External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G02280 129 / 5e-36
ATSUS3, SUS3
sucrose synthase 3 (.1)
AT5G49190 123 / 6e-34
ATSUS2, SSA, SUS2
SUCROSE SYNTHASE FROM ARABIDOPSIS, sucrose synthase 2 (.1)
AT3G43190 115 / 2e-31
ATSUS4, SUS4
ARABIDOPSIS THALIANA SUCROSE SYNTHASE 4, sucrose synthase 4 (.1)
AT5G20830 106 / 4e-28
ASUS1, ATSUS1, SUS1
sucrose synthase 1 (.1.2)
AT1G73370 79 / 3e-18
ATSUS6, SUS6
ARABIDOPSIS THALIANA SUCROSE SYNTHASE 6, sucrose synthase 6 (.1.2)
AT5G37180 62 / 2e-12
ATSUS5, SUS5
ARABIDOPSIS THALIANA SUCROSE SYNTHASE 5, sucrose synthase 5 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013417
144 / 2e-41
AT4G02280 1162 / 0.0
sucrose synthase 3 (.1)
Lus10010308
139 / 2e-39
AT5G49190 1318 / 0.0
SUCROSE SYNTHASE FROM ARABIDOPSIS, sucrose synthase 2 (.1)
Lus10001468
134 / 6e-38
AT4G02280 1385 / 0.0
sucrose synthase 3 (.1)
Lus10020506
115 / 3e-31
AT3G43190 1420 / 0.0
ARABIDOPSIS THALIANA SUCROSE SYNTHASE 4, sucrose synthase 4 (.1)
Lus10012454
115 / 3e-31
AT3G43190 1374 / 0.0
ARABIDOPSIS THALIANA SUCROSE SYNTHASE 4, sucrose synthase 4 (.1)
Lus10041979
79 / 3e-18
AT1G73370 1190 / 0.0
ARABIDOPSIS THALIANA SUCROSE SYNTHASE 6, sucrose synthase 6 (.1.2)
Lus10007372
75 / 6e-17
AT1G73370 1195 / 0.0
ARABIDOPSIS THALIANA SUCROSE SYNTHASE 6, sucrose synthase 6 (.1.2)
Lus10020791
75 / 7e-17
AT1G73370 1200 / 0.0
ARABIDOPSIS THALIANA SUCROSE SYNTHASE 6, sucrose synthase 6 (.1.2)
Lus10017984
74 / 1e-16
AT1G73370 1117 / 0.0
ARABIDOPSIS THALIANA SUCROSE SYNTHASE 6, sucrose synthase 6 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0113
GT-B
PF00862
Sucrose_synth
Sucrose synthase
Representative CDS sequence
>Lus10008204 pacid=23164920 polypeptide=Lus10008204 locus=Lus10008204.g ID=Lus10008204.BGIv1.0 annot-version=v1.0
ATGAATCTTCGAATAATCTTGCTGTTCAGTTTGAAATTTGAAATTTGTGGCGCGCGGTTGACACAGGAAGCCATAGTACTACCTCCATTCATAGCAATTG
CAGTTCGTCCAAGGCCTGGTGTTTGGGAGGATGTGCGCGTTAATGTTAATGAACTCAGCGTGGATCAATTGACTGTCTCCGAGTACCTTAAGTTCAAAGA
AGAGCTTGTGGACGGGCAGGCTAGTGATCCGTATGTTCTGGAGCTGGATATGGAGCCATTCAACGCTAGCTTCCCTCGGCCCAGTCGTTCAACGCCATTC
AACGCCATTCAACGCTAG
AA sequence
>Lus10008204 pacid=23164920 polypeptide=Lus10008204 locus=Lus10008204.g ID=Lus10008204.BGIv1.0 annot-version=v1.0
MNLRIILLFSLKFEICGARLTQEAIVLPPFIAIAVRPRPGVWEDVRVNVNELSVDQLTVSEYLKFKEELVDGQASDPYVLELDMEPFNASFPRPSRSTPF
NAIQR
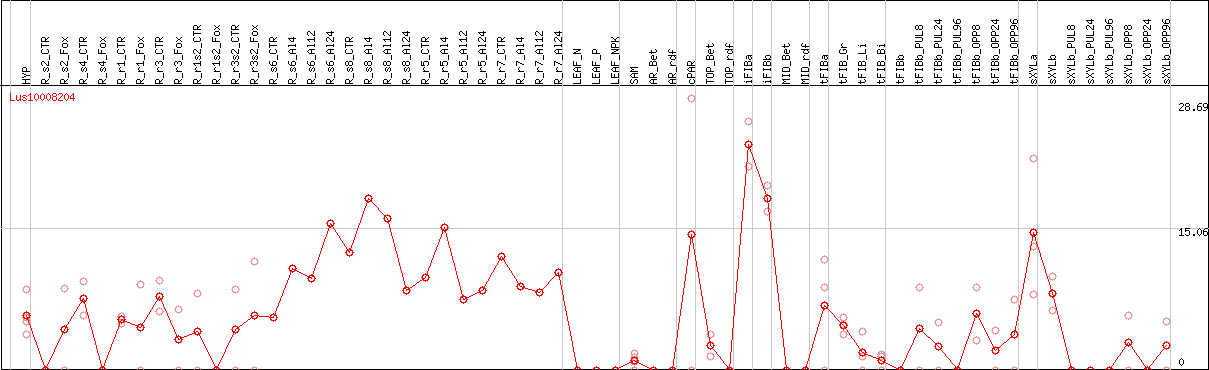
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10008204 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.