External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G63010 206 / 7e-67
ATGID1B, GID1B
GA INSENSITIVE DWARF1B, alpha/beta-Hydrolases superfamily protein (.1)
AT5G27320 198 / 6e-64
ATGID1C, GID1C
GA INSENSITIVE DWARF1C, alpha/beta-Hydrolases superfamily protein (.1)
AT3G05120 192 / 8e-62
ATGID1A, GID1A
GA INSENSITIVE DWARF1A, alpha/beta-Hydrolases superfamily protein (.1)
AT5G23530 86 / 8e-21
ATCXE18
carboxyesterase 18 (.1)
AT5G16080 63 / 1e-12
ATCXE17
carboxyesterase 17 (.1)
AT5G06570 63 / 1e-12
alpha/beta-Hydrolases superfamily protein (.1.2)
AT1G68620 55 / 8e-10
alpha/beta-Hydrolases superfamily protein (.1)
AT5G14310 45 / 4e-06
ATCXE16
carboxyesterase 16 (.1)
AT2G45600 38 / 0.0007
alpha/beta-Hydrolases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10027969
265 / 6e-90
AT3G63010 566 / 0.0
GA INSENSITIVE DWARF1B, alpha/beta-Hydrolases superfamily protein (.1)
Lus10002254
189 / 2e-60
AT5G27320 550 / 0.0
GA INSENSITIVE DWARF1C, alpha/beta-Hydrolases superfamily protein (.1)
Lus10000928
188 / 4e-60
AT5G27320 553 / 0.0
GA INSENSITIVE DWARF1C, alpha/beta-Hydrolases superfamily protein (.1)
Lus10013774
106 / 2e-28
AT5G23530 315 / 2e-106
carboxyesterase 18 (.1)
Lus10039162
96 / 2e-24
AT5G23530 324 / 3e-110
carboxyesterase 18 (.1)
Lus10018963
88 / 5e-22
AT5G27320 365 / 3e-127
GA INSENSITIVE DWARF1C, alpha/beta-Hydrolases superfamily protein (.1)
Lus10013771
76 / 1e-18
AT5G23530 96 / 4e-25
carboxyesterase 18 (.1)
Lus10036168
71 / 2e-15
AT5G06570 321 / 2e-109
alpha/beta-Hydrolases superfamily protein (.1.2)
Lus10013377
69 / 2e-14
AT3G44050 1290 / 0.0
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0028
AB_hydrolase
PF07859
Abhydrolase_3
alpha/beta hydrolase fold
Representative CDS sequence
>Lus10008189 pacid=23173766 polypeptide=Lus10008189 locus=Lus10008189.g ID=Lus10008189.BGIv1.0 annot-version=v1.0
ATGTTCGGCGGGCAGGAGAGGACCGAATCGGAAAGGTGCTTAGACGGGAAGTACTTCGTCACGATTCAAGATAGGGATTGGTACTGGAGGGCCTTTTTGC
CGGAAGGTGAGGATAGAGATCATCCAGCCGTTAATATCTTCGGTCCCAGGAGTAAGAAATTGGACGGTGGGATTAAGCTTCCGAAGAGTTTGGTAGTTGT
GGCCGGTTTGGACCTGATCCAAGATTGGCAATTGGCGTACGTTGATGGCATGGAGAAAGCTGGGCATCAAGTGAAGCTGATTTTTCTCCAGGAAGCTACA
ATTGGGTTCTATTTCTTGCCCAATAACAAGCATTTCTACTGCCTTATGGAGGAAATCAAGAACTTTGTCAATCCTTCTAACTAA
AA sequence
>Lus10008189 pacid=23173766 polypeptide=Lus10008189 locus=Lus10008189.g ID=Lus10008189.BGIv1.0 annot-version=v1.0
MFGGQERTESERCLDGKYFVTIQDRDWYWRAFLPEGEDRDHPAVNIFGPRSKKLDGGIKLPKSLVVVAGLDLIQDWQLAYVDGMEKAGHQVKLIFLQEAT
IGFYFLPNNKHFYCLMEEIKNFVNPSN
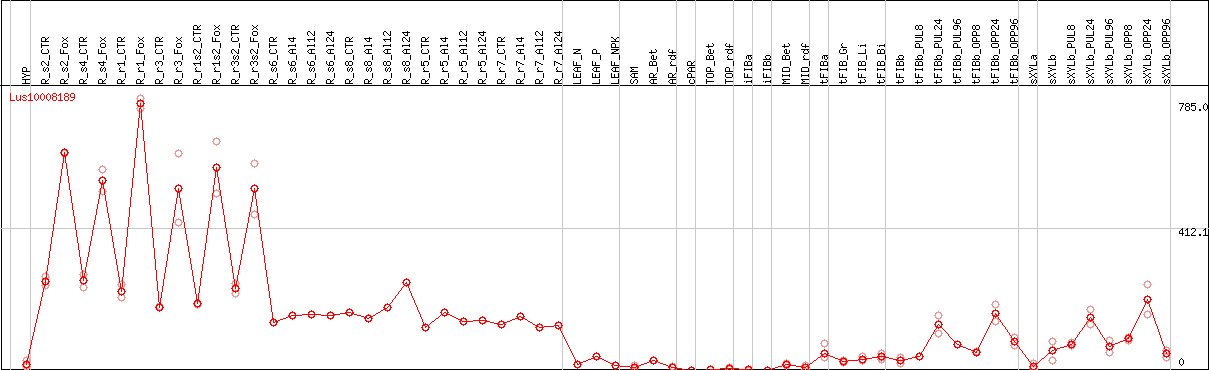
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10008189 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.