External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G60910 290 / 2e-99
MADS
FUL, AGL8
FRUITFULL, AGAMOUS-like 8 (.1.2)
AT1G69120 285 / 3e-97
MADS
AGL7, AP1
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
AT1G26310 261 / 9e-88
MADS
CAL1, AGL10, CAL
CAULIFLOWER, AGAMOUS-like 10, K-box region and MADS-box transcription factor family protein (.1)
AT3G30260 204 / 2e-65
MADS
AGL79
AGAMOUS-like 79 (.1)
AT1G24260 174 / 1e-53
MADS
AGL9, SEP3
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
AT5G15800 174 / 2e-53
MADS
AGL2, SEP1
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
AT3G02310 172 / 8e-53
MADS
AGL4, SEP2
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
AT2G45650 168 / 2e-51
MADS
AGL6
AGAMOUS-like 6 (.1)
AT3G61120 163 / 2e-49
MADS
AGL13
AGAMOUS-like 13 (.1)
AT2G03710 160 / 7e-49
MADS
AGL3, SEP4
SEPALLATA 4, AGAMOUS-like 3, K-box region and MADS-box transcription factor family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021140
441 / 1e-158
AT5G60910 291 / 1e-99
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10034662
340 / 6e-119
AT5G60910 284 / 3e-97
FRUITFULL, AGAMOUS-like 8 (.1.2)
Lus10017871
318 / 3e-111
AT1G69120 264 / 2e-90
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10026679
272 / 2e-92
AT1G69120 338 / 1e-118
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10004637
271 / 2e-92
AT1G69120 318 / 2e-111
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10005081
242 / 9e-80
AT1G69120 223 / 1e-72
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10034370
231 / 4e-76
AT1G69120 224 / 1e-73
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Lus10034663
189 / 3e-59
AT5G15800 267 / 4e-90
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Lus10003124
176 / 5e-54
AT1G24260 355 / 6e-125
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G062300
322 / 6e-112
AT5G60910 288 / 5e-99
FRUITFULL, AGAMOUS-like 8 (.1.2)
Potri.010G154100
284 / 7e-97
AT1G69120 324 / 7e-113
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Potri.008G098500
282 / 3e-96
AT1G69120 336 / 9e-118
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
Potri.004G115400
269 / 6e-91
AT5G60910 230 / 7e-76
FRUITFULL, AGAMOUS-like 8 (.1.2)
Potri.017G099800
268 / 1e-90
AT5G60910 255 / 7e-86
FRUITFULL, AGAMOUS-like 8 (.1.2)
Potri.008G098400
192 / 5e-61
AT5G15800 275 / 2e-93
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Potri.004G115500
174 / 8e-54
AT3G02310 357 / 4e-126
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Potri.014G074100
174 / 1e-53
AT2G45650 311 / 5e-108
AGAMOUS-like 6 (.1)
Potri.017G099700
173 / 3e-53
AT3G02310 372 / 8e-132
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Potri.003G169600
172 / 3e-53
AT1G24260 359 / 7e-127
SEPALLATA3, AGAMOUS-like 9, K-box region and MADS-box transcription factor family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01486
K-box
K-box region
PF00319
SRF-TF
SRF-type transcription factor (DNA-binding and dimerisation domain)
Representative CDS sequence
>Lus10007983 pacid=23162332 polypeptide=Lus10007983 locus=Lus10007983.g ID=Lus10007983.BGIv1.0 annot-version=v1.0
ATGGGGAGAGGGAGAGTGCAGCTGAGGAGGATCGAGAACAAGATCAACAGGCAAGTGACGTTCTCCAAGAGGAGGTCAGGTCTTCTGAAGAAGGCTCATG
AGATCTCTGTACTCTGTGATGCTGAAGTGGCTCTCATCGTCTTCTCCACCAAGGGCAAGCTCTTTGAATACTCCACTGATTCCTGTATGGAGAGGATCCT
GGAACGTTATGAGAGATACTCCTATGCTGAGAGGCAGCTTCTTGCCAATGATACTGAGACATGCGGTAGCTGGACTCTAGAATATGCCAAGCTCAAGGCT
AGGATTGAAGTCCTACAAAAGAACCTAAGGAACTACAACGGCCAAGACCTTGATTCCTTGAGCCTTAGGGAGCTTCAGAACCTGGAACAACAACTTGATT
CTGCTCTTAAACATATCCGCTCCAGAAAGAACCAACTGATGTATGAATCCATCTCGGAGCTTCAGAAAAAGGACAAGGCATTGCAGGACCAAAACAACCA
ACTGGCAAAAAAGGTTAAAGAAAGGGAGAAGGAAATATTGGAGCAACCTCAGTGGGAGCAACAACAGGATCCACCCTCTCTGGTTACTCAACAAATGGAG
GACTCACCCTCCAGCATCGTCGTTGCACCACAGCCATTCCAGCCTCTAGACATAAGTGGTGGCAGCGGTGGAGGTGGCGGAGGAGGTTATCAGCTGGCAA
GGGGAAACACAGGAGTTGAAGAAGGAACGAGTAGTTCGTCGTCCCAGCATCGACCCAGTGCAGTGCTGCCTTCATGGATGATGGGAAACCATATATAG
AA sequence
>Lus10007983 pacid=23162332 polypeptide=Lus10007983 locus=Lus10007983.g ID=Lus10007983.BGIv1.0 annot-version=v1.0
MGRGRVQLRRIENKINRQVTFSKRRSGLLKKAHEISVLCDAEVALIVFSTKGKLFEYSTDSCMERILERYERYSYAERQLLANDTETCGSWTLEYAKLKA
RIEVLQKNLRNYNGQDLDSLSLRELQNLEQQLDSALKHIRSRKNQLMYESISELQKKDKALQDQNNQLAKKVKEREKEILEQPQWEQQQDPPSLVTQQME
DSPSSIVVAPQPFQPLDISGGSGGGGGGGYQLARGNTGVEEGTSSSSSQHRPSAVLPSWMMGNHI
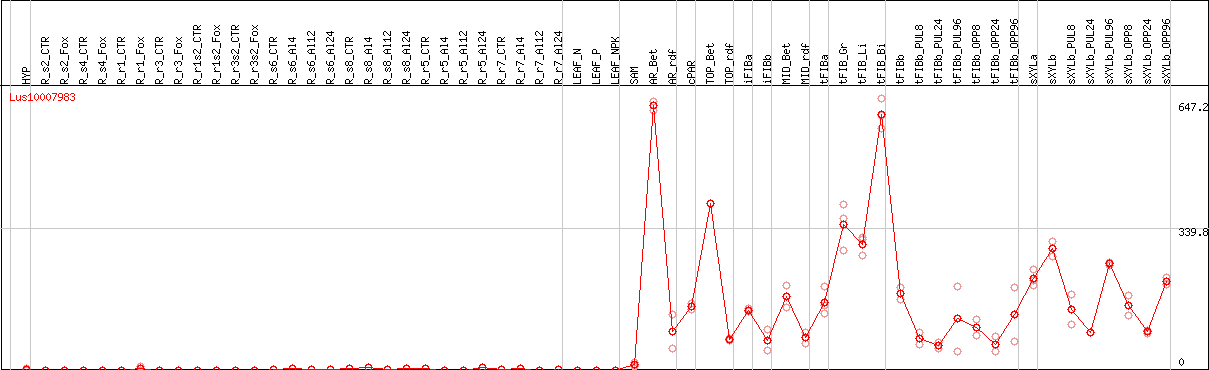
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007983 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.