External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G56680 267 / 3e-87
SYNC1ARATH, SYNC1, EMB2755
EMBRYO DEFECTIVE 2755, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
AT1G70980 266 / 5e-87
SYNC3, SYNC1
Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
AT4G17300 204 / 6e-63
ATNS1, NS1, OVA8
ovule abortion 8, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
AT3G07420 179 / 3e-53
ATNS2, SYNC2_ARATH, NS2
SYNTHETASE C2, asparaginyl-tRNA synthetase 2 (.1)
AT4G31180 65 / 2e-12
Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1.2)
AT4G26870 62 / 2e-11
Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012920
287 / 2e-95
AT5G56680 799 / 0.0
EMBRYO DEFECTIVE 2755, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
Lus10032704
285 / 7e-94
AT5G56680 907 / 0.0
EMBRYO DEFECTIVE 2755, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
Lus10038228
197 / 1e-62
AT3G07420 353 / 7e-118
SYNTHETASE C2, asparaginyl-tRNA synthetase 2 (.1)
Lus10025874
201 / 3e-61
AT3G07420 604 / 0.0
SYNTHETASE C2, asparaginyl-tRNA synthetase 2 (.1)
Lus10007470
164 / 1e-47
AT4G17300 873 / 0.0
ovule abortion 8, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
Lus10028945
157 / 9e-46
AT4G17300 740 / 0.0
ovule abortion 8, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
Lus10028555
65 / 2e-12
AT4G31180 606 / 0.0
Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1.2)
Lus10018862
61 / 6e-11
AT4G31180 667 / 0.0
Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1.2)
Lus10028557
57 / 3e-10
AT4G31180 235 / 8e-76
Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G070000
274 / 6e-90
AT5G56680 931 / 0.0
EMBRYO DEFECTIVE 2755, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
Potri.006G153900
273 / 2e-89
AT5G56680 929 / 0.0
EMBRYO DEFECTIVE 2755, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
Potri.002G249700
197 / 2e-59
AT3G07420 654 / 0.0
SYNTHETASE C2, asparaginyl-tRNA synthetase 2 (.1)
Potri.007G129800
193 / 1e-58
AT4G17300 872 / 0.0
ovule abortion 8, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
Potri.017G028800
192 / 3e-58
AT4G17300 888 / 0.0
ovule abortion 8, Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1)
Potri.006G085400
64 / 4e-12
AT4G31180 709 / 0.0
Class II aminoacyl-tRNA and biotin synthetases superfamily protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0040
tRNA_synt_II
PF00152
tRNA-synt_2
tRNA synthetases class II (D, K and N)
Representative CDS sequence
>Lus10008180 pacid=23173779 polypeptide=Lus10008180 locus=Lus10008180.g ID=Lus10008180.BGIv1.0 annot-version=v1.0
ATGGATCAAATCCAAAATCCATTTCTTTGGCTCCAGATTCGGACTCAGGATGACAAGAACTGTGCAGAAGCATACGTAAAGTATATGTGCAAATGGCTGC
TTGATAAGTGTTACGACGACATGGAGGCTGTGAAGGATAGCACTAAGACGTTTGATAACCAAGTTGAATGGGGCATTGATCTCGCATCTGAACATGTGAG
GTTTCTGGCAGAAGAAAAGTTCGAGAAGCCGGTTATTGTGTACGATTATCCTAAAGGAATCAAAGCCTTCTACATGAGGCTGAACGATGATAAGAAGACG
GTGGCTGCCATGGATATTCTTGTACCAAAGGTGGGAGAATTGATTGGTGGAAGTCAAAGGGAGGAGCGATATGAAGTGATGGAAGAGAGGATTGTGGAGA
TGGGGCTGCCACTGGAGGCATACAAGTGGTACCTCGACTTGAGGCAGTTTGGGACCATAAAGCATAGTGGGTTCGGGTTAGGATTCGACGGGATGATACT
GTTTGCGACTGGTATTGACAATATCAGAGAAGTTATTCCCTTCCGCAGAAGTGCACCACCCAGAAGTACAGAGAAAGCAGATTTGTGA
AA sequence
>Lus10008180 pacid=23173779 polypeptide=Lus10008180 locus=Lus10008180.g ID=Lus10008180.BGIv1.0 annot-version=v1.0
MDQIQNPFLWLQIRTQDDKNCAEAYVKYMCKWLLDKCYDDMEAVKDSTKTFDNQVEWGIDLASEHVRFLAEEKFEKPVIVYDYPKGIKAFYMRLNDDKKT
VAAMDILVPKVGELIGGSQREERYEVMEERIVEMGLPLEAYKWYLDLRQFGTIKHSGFGLGFDGMILFATGIDNIREVIPFRRSAPPRSTEKADL
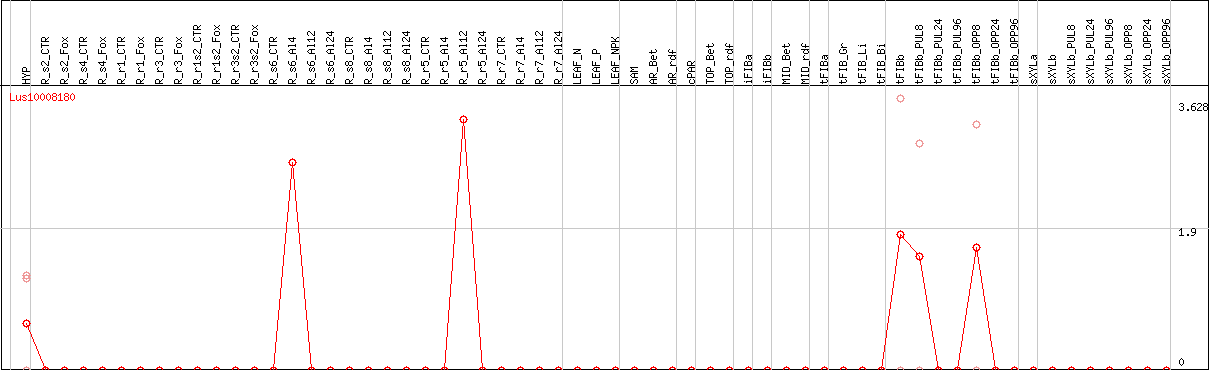
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10008180 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.