External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G15140 237 / 1e-79
FAD/NAD(P)-binding oxidoreductase (.1), FAD/NAD(P)-binding oxidoreductase (.2), FAD/NAD(P)-binding oxidoreductase (.3)
AT1G20020 45 / 6e-06
ATLFNR2
ferredoxin-NADP\(+\)-oxidoreductase 2, LEAF FNR 2, ferredoxin-NADP(+)-oxidoreductase 2 (.1), ferredoxin-NADP(+)-oxidoreductase 2 (.2), ferredoxin-NADP(+)-oxidoreductase 2 (.3)
AT5G17770 41 / 0.0001
CBR1, ATCBR
NADH:cytochrome B5 reductase 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013488
280 / 5e-99
AT1G15140 244 / 1e-82
FAD/NAD(P)-binding oxidoreductase (.1), FAD/NAD(P)-binding oxidoreductase (.2), FAD/NAD(P)-binding oxidoreductase (.3)
Lus10035265
215 / 4e-66
AT1G15130 836 / 0.0
Endosomal targeting BRO1-like domain-containing protein (.1)
Lus10034633
187 / 5e-60
AT1G15140 316 / 3e-108
FAD/NAD(P)-binding oxidoreductase (.1), FAD/NAD(P)-binding oxidoreductase (.2), FAD/NAD(P)-binding oxidoreductase (.3)
Lus10014491
41 / 0.0001
AT5G20080 475 / 1e-170
FAD/NAD(P)-binding oxidoreductase (.1)
Lus10030059
40 / 0.0002
AT5G20080 516 / 0.0
FAD/NAD(P)-binding oxidoreductase (.1)
Lus10005682
40 / 0.0002
AT5G17770 474 / 6e-171
NADH:cytochrome B5 reductase 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G126700
238 / 2e-80
AT1G15140 395 / 9e-140
FAD/NAD(P)-binding oxidoreductase (.1), FAD/NAD(P)-binding oxidoreductase (.2), FAD/NAD(P)-binding oxidoreductase (.3)
Potri.010G116500
213 / 6e-70
AT1G15140 368 / 1e-128
FAD/NAD(P)-binding oxidoreductase (.1), FAD/NAD(P)-binding oxidoreductase (.2), FAD/NAD(P)-binding oxidoreductase (.3)
Potri.013G067300
44 / 1e-05
AT5G17770 461 / 6e-166
NADH:cytochrome B5 reductase 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0091
NAD_Ferredoxin
PF00175
NAD_binding_1
Oxidoreductase NAD-binding domain
Representative CDS sequence
>Lus10007955 pacid=23162303 polypeptide=Lus10007955 locus=Lus10007955.g ID=Lus10007955.BGIv1.0 annot-version=v1.0
ATGGGGAAAGGATTCGGTATCAATCAAATCGATCCGCCTGTGAAGTACTCTACCGTTCTAATTTTTGCCACCGGCTCCGGAATCAGTCCAATTCGATCTC
TAATTGAGACAGGGTTTAGTGCTGACCAAAGATCTGATGTGAGGCTTTACTATGGAGCTAGAAACCTTAAGAGGATGGCTTACCAGGATAGGTTCAAAGA
ATGGGAGTCGTCTGGTGTTAAGATTATCCCAGTACTATCACAACCAGATGGTACCTGGAAGGGTGAAGCTGGCTATGTACAGACTGCTTTCGCTAGCGCC
ACGCAGATATCCAGCCCAGCAGGCACAGGTGCTGTGCTTTGTGGGCAGAAACAGATGGCAGAGGAGGTGACATCGATTCTTTTAGCAGAGGGAGTCTCGA
CAGAGAGAATACTGAAGAACTTCTGA
AA sequence
>Lus10007955 pacid=23162303 polypeptide=Lus10007955 locus=Lus10007955.g ID=Lus10007955.BGIv1.0 annot-version=v1.0
MGKGFGINQIDPPVKYSTVLIFATGSGISPIRSLIETGFSADQRSDVRLYYGARNLKRMAYQDRFKEWESSGVKIIPVLSQPDGTWKGEAGYVQTAFASA
TQISSPAGTGAVLCGQKQMAEEVTSILLAEGVSTERILKNF
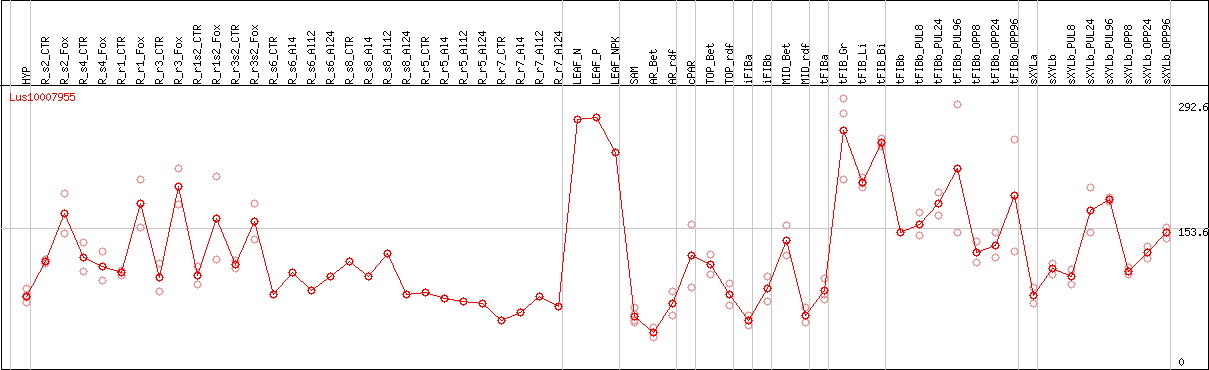
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007955 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.