Lus10008959 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10008959 pacid=23175170 polypeptide=Lus10008959 locus=Lus10008959.g ID=Lus10008959.BGIv1.0 annot-version=v1.0
ATGGGAAAAATAGCCAGCCCTACGAGGATGGAANNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNTCGACACCCTGCCCCCCAGACAGCAAACGGCAGCCGTAGCGGTTGGTTCTCGAAACTGGTCGA
TCCGGCGCAGAAGCTCATCTCCTCTGGTGCTCAAAGATTTTTTGCATCTGTCTTCCGCAAGCGCCTTGCCCCTCCTCCACCGACGCGGCCGCAGGAAGCA
GAAGCAGAGCAGGGAACAAAGGACAAAGAAGTCAAAACGACTATCAAGGACAAGCAAGTCATAACAATTCCCAAGGATAAACAAGTTGAAACATTTCCCA
TGCTTTCCTCCAAGGTTCACGGAGACTCTCGGGGAGAAACTAGCCACGTGTATGATGGCCCAAGTAGCTCAAATGGAGGCGGGGCTACTGATTTGGAGTC
GATGTTGAAGCATAAAAGTTTTACCAGGCATGAGATTGATAGATTGACTGCTCTATTGCAGTCAAGAATGGTTGATTCACCTCCAGAAACTGGAAAGAAT
GCTGAAGTTGCTCCATCAAATTTATTTCTGCTGGGGGACAAGGATGAACTGGCCATCACTCCGGTTAAAAATCGTGGGACTATGAACAACTTATTCTCAA
CCCCTGTTGTACTTGATCAAGATGTTACTTCACCAGCGGAGCTTGCAAAAGCTTACATGGGCAGTAGACCTTCCAAGGTATCTCCATCAATGTTAGGTTT
CCCCAGTCAACCATGGAAGGAAGATTCAGCTCCGCAGTTAAGTCCAAACTTTGTTTCGAAAGGGCCGACTCCATCCGCCATTTACAGGTCTTCCAGTCAA
GCTGGATTGACAGATAATGGATTCATGACACCAAATTCACGTGGTAGATCTGCTGTTTACAGCATGGCTAGAACTCCATATTCCAGAACCCAGTCAACTA
AAATGATTCAGGCCATAGGTACCTCAAGTAATGCTTGGGGTGGGCAAGATTCTTCTTCTCTTAGTGCTGTGGAGAATGGCAGAGTTTATGGATCGAAACA
AGGGACTTTGAAGAGAAGGAGTTCTGTACTGGACAACGATATAGGATCTACTGGTCCTATTCGCAGGATTCGCCAAAAATCAAATCTAATGCCTACTGGA
GCTCTCTCCGCCCGTGGAACTCGTTTCACTTCGAATGCAGAGAGACAGCTATCCTTGACCTCAAAGCCTGCTCTAGACAATAAACCTTCCAAGGAAAATA
CCGACTATAACATTCATAATTCAGGATTTGCCCCCATCTCTTCTCAATCCAGTGAGATGGCTTCAAAGATACTCAAGCAGCTAGATGTGTTGGTCTCATC
TAGGGAAAAGTCCCCTAGTAAATTATCCCCATCGAAGTTACGTGGACAAGCTCTTAGAAGCCTAGAGAAAATAGATTCTTCGAAATTTTTGGAGAGTTTT
CAGAATCCGGGGATGGATGCTCCAAATGCTGGTATGTTAGATGATTCTCAACACTGTATGCCTCGAAATGAGGACATGAGTGAACAAATTGGTCCGGGCA
AAGTAAACTATCTTAGTAGCAAGTTGGGTACTGGAAAAACTGTAGAGACACCTACTTCAGCTGGTGATGGAGCAGCTAAAGGAGTTGATTCTACCATCAA
GCCCCCTCAAGCGAGAAAACCAGGATTTCGTATAAGTGCAGATGAGGATTTCCGGGAGATGGATGACGATGACGGTGTCAGGAAAAATGCTACCGGTGAC
ATGGACAATATGAAGTTCATCTCTAGTTCTTCTAATGGTAAAGCTGCTGAAACTGTTACGACGGGAAACCCTTCAGTTCTTTATGAAGCGACTCCCCAGA
CAAATTCTACACTAATCAAGAATGATGATTCGACAATCCTTGGAAATAGCAATACTTTTGCAGTTGGTGCAGCTTCATCCATGGGGATTGTGAAGCATGC
TGCCTCCTTAAGTCAATCAGCTGCAGCTTCTGATAAGACTGACGGCAGAGTATCCAGGGAGAAGGTTGCTTTTCAAATTGGGCTTAATGTTGCCCATGAG
AGAACTCCTGTGTTTTCGTCTTCATCTCCGAAAGCCGCAAATGATGGATTTGGTGCTTCATCAAACTCCCAATCAGAAAGCTTAGAGAGGACTGTTACTG
CTGCTTCCACAATAGTTGATCCTGTGGCAAAACAACCTGAAGCTGATAAAGATGTTGAGAGAAACAAATCTAGCCTCTCCTTCCTGAATAATAAGGAAAC
TGTACCTGCATCTTTGTTGGCCTCATCACCTGCTGGAGGCATATTCTCATTTGGGTCCAGTCTACAAAATGGAACTCTTCCAGGGTCAGCAACTGCTCAA
ACTGTATCCAACAGTTCCGTCTCGGTTGTTCCTAGTGGCACCATTGGCGCAGACATCTCTGCGAATGCGGCAAAAACTGATAGTAAAGGCGATTTTAAAT
CAGCTTTAGCACCTCCCGCGAGTTCATCCATTTTCAAATTTGGGTCATCTCCTACTTCAGGTTCAGTTTCTTTGGCTTCAACAAATGCTGGTAGTGTTAT
CAGCCAAAAAAAAGCAGACACTAGCTTTGGAAGTTTTATCAGTGCTGCCTCCGGTGATTCCAAACCTGCAGTCACAATTGCTGAGAACAGTACAGCCAGC
CCCTCAGGTGATATTTCATCTACGGAAAAGGGAAGTGCCTTAATTGGTGGTACTGGTTTTTCAAGTACAAGCGCTGGAAACAGTCTTTTTGGTGGTAAAC
CTACTTCTGGTACAATCACCAGCAGTAACCCTTTTGGTGCTTCTAGCTCTGCTGGTACGAGCACTGGAAGTGGTCTCTTTAGTACTACTGGTGGTTCAAG
CAGTGGACACAGCCTCTTCAGCGCAACAGCTCCAAGTACGACTATCGGGAGTAGTCCTTTTAGTTTCAGTGCTAAATCAGTACCGTCAGGTGTGAGCAAT
CAGTCTCAGAATATGAATAATACTCTAGGCAGCACCCACGTAGCTGCAACTGGCTCTGCCTTGTCAAACCCGAGTACCACTATGTTCAGTTCTACTCTGC
CGTCTCCTTTTCAGTTGTCAAACTCGTCTGCTGCTGCTCCCGGCAATTCACTTTTCAGCTCATCTTCTGCTTCTAATTTGTTCAGTTCTGGTCCTTCTTT
TGGAGCAACACCTTCCGGCCCAGCTTCAGAACCAAACTCGGTTAGCTCAAGTGCTAGCACATCATCTCCAATATTTAGTACAAGCTGGTCAACACCTACA
CCATCCGTGTTTAGTACTATGCCCAACTCCACAGCTTCGTCAAGTGGGTTTGTCTTCGGAGCGTCATCAAGTGCAGTTACTGCAGGCACCTCCGTCTTTG
GATCTTCTACATCCAGTTCCCCATTCTCCTTCAGTGCACCTGCAGCAAATCCTCCTGCTCAATCTGTGTTTGGTGGCCCTAGTACATCTTTTGCGTTCAA
TTCGACTCCATCTTTTGGCTCGACACCGTCAGGTAGCAATGATCAAATGAGCATGGAGGACAGCATGGCAGAAGACACAGTTCAGGCAACCACACATTCT
GTTCCCTTCTTCAGTCAGCCAACATCTGCACCCCCTTCCTCTGGCTTTGTATTCGGTTCGGAATGCAGCTCCATAACTCCGGCAAATGCTCAGTTTGGTG
CAGCATCACCATTCCAATTTGGCGGCCAGCAAAGTATGCCGGCAGCTCATCAACAAAATGCTTCACCATTTCAAGCCAGCGGGAGCCTGGAGTTCAACGC
AGGAGGCAGCTTTTCATTAGGGACAGGTGGTGATAAAAGTAACAGAAGAATCATTAAAGTGAAGAAAAGCAACCGGAAGAGGTGA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10008959 pacid=23175170 polypeptide=Lus10008959 locus=Lus10008959.g ID=Lus10008959.BGIv1.0 annot-version=v1.0
MGKIASPTRMEXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXRHPAPQTANGSRSGWFSKLVDPAQKLISSGAQRFFASVFRKRLAPPPPTRPQEA
EAEQGTKDKEVKTTIKDKQVITIPKDKQVETFPMLSSKVHGDSRGETSHVYDGPSSSNGGGATDLESMLKHKSFTRHEIDRLTALLQSRMVDSPPETGKN
AEVAPSNLFLLGDKDELAITPVKNRGTMNNLFSTPVVLDQDVTSPAELAKAYMGSRPSKVSPSMLGFPSQPWKEDSAPQLSPNFVSKGPTPSAIYRSSSQ
AGLTDNGFMTPNSRGRSAVYSMARTPYSRTQSTKMIQAIGTSSNAWGGQDSSSLSAVENGRVYGSKQGTLKRRSSVLDNDIGSTGPIRRIRQKSNLMPTG
ALSARGTRFTSNAERQLSLTSKPALDNKPSKENTDYNIHNSGFAPISSQSSEMASKILKQLDVLVSSREKSPSKLSPSKLRGQALRSLEKIDSSKFLESF
QNPGMDAPNAGMLDDSQHCMPRNEDMSEQIGPGKVNYLSSKLGTGKTVETPTSAGDGAAKGVDSTIKPPQARKPGFRISADEDFREMDDDDGVRKNATGD
MDNMKFISSSSNGKAAETVTTGNPSVLYEATPQTNSTLIKNDDSTILGNSNTFAVGAASSMGIVKHAASLSQSAAASDKTDGRVSREKVAFQIGLNVAHE
RTPVFSSSSPKAANDGFGASSNSQSESLERTVTAASTIVDPVAKQPEADKDVERNKSSLSFLNNKETVPASLLASSPAGGIFSFGSSLQNGTLPGSATAQ
TVSNSSVSVVPSGTIGADISANAAKTDSKGDFKSALAPPASSSIFKFGSSPTSGSVSLASTNAGSVISQKKADTSFGSFISAASGDSKPAVTIAENSTAS
PSGDISSTEKGSALIGGTGFSSTSAGNSLFGGKPTSGTITSSNPFGASSSAGTSTGSGLFSTTGGSSSGHSLFSATAPSTTIGSSPFSFSAKSVPSGVSN
QSQNMNNTLGSTHVAATGSALSNPSTTMFSSTLPSPFQLSNSSAAAPGNSLFSSSSASNLFSSGPSFGATPSGPASEPNSVSSSASTSSPIFSTSWSTPT
PSVFSTMPNSTASSSGFVFGASSSAVTAGTSVFGSSTSSSPFSFSAPAANPPAQSVFGGPSTSFAFNSTPSFGSTPSGSNDQMSMEDSMAEDTVQATTHS
VPFFSQPTSAPPSSGFVFGSECSSITPANAQFGAASPFQFGGQQSMPAAHQQNASPFQASGSLEFNAGGSFSLGTGGDKSNRRIIKVKKSNRKR
|
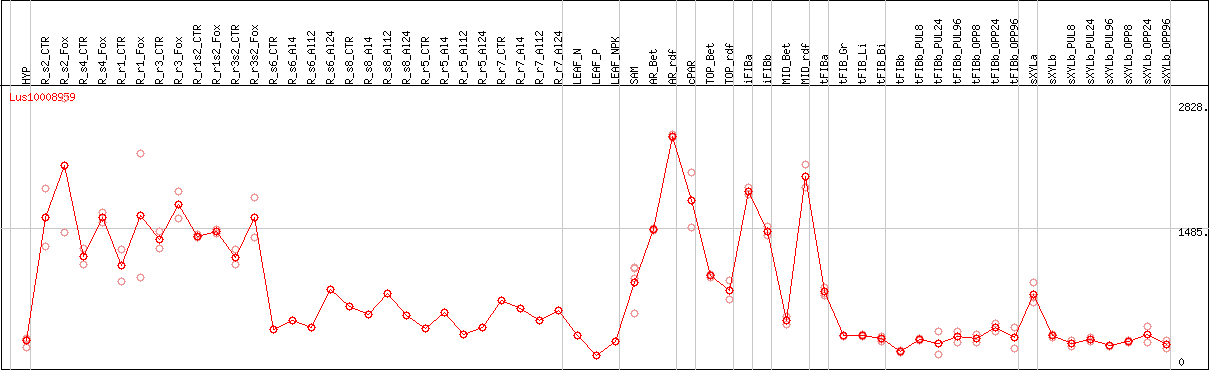
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10008959 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
