Lus10009028 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10009028 pacid=23173348 polypeptide=Lus10009028 locus=Lus10009028.g ID=Lus10009028.BGIv1.0 annot-version=v1.0
ATGAGCACATCAAAGTCAAAGTCAGAGTTTTTAGCGAGACAATATGGGAGAGTCAACAGTGCAAAACATCTTTCAACAGACTGGGATTGCTCCCAATCAG
CTTCCTCCTTATGTGATTCTGATGAGACTGAAGAGAGGAAGCTAGCAGTGTTGATAGAGTGGCTAAACACCATTCTACCTGATCTGAAGCTGCCACTAAA
GGCTTCAATTGATGAACTCAGAGACTGCTTAGCTGATGGCACTGTCCTTCTTCAAATCTTGAGGAAAGTCAAACCAAGATCCTTCACTAAAGATCTAGAA
GGGGTTTTGGAAGCTGAGATAGTATCACCAAGCCAAAATGTACACCTATTCTTGGACAACATTGATGAAAGGGGCATCCCCAAGTTCGAGATGGGAGACT
TGGAGAAGGGCTCGATGAAGAGAGTGGTGGATTGCCTTCTGTCAATAAGAGTAAGGTACCCAACAGGGAGGAACAACTTCTCATCAGTAAATGGTAGAAT
GACTAGAAGTGGTAGCCTAAGGGAGTTCTCACCTTACCAAAGCCTTCACCTTGGTACATTCTCATCCTCACTCTCGTCTCCTGGTGCCGAAAGGCAGCCA
CTCTACTCCGAGTCGAAATTCCAGCAGTTCTTGCGTACCAGTCCCGCCCTTTCAGAGTCATCTGCCGCGTGGATGCACCACGCTGGACACCGATTCCACG
AGGTGTTCCAGGTCAGGCAAGGTCGATACGCTGACCTGTCGCCTTCGAAGATATCGGAGATGATGAAGTCAACCAGCTTAGATAATGCTCCAACTCAGTC
TCTTCTAAGTGTTGTCAATGGCATCTTGGATGAAAGCATTGACAGAAAGAACGGCGAAATCCCTCATAGGGTAGCTTGCTTGCTGAGGAAGGTAGTGCAA
GAGATTGAAAGGAGGATATCAACTCAAGCAGAACACTTAAGAACACAAAACAACCTTTTCAAGGCTAGAGAGGAGAAATACCAGTCCAGAATTAGGGTTC
TTGAGACACTTGCTTCCGGTACAGGTGAAGAACCTGCGGGAGTGAAGGTGAAGATGGAAGATAAAAGCAAGGTGAATGAAGAGATATCAAAGTTGATGAA
AGAAAAGGAACAAGAAAGCATTGAGCTTGCAAGGGTAAGGCAAGAGCTGGAAACAACCAAGACCGTATATGATGATATGGTGAAGCTAATGAGACAAAAG
GATGAGACAACAACTAGTTTCAAACTCTCATCTTCAAACCAGCAGCAGAGAACGAATGCCGAAATCTCAGCGTTGCAGCAGGAAGTTGCAGCAGGGAAGA
AAGAGCTCAACCACATAGTTGACTCGGTTAGGGAGACAGACCGGACGAAGAATGAGCAGACCAATGCTGAGGTTCTAGCATTGCGGAAGGAACTGGCTTC
GACGAGAAAGGAGCTTGATGGAATTTTCGGTTCAATGAAGGAAAAGGACCAAATGAATAAAGTTGGCGGTGATAATGTGGTTAGATTATTGAAGGAGAAA
GAGGAGATGAGTCTTCAAGTTGTGGCATTGAAGCTAGAGCTCGAAAGGGTTTTGAGGCAAGGAAATGTCGGTAATGATGTCGATGTGGTTAGATTAGTGA
AGGAGAAAGATGATTTGAGTGTTCAAGTTATGGCATTGAAGCAGGAGCTTGAAAGGGTTTCGAGACAAAGCATTAATGTCGGTGCTGCTGATGTTGTTAG
ATTAGTGAAGGAGAAAGAGGAGATGAGTCTCCAAATCGCGACATTGAAGCAAGAACACGAAAGAAAAAAGCAATTAGGACATGGAAATGTGGAGAATGGT
GATGTGGTTAGATTAGTGAAGGAGAAAGAGGACATGAATGGTCAAATCGCGACATTGAAGCAAGAGCTCGAAAGGGTAAAGGGATCACAGGGGGATGTTG
ATAATGATGTCGACGTGGCTAGATTAGTGAAGGAGAAGGAGGAAATGAGCCTAAGACTGTCCAAATTGCAGCAAGAAATGGAGACATTAACACAAGGAAA
TAACAATGATGCTGATGTGGCTAGATTGATGAAGGAAAATGAAGAAATGAGCCTAAAGATCTCATCTTTGCAGCAAGAAGGAGACTTGGTTAGCAAGCAA
CTCACAACTCTGCAGCAAGAACATGAGGAAATGGTCAGATCAATGAAGGAGAAAGACCAAACTAGCCATGATCAGCTCTCAAAATTGCAGCAAGAGATGG
ATTCGGCAAAAAGGGAGCTCGACGGCATGGCTAGATTGGTGAAGGAGAAAGACCAGGCAAACCTTAGACTTGCCATGTCAATCCAAGAGCTAGAGGTGGC
TAACAAGGCAAAAGAACAGAACTACTTGCAACