Lus10008628 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10008628 pacid=23180071 polypeptide=Lus10008628 locus=Lus10008628.g ID=Lus10008628.BGIv1.0 annot-version=v1.0
ATGTGCGGCTCCGGCGGCGGCGACGGCGACGTCGTCGATGATGGCTCGATCAAATCTCCTATTGACGACCACGAAAACGAGGAAAATCGAGAAGAAGAAG
AAGGCGACGAAGATGAAGTGAACTACGAGGCGGAGTACGAGGTCGAAGGGGAGCTGTTCTCCGGTGTGCGTTTGGCTGATGCTCCTATACTATTCCTCCT
CTGTTTCCATAGGGCGTTCCGTCGGGAACTGGCGGAGCTGTGTCGGATTGCTCTTGTGGCTTCCGAGAGTGGAGGAGGAGGAAGCGCGATTGATGAGCTT
CGCTGCAGATTCGAGTTCCTTAAACTTGCTTATGACTATCACTCTGCCGCTGAAGACGAGGTTATCTTTCTAGCGTTAGACGTCCACGTTAAGAATGTTG
TCTGTGCATATTCACTAGAGCACAAAAGCATAGATGATGCATTTGCCACGGTTTCCCGTTGCTTGAAAGTGTTGATGGAGGAAAAACAAGAAACTAGTTT
TGAGCCATTTCAAGAACTTGTATCTTGCCTCAGCATTATCCATTCCTCCATATGCCACCATATGCTAAAAGAAGAAAAACAGGTTTTTCCTTTACTCGTG
GAGCGGTTCTCTCCGAAAGAACAAGCTTCACTTGTGTGGCAATTCCTTTGCAGCATACCAGTAATTTTGTTGGAGGATTTCCTGCCATGGATGATCTCTC
TTTCTCCTGAAAACCAAGATGATGTTGCACTGTGCATTAAAGAAATTGTACCTGATGGACCACTCCAAGAGGTAATAATTTCTTGGCTCCGGAGACCCGC
TCAATTCCCATCTAGGGATTTGACGAACACAGCTGAACGGGGACATATGCCACACCTGCTCTTTTCTGCTAGGTGTTCTGAAGAAAATTGGCAATCGAGG
ACAACAGATTCTGTCCTTGCCAAAAAGGGGCACATTGTTCATGACTGTTTCCATCTCTGGCATGGTGTGATTATAAAATCATTATGGGAAATTCTTGAAG
AATCATACCAAATAAGATCAGGCAGTTTGTTAAATGTAGATCCAACCTTATTGAGGCTGAAGTTTCTTGCTGATGTATTTACCTTCTACAGCAATGCATT
GAAGAAATTCTTTGATCAGATGTTAAATAGACTGGCCAGTGACCACTTCACTGGCTTATATGCTGACCAGTCCCAGGCTGAAAGTCACCTGGAGAGATTA
CAGCAGTTGCTCCATGATAATGCTCAGAATATTTTACCTGTGCATGAGTTTGTGGACAAGCTTTGCGAGGGAATGGAGCCCTTTGTCGTGGAAGTCGGAA
AACAATTCAAATTTCTAGAAGCAAAGGTACTTTCCCTTTTAATCGATAACTGTGGCCAAGAAATGCAGTTAGAGCTCTTTTTGTACATGAGCTTTGGTAT
GATGCCTCTAGGATTACTGAAGTTTGTAATCACTTGGTTCGCAGCCTGCTTGTCCAAACATGAGTTGGGACTCATCCTTCAAGGGTTGAAACAAGATGAA
AGTTTGGCCAGTAAATCTTTTCTGTCCCTCTTGCATGAGTGGTTGCTCATTGGTTATTCTGGAAAAATCTCTGTCGATGATTTTGGAAAGGATTTGCAGA
GAATGACTAAATCTTCATTTCTGATTGAAAGATCCAAGGAGGATGTTAAATTCTCTTCCTTACCCTTGAACTTGAAACCTTATGAAGCATCTCCCCATTT
