCTATCCATGGTTAGTTTTTAACGAGAAGGTCAAAGTCAATGCTGTCTTCCTCCGCGACTCCAC
AGCTGTTTCAGACTCAATGTTGCTTCTTTTCGGTGGAACCGTCACGAAAGGGGATGCTGATGGTCACTTGAAGATGCTAGGAGGATATCTGGAATTCTTC
ATGAAACCGGCAGTGGCGGAAATGTATACCAACTTGAAAAGAGAACTCGATGAGCTTATTCAGATGAAACTACTCAATCCGAAAATGAGCTTAAAGCCAC
ACGACGAGCTTTTCTCAGCAGTACGGTTACTCCTCTTTGAGGATCGCTCCGATGGAAGATTCTTATTTGGCCATCAGCTGCTCAACCTTACGAAACCCCC
CGTGGTGGCTTCAACGACAAGAAAGCCGATTCTGGCATCAGGAGGTGACAGTGGACCCGGAGGCGATAACCCAAAGAGCCAACTCCAAACGCTGCTGACG
AGAGCCGGGTACACTGCGCCGGTATACAAAACGAAGCAACTGAAGAACAACCAGTTCCGGTCCACCGTGGAGTTCAACGGCATGGAAGTGATGGGGAACC
CTTACGGAAACAAGAAGAGCGCGGAGAAGGATGCCGCGGCAGAGGCCCTGCTGTGGTTGATGGGCGGATCTCAGACCGGTCGTGATTACATCGACCAAAT
GTCGATTCTGCAGAAGAGAAGCAAGAAGAATCATTTCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10007938 pacid=23162297 polypeptide=Lus10007938 locus=Lus10007938.g ID=Lus10007938.BGIv1.0 annot-version=v1.0
MKDRPPSPYGAVYVPPHQRLRSVIVSPNFNSSSASSHLLESRTTDSRCAALSRSKASSPSPTEHQQRNGGAAEVCNKNPIRKFASACVNAVSVDGSDHDM
ESLPAHPSGTYMSDSTEEWRWKFTMLLRDKEKKEVVSREKKDRRDFEQLAALASKMGLYSRVYVKVVVFSKVPLPNYRFDLDDKRPQREVSLPLGVLKRV
DTYLGDYVFQKSKTKECCLDMSRSNSSGSLGADEGLFELPESLASSNIVLEKALRRRSQHLYDKQLSWQESPEGRKMMEFRKTLPAYKEKNAILTAVAQN
QVVIISGETGCGKTTQIPQFILESVIDAARGAACSIICTQPRRISAISVSERVATERGEKLGESVGYKVRLEGIKGRETHLLFCTTGILLRRLLVDRSLK
GVSHVIVDEIHERGMNEDFLLIVLKDLLSRRPDLRLVLMSATLDAELFSSYFNRAPILRIPGFTFPVKTHFLENILEMTGYRLTPYNQIDDYGQERAWKL
NKQGPKRRKSQIASAVEDALRTADFRKYRKETRDSLSLWNPDCIGFNLIEYLLSYICENERPGAILVFLTGLDDISSLKDKLQAHTILGDPRRVLLLTCH
GSMASSEQRLIFDVPPYGVRKIVLATNIAETSLTINDVVYVIECGKAKETSYDALNNTPCLLPSWISKVSAQQRRGRAGRVQPGECYHLYPRCVYDAFAE
YQLPEILRTPLQSLCLQIKSLKLGSISEFLSKALQSPELLAVEKAIEYLKVIGALDKKEDLTVLGKYLTRLPVEPKLGKMLVLGAIFNCLDPISTIVAGL
SFRDPFLTPLDKKDLADAAKAQFSRDYSDHMALVRAFNGWKDAERNFAGHDYCWKNFLSVQSMKAIESLRKEFFSLIRDIGLIDSNAHSNGLSHEEHFVR
AVICYGLYPGVCSVVNNEKSFSLKTMEDGQVLLYSNSVNAQNSSIPYPWLVFNEKVKVNAVFLRDSTAVSDSMLLLFGGTVTKGDADGHLKMLGGYLEFF
MKPAVAEMYTNLKRELDELIQMKLLNPKMSLKPHDELFSAVRLLLFEDRSDGRFLFGHQLLNLTKPPVVASTTRKPILASGGDSGPGGDNPKSQLQTLLT
RAGYTAPVYKTKQLKNNQFRSTVEFNGMEVMGNPYGNKKSAEKDAAAEALLWLMGGSQTGRDYIDQMSILQKRSKKNHF
|
DESeq2's median of ratios [FLAX]
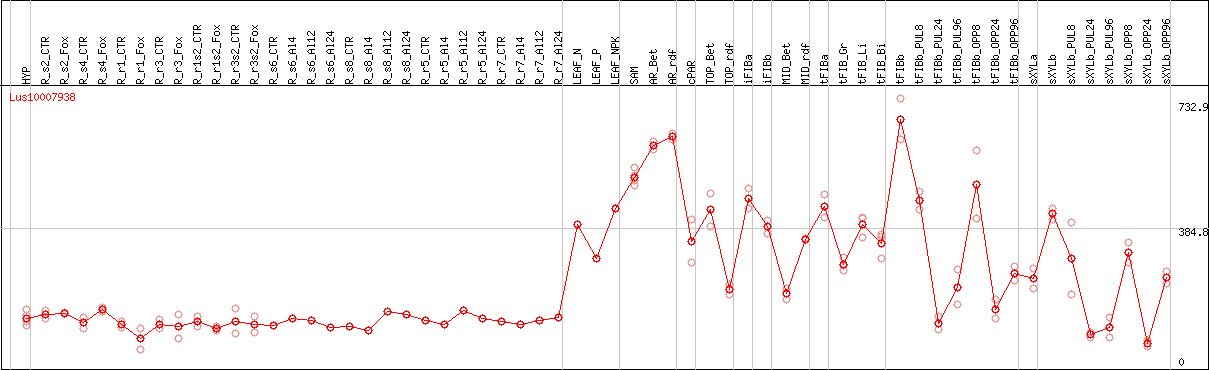
Coexpressed genes
Lus10007938 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
