Lus10007993 [FLAX]

| External link |
|
||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||
|
Representative CDS sequence |
>Lus10007993 pacid=23164440 polypeptide=Lus10007993 locus=Lus10007993.g ID=Lus10007993.BGIv1.0 annot-version=v1.0
ATGTTCAATGCTTGCTGGAAAGTGTATGTCGGGCGCTGGAGCGGGAGTTCCTGTGGAGTGGAGTTTGATGTCCCATCTTCCGATTCACGAACACAGTACA
AGAGAAGGCGAATTTCCTCCCGGGACTTAGGAGAGGTAGAAGCTTCGTTACCCCTTTTAACTTCACCAGATTATTACACGGAGCCTTGCTTGGCCAATTT
GGCTGCCAGGGAATTTCTAGATCCTGGTTATTGTGGTCGAGTTCCCAATTTCACTGTTGGGAAGTTTGGTTATGGGTGTGTCAAGTTTCTTGGAAACACT
GATGTTAGGAAGCTAGAAGTGGATCACATTGTTAAGTTCAATAGGCATCAAATTGTCGTATATGGAGATGAAAGTACCAAGCCTGCAGTTGGTCAGGGGC
TCAACAAGCCTGCTGAAGTGACTCTGAAATTAAAATTAAGACCATTTGATTCATCTAAACAGCATTACAATGCTATTGTGAAGAAACTGAGGTTAAGCTT
GGAGAGGCAGGGAGGTAGGTTCATTTCTTTCAACCCCAATAATGGTAATTGGAGTTTCTTAGTCCAACATTTTAGTCGATTTGGATTGACTGAAGATGAT
GAAGAAGACATCGGTATGGATGATGTTGAGGAAGTTCATGATCCTTCCAAGATCAACGATGCTGAGATCATGGAGAATGGGGAATCACAAGAAGAAATGG
AACCTACTGGGCCTATGCTTTATCATTCTCTACCTGCACATCTTGGCCTTGACCCAATAAAGATGAGAGAGATGAAGATGTTGATGTTTCCTGTTGAGGA
TGAGGAAGAGGTTGAAGATTTGACTGACCTCTCTCATCCCAACTTGTCCAGAAGTAAAGAACATATGAGACGTTCATTTCTGAATAGTTCTCAGAGAATG
AGTCACAGATCTAGTATGCCTGTGGTGAGGAAACCACCATTGCCTCTGCTTGAATACAATCTTGCTGAGAACAACTCAGATTCTCCTGGAAACATATTAA
TGGCCCAACAAAACAAGGGCTTATCTTTAAAGCCAATTAGAGGTGAGGGCTTCAAGCTGAACCTGGAACGCCAGACACCAGTAAGTGGAAGTCATTCTCG
CAACATAGTTGATGCTGGTTTGTTCATGGGCAAATCATTCCGCGTTGGGTGGGGTCCAAATGGAGTTCTTCTCCATTCGGGTACTCCTGTAAGAAGTAAC
AGCTCTGAAAGATGGTTATCTTCTGTCATAAACATAGAGAAGGTTGGCATTGACAAAGTTGTTAGAGATGAAAATGATAAAACCAGAAAGGAACTGGTTG