TAAAGCAGAACCAGTTCCCCTTTATGAGAACAAATTCCTTTCTCCATCTTCTTTGTCCTTTGGTTTTGACAAAGCTAAGAGTTACCAGACATCTTATTCT
AGAGAAATAAACCTACATGTGTGTTGTCCTCAAACGATAAAGCTAAGGAACTCTGTTCCTAAAATACCTGGTGTGGAGAGCTCTTCTGCTTCCACTATGG
ATGAGCCAATACCTTTGGATTTCATGCTTTTCTTCAATAAGGCTCTTAAGAAGGATTTGGAGAACATTGTCCTGTATTCAGCTCGATTGGCTGAGAACTT
TGGGTTTCTCAAAGAATTCTGCCAACACTTCAATCTATTGCGGCACCAATATAAGTTCCACAGTAATGCCGAGGATGAATGGATATTTCCGTTACTTGAG
GCTAAGGGAAAAGCTGAGAATATTAGTCACTCCTACACAATTGATCATCAGCTTGAAGTCGAGAATTTCCAACAAATTTCCATCATTCTTGAAGAGATGG
TTGGCTTGCAGATGTCAGTCTCTATTGGCACTCATAAGCAGGCTCGGAGAATTATTAGGTGCAAGCATCTCTGTCTGAAACTCCAGCGCAGGTGCAAATC
TATTCAAAAACTACTCTCTGATCATATTCATCATGAAGAAATTGAACTTTGGCCAGTAATCAAGCAATGCTTGACAATCCAAGAGCAGGAAAAGGTTGTG
GCAAGCATATTGGGGAGGACTAGAGCAGAAACATTACAAGTCATGATACCATGGTTGATTGGCTCTCTAAAACCAGAAGAACAGCTTACAGTTATGTCCT
TATGGCGTAAGGTCACAAAGGACACAAACTTCAGTGATTGGTTAGGAGAATGGTGGGAAGTGTGTGATAAATCTTCTATCTCTGAGGAGTTTTCTTCATG
TGATACAGACTCGCTGGAGACTGTCTCCAGATATTTGTCTAGTGAATCCCCTGATCAACAAGATGGCTTCCTCAACAACATAAGCATCAATATGCCACAG
AAAGATTGCAATAGTGCTAACATCGTGGGAGATAGTGCATCCAAAGATAATACGAAAACTGCTGCTGTGGATGAGGAAAATGAGTGTCTAGACTGCATAA
AGATTCTCAATCAAGGTGATAAAAAGACATGCAATGAAGTAGGGGAAATCACAGATGAGAAGGCAAATAAAGCTCAACCTTTCCAAGAAACTCTAAAATC
CGTTGATCATGAGCGCCTTTTATCAATTGGTCAAGAAGACTTCGAAGCAGCATTAGGAAGAATTTCTCACGACTCCTCTCTAGATCTTCAGAAGAAATCA
CGTCTAATGCAGAACCTGCACACGAGTCGTTGGCTTATTCAGAAACAGATGTATCATACTGATGAAATTCCCACGACTAATGAGAAAGTACTTCAGGGTC
AGCATCCATCATACCAAGACCCTAAAAAGTCAACCCTGGGTTGTAAACACTACAAGAGGAGATGCAAACTTTTTGCTGAATGTTGCAATCAACTTTATAC
ATGCCTACGTTGCCATGATGAGTCAGCTGATCACACTGCAGACAGAAGGGATATTACAAAGATGATGTGCATGAATTGTTTAACTATCCAATCTATACAG
CAGACATGTGCAACCATCTCCTGCAACAATTTGTCGATGGCAAAATACTATTGCCAAATATGCAAATTGTTTGACGATGGAAGGGAAATCTACCACTGCC
CTTTCTGTAACATCTGCCGTGTTGGAAAGGGTTTAGGGATTGACTACTTCCATTGCATGAAGTGCAATGCCTGCATGTCCAAGTCTTTGTCCAAGCATAC
ATGCAGAGAGAAATGCTTCGACGATAACTGCCCCATTTGCCGGGAATCCATCTTCACTTCCAGCATACCAGTGAAAGCTCTCCCTTGTGGTCATCTGATG
