External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G04380 216 / 1e-68
SDG31, SUVR4
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
AT5G43990 151 / 9e-43
SDG18, SUVR2
SET DOMAIN PROTEIN 18, SET-domain containing protein lysine methyltransferase family protein (.1.2.3.4.5)
AT1G04050 144 / 7e-40
SDG13, SUVR1
SET DOMAIN PROTEIN 13, homolog of SU(var)3-9 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005476
356 / 5e-123
AT3G04380 503 / 3e-176
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Lus10012513
168 / 2e-50
AT3G04380 419 / 2e-143
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Lus10022657
160 / 6e-48
AT3G04380 412 / 2e-142
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Lus10022879
157 / 1e-44
AT5G43990 455 / 2e-149
SET DOMAIN PROTEIN 18, SET-domain containing protein lysine methyltransferase family protein (.1.2.3.4.5)
Lus10024946
155 / 1e-43
AT5G43990 488 / 7e-163
SET DOMAIN PROTEIN 18, SET-domain containing protein lysine methyltransferase family protein (.1.2.3.4.5)
Lus10029470
44 / 3e-05
AT1G73100 762 / 0.0
SET DOMAIN PROTEIN 19, SU(VAR)3-9 homolog 3 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G048100
213 / 6e-66
AT3G04380 471 / 5e-161
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Potri.009G138600
183 / 9e-55
AT3G04380 427 / 2e-144
SET DOMAIN PROTEIN 31, SET-domain containing protein lysine methyltransferase family protein (.1.2)
Potri.002G257400
172 / 9e-50
AT5G43990 524 / 8e-176
SET DOMAIN PROTEIN 18, SET-domain containing protein lysine methyltransferase family protein (.1.2.3.4.5)
Potri.014G192500
171 / 2e-49
AT5G43990 531 / 3e-178
SET DOMAIN PROTEIN 18, SET-domain containing protein lysine methyltransferase family protein (.1.2.3.4.5)
Potri.001G036800
42 / 0.0001
AT1G73100 796 / 0.0
SET DOMAIN PROTEIN 19, SU(VAR)3-9 homolog 3 (.1)
Potri.003G188700
42 / 0.0002
AT1G73100 806 / 0.0
SET DOMAIN PROTEIN 19, SU(VAR)3-9 homolog 3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF05033
Pre-SET
Pre-SET motif
Representative CDS sequence
>Lus10008198 pacid=23173786 polypeptide=Lus10008198 locus=Lus10008198.g ID=Lus10008198.BGIv1.0 annot-version=v1.0
ATGTGTAGAAAACTGCTTCATTGGTTGGTTTTAGCAGATGAAACCAGGAACATAAATCTTCGAAGGAACTATCATCACTACATACCCGACATAACGAAAG
GAGCTGAGAACGTCATGATTTTGTTAGTTGATGAATATGGCAGAGAGGAGGAGCTGCCGAAGTTCACCTACATACAGAACACCATCGTGCACCAAGACGC
TAAGTTGCACATTTGCTTGGCGCGGATTTCCGACATGGACTGCTGCTTGAGTTGCGTGGGAGATTGTGTTGCACAACTGGTGCCTTGCGCATGTTCCCGT
GAGACTGGTGGGGGATTCGCTTATACGAAACAAGGCCTTGTCAACGAAGAGTTTCTTGAAGCCTGTTTATCCATGGCTACCGATCCTGAAGAGCATCATA
TGTTTTTTTGCCAAGATTGTCCCTTGGCCAGGGTTAAGCGCCAGAATGTGCATGCAAGATGTGATGGCCATTTGATCAGGAAGTTCATCAAAGAGTGTTG
GGTCAAATGTGGATGTGAAATGAGCTGTGGAAATCGAGTTCTACAACGGGGTATCACTTGCAAATTACAGGTTAGAATTCTGTAG
AA sequence
>Lus10008198 pacid=23173786 polypeptide=Lus10008198 locus=Lus10008198.g ID=Lus10008198.BGIv1.0 annot-version=v1.0
MCRKLLHWLVLADETRNINLRRNYHHYIPDITKGAENVMILLVDEYGREEELPKFTYIQNTIVHQDAKLHICLARISDMDCCLSCVGDCVAQLVPCACSR
ETGGGFAYTKQGLVNEEFLEACLSMATDPEEHHMFFCQDCPLARVKRQNVHARCDGHLIRKFIKECWVKCGCEMSCGNRVLQRGITCKLQVRIL
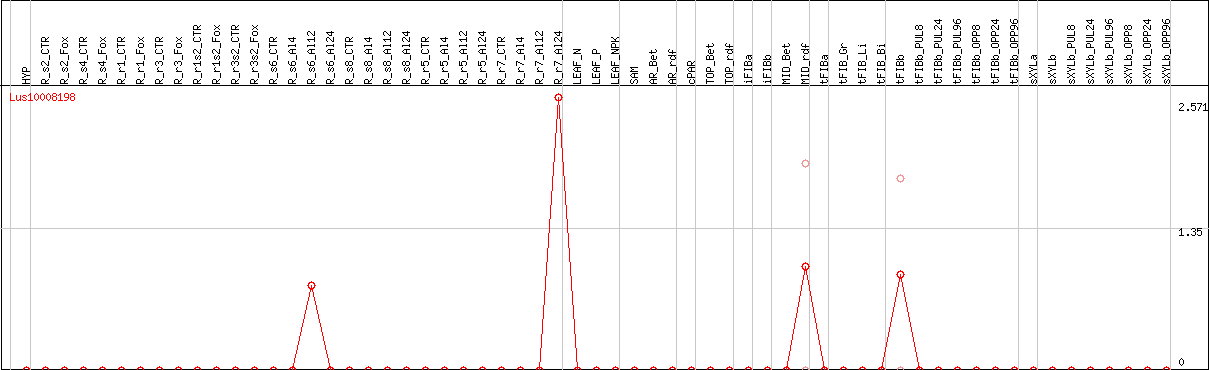
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10008198 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.