AGTTTGCGTTTGATGCACCCTTGAATCTCCATAAGTCACTAAATCATGATCTGAAACAAGTTCAAGTTGGGTCATATGCTCTGAAGCTTCAAAAGGTAAT
CACGAATAAGTTGGCTCTTGCTGAAATTTGTCGGAGTTATATAGATATAATTGAGGGGCAGCTGGAGGTTCCTGGACTGTCATCATCCTCACGTTTGGTT
TTGATGCATCAAGTGCTTATCTGGGAGTTGATCAAAGTTCTCTTTTCCGAGAGAGAAAATAGTGACCCACCAAAATCTTTGGCTGCTGATAACGAGGAAG
ATATGATGCCAGATGTGAAAGAAGGTAGTCAACAAATAGACCCAGAAGCAGTTTCGCTTATTAGAAGAGCAGAGTTCAGCTGTTGGTTGCAAGAAAGTGT
ATGCCATCGTGTCCAAGAGGATGTGAGCGCTTTGGATGAATCCAGTTATCTCAGACACATTTTTTTACTTCTTACTGGGAGGCAACTCGATGCTGCTGTT
GAAATGGCTGTTTCTAGAGGGGATGTAAGGTTGGCTTGTTTGCTCAGTCAGGCTGGTGGATCGACTGTTAATCGTGATGATATTGCACACCAGCTTGATC
TTTGGAGGACTAATGGGCTGGATTTCAATTTTGTTGAGAAGGAGAGGACTAGGCTTTATGAGCTGCTTTCTGGTAATATTCACAATGCCTTGGAGGGTGT
TAAAATTGACTGGAAAAGATTCTTAGGATTGTTGATGTGGTATCGTCTACCACCTGATACACCATTGCCTGTCATTTTCCAAAATTATCAACTTCTCCTC
CATGAAGGTAAAGCTCCATTTCCTCTTCCAATTTATATTGATGAAGGCCCAGTGGAAGAGGCACTGGGAGTTGCGGAAAAGAACATTGATCTCACATATT
ATCTCATGCTTCTTCATGCCAATGGGGAGGGTAAATTTGGCTATCTGAAGACCATGCTCAGTAATTTCTCTTCAACATATGATCCACTGGACTACCATAT
GATTTGGCATCAGAGAGCAGTGCTAGAGGCAGCTGGTGTGTTTACTTCTAGTGATCTTCAAGTGCTTGACATGGGGATTGTTTCCCAGTTGTTGGGTACT
GGACAATGTCATTGGGCCATCTATGTGGTGCTTCACATGCCACAGTGTGATGATTATCCACACTTCCATGCTTCTGTTATTCGAGAAATATTATTTCAGT
ATTGTGAAACTTGGAGTTCAGATGAATCGCAACACCAATTCATTCAGAACCTTGAAATCCCATCAGAGTGGCTGCATGAAGCTATGGCAGTATATTTCAG
TTATTACGGGAATCTGTCCAAGGCCCTTGAACACTTCCTTGAATGTGCAAAATGGCAGAAGGCTCATTCTATTTTTACAACCTCGGTTGCTCATACATTG
TTTTTGTCTGGTGATCATTCAGAAGTTTGGAGGCTTGCTACATTAATGGAGGAGCACAAATCTCAAATTGAAGACTGGGAATTGGGAGCAGGAATCTACA
TTTCCTTCTTCGTGCTTAAAAGTTCATTTGAAGAGGATAACAACACAATGAACGAAGAGGACCCTCTTGCAAGCAAAATTTCCGACTGTGACGACTTCCT
TAGTCAACTTAATGAATCTCTGGCAGTTTTGGGTGACCGGTTGCCTACTGATGCAAGGATTGCGTACTCAAAGATGGGAGAGGAGGTATCTGAGATGCTG
CTCTCAATTAGTAGTTTGGGCGAGTCACGGGACAGCCAGTTGAAATGCTTTGACACAGTTCTCAAGGCTCCAGTACCTGAAGATGTATGCTCAAATCATC
TGCAGGATGCCGTCTCTCTTTTTACATGTTATCTATCGGAGATGGCCGCACAATCCGTTTAA
|
||||||||||||||||||||
|
AA sequence
|
>Lus10007993 pacid=23164440 polypeptide=Lus10007993 locus=Lus10007993.g ID=Lus10007993.BGIv1.0 annot-version=v1.0
MFNACWKVYVGRWSGSSCGVEFDVPSSDSRTQYKRRRISSRDLGEVEASLPLLTSPDYYTEPCLANLAAREFLDPGYCGRVPNFTVGKFGYGCVKFLGNT
DVRKLEVDHIVKFNRHQIVVYGDESTKPAVGQGLNKPAEVTLKLKLRPFDSSKQHYNAIVKKLRLSLERQGGRFISFNPNNGNWSFLVQHFSRFGLTEDD
EEDIGMDDVEEVHDPSKINDAEIMENGESQEEMEPTGPMLYHSLPAHLGLDPIKMREMKMLMFPVEDEEEVEDLTDLSHPNLSRSKEHMRRSFLNSSQRM
SHRSSMPVVRKPPLPLLEYNLAENNSDSPGNILMAQQNKGLSLKPIRGEGFKLNLERQTPVSGSHSRNIVDAGLFMGKSFRVGWGPNGVLLHSGTPVRSN
SSERWLSSVINIEKVGIDKVVRDENDKTRKELVEFAFDAPLNLHKSLNHDLKQVQVGSYALKLQKVITNKLALAEICRSYIDIIEGQLEVPGLSSSSRLV
LMHQVLIWELIKVLFSERENSDPPKSLAADNEEDMMPDVKEGSQQIDPEAVSLIRRAEFSCWLQESVCHRVQEDVSALDESSYLRHIFLLLTGRQLDAAV
EMAVSRGDVRLACLLSQAGGSTVNRDDIAHQLDLWRTNGLDFNFVEKERTRLYELLSGNIHNALEGVKIDWKRFLGLLMWYRLPPDTPLPVIFQNYQLLL
HEGKAPFPLPIYIDEGPVEEALGVAEKNIDLTYYLMLLHANGEGKFGYLKTMLSNFSSTYDPLDYHMIWHQRAVLEAAGVFTSSDLQVLDMGIVSQLLGT
GQCHWAIYVVLHMPQCDDYPHFHASVIREILFQYCETWSSDESQHQFIQNLEIPSEWLHEAMAVYFSYYGNLSKALEHFLECAKWQKAHSIFTTSVAHTL
FLSGDHSEVWRLATLMEEHKSQIEDWELGAGIYISFFVLKSSFEEDNNTMNEEDPLASKISDCDDFLSQLNESLAVLGDRLPTDARIAYSKMGEEVSEML
LSISSLGESRDSQLKCFDTVLKAPVPEDVCSNHLQDAVSLFTCYLSEMAAQSV
|
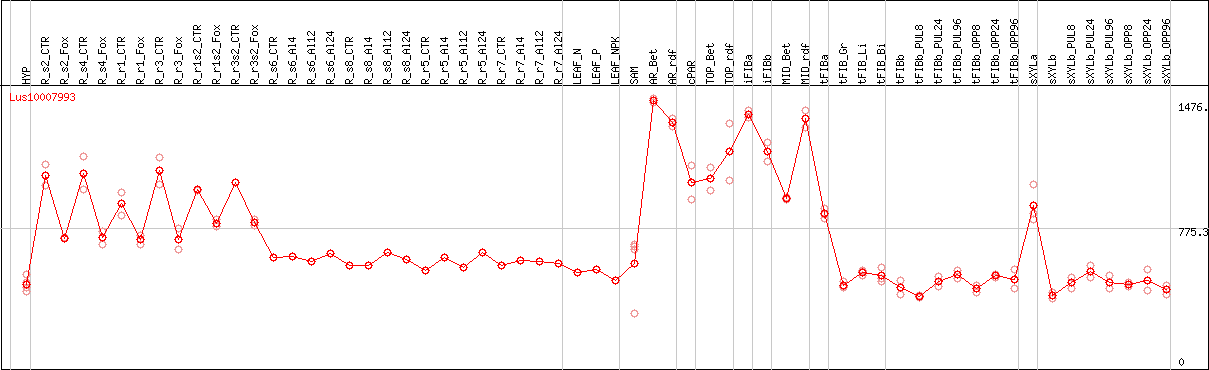
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10007993 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