CACTCCCCATGTTTTCAGGACTACACTTGCAATCATTATATCTGCCCGATCTGTAGCAAGTCACTTGGCGACATGAAGGTGTACTTTAAGATGCTGGATG
CCCTTTTGGCTGAAGAGAGAACTCCAGATGAATACGCAAGCCAAACTCAGGCCATCCTTTGCAATGACTGTGAGGAGAAAGGATCTGCACGTTTCCACTG
GTTTTATCACAAATGTTCGTATTGTGGGTCCTACAACACAAGACGTATATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10008628 pacid=23180071 polypeptide=Lus10008628 locus=Lus10008628.g ID=Lus10008628.BGIv1.0 annot-version=v1.0
MCGSGGGDGDVVDDGSIKSPIDDHENEENREEEEGDEDEVNYEAEYEVEGELFSGVRLADAPILFLLCFHRAFRRELAELCRIALVASESGGGGSAIDEL
RCRFEFLKLAYDYHSAAEDEVIFLALDVHVKNVVCAYSLEHKSIDDAFATVSRCLKVLMEEKQETSFEPFQELVSCLSIIHSSICHHMLKEEKQVFPLLV
ERFSPKEQASLVWQFLCSIPVILLEDFLPWMISLSPENQDDVALCIKEIVPDGPLQEVIISWLRRPAQFPSRDLTNTAERGHMPHLLFSARCSEENWQSR
TTDSVLAKKGHIVHDCFHLWHGVIIKSLWEILEESYQIRSGSLLNVDPTLLRLKFLADVFTFYSNALKKFFDQMLNRLASDHFTGLYADQSQAESHLERL
QQLLHDNAQNILPVHEFVDKLCEGMEPFVVEVGKQFKFLEAKVLSLLIDNCGQEMQLELFLYMSFGMMPLGLLKFVITWFAACLSKHELGLILQGLKQDE
SLASKSFLSLLHEWLLIGYSGKISVDDFGKDLQRMTKSSFLIERSKEDVKFSSLPLNLKPYEASPHFKAEPVPLYENKFLSPSSLSFGFDKAKSYQTSYS
REINLHVCCPQTIKLRNSVPKIPGVESSSASTMDEPIPLDFMLFFNKALKKDLENIVLYSARLAENFGFLKEFCQHFNLLRHQYKFHSNAEDEWIFPLLE
AKGKAENISHSYTIDHQLEVENFQQISIILEEMVGLQMSVSIGTHKQARRIIRCKHLCLKLQRRCKSIQKLLSDHIHHEEIELWPVIKQCLTIQEQEKVV
ASILGRTRAETLQVMIPWLIGSLKPEEQLTVMSLWRKVTKDTNFSDWLGEWWEVCDKSSISEEFSSCDTDSLETVSRYLSSESPDQQDGFLNNISINMPQ
KDCNSANIVGDSASKDNTKTAAVDEENECLDCIKILNQGDKKTCNEVGEITDEKANKAQPFQETLKSVDHERLLSIGQEDFEAALGRISHDSSLDLQKKS
RLMQNLHTSRWLIQKQMYHTDEIPTTNEKVLQGQHPSYQDPKKSTLGCKHYKRRCKLFAECCNQLYTCLRCHDESADHTADRRDITKMMCMNCLTIQSIQ
QTCATISCNNLSMAKYYCQICKLFDDGREIYHCPFCNICRVGKGLGIDYFHCMKCNACMSKSLSKHTCREKCFDDNCPICRESIFTSSIPVKALPCGHLM
HSPCFQDYTCNHYICPICSKSLGDMKVYFKMLDALLAEERTPDEYASQTQAILCNDCEEKGSARFHWFYHKCSYCGSYNTRRI
|
DESeq2's median of ratios [FLAX]
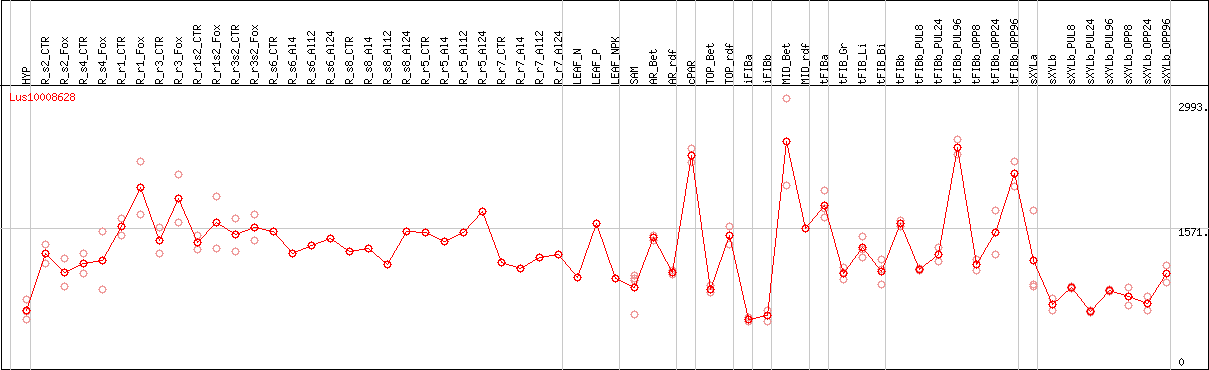
Coexpressed genes
Lus10008628 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
