External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G19510 85 / 2e-19
Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
AT4G12010 82 / 3e-18
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT4G11170 80 / 1e-17
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT4G16990 76 / 4e-16
RLM3
RESISTANCE TO LEPTOSPHAERIA MACULANS 3, disease resistance protein (TIR-NBS class), putative
AT4G19530 75 / 4e-16
disease resistance protein (TIR-NBS-LRR class) family (.1)
AT2G20142 74 / 5e-16
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT5G45060 75 / 6e-16
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G57670 74 / 1e-15
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT1G72920 72 / 1e-15
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT4G16950 73 / 3e-15
RPP5
RECOGNITION OF PERONOSPORA PARASITICA 5, Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028058
254 / 5e-79
AT4G12010 367 / 3e-108
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10008221
183 / 7e-54
AT4G12010 400 / 8e-120
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10028060
160 / 1e-45
AT4G12010 363 / 1e-106
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10008320
158 / 4e-45
AT4G12010 402 / 1e-120
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10008222
154 / 2e-43
AT4G12010 327 / 4e-96
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10023051
152 / 5e-43
AT4G12010 415 / 2e-125
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10012246
138 / 6e-38
AT4G12010 396 / 9e-122
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10009107
137 / 9e-38
AT4G12010 347 / 4e-101
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10008209
125 / 3e-37
AT4G19510 107 / 8e-28
Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G069866
100 / 1e-26
AT4G12010 179 / 4e-52
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Potri.019G070436
96 / 2e-25
AT2G20142 160 / 7e-49
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
Potri.019G070700
101 / 3e-25
AT5G17680 722 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G069500
100 / 8e-25
AT5G17680 711 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G070393
100 / 8e-25
AT5G17680 629 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G069600
99 / 2e-24
AT5G17680 647 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.019G070651
97 / 1e-23
AT5G17680 630 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Potri.013G098100
89 / 2e-22
AT5G36930 159 / 1e-45
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.013G096849
89 / 3e-22
AT5G36930 167 / 4e-48
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G098600
88 / 6e-22
AT2G20142 138 / 4e-40
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0173
STIR
PF01582
TIR
TIR domain
Representative CDS sequence
>Lus10008318 pacid=23178499 polypeptide=Lus10008318 locus=Lus10008318.g ID=Lus10008318.BGIv1.0 annot-version=v1.0
ATGGCCTCTTCATCTTCCTCTGCTCCTTCACGGCCGTACAGTGGGAAATGGGAATACGACGTTTTCCTATGTTTCAGAGGGGACACGAGGTACGATTTCA
AGAGTCACCTTGAATCAGCTATGAAGGCTAAGAAAATCAGAGTCTTCGTCGACTCAATGCTAAAGAAAACGCAGGACAACGGAGAGCTGCTCGGAATCCT
CGAAAGGACTGCGGTTTCGGTGGTGATTTTTTCGGGCAAGTTCGCTGGTTCGCCCTGGTGCTTGGACGAGGTTTGCACCATCGTTGAGAGCATTGGGAAA
TATGGACACAGCGCACTTCCGGTATTTCACAAAGTGGATTGGATGACTGTTGCTGGCGACTACCATTGCTCCGATTGTTTGAAGTCGGCGATCTGCTGCT
CTTGCTTTCTCGATCGCCAGCCACTGATCAAAGTAACAAGTAAAATTGAAACTGGATCTCATGTGTCGATTTTGTCATCACTAGTAAATCGTGGTAAGAA
ATTAGGTGTGAAGGCACAGATCTGTCTATAG
AA sequence
>Lus10008318 pacid=23178499 polypeptide=Lus10008318 locus=Lus10008318.g ID=Lus10008318.BGIv1.0 annot-version=v1.0
MASSSSSAPSRPYSGKWEYDVFLCFRGDTRYDFKSHLESAMKAKKIRVFVDSMLKKTQDNGELLGILERTAVSVVIFSGKFAGSPWCLDEVCTIVESIGK
YGHSALPVFHKVDWMTVAGDYHCSDCLKSAICCSCFLDRQPLIKVTSKIETGSHVSILSSLVNRGKKLGVKAQICL
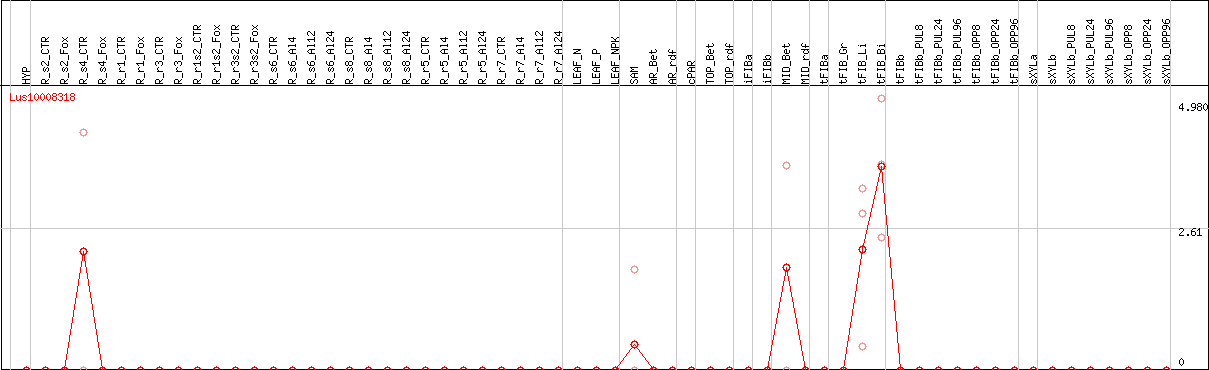
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10008318 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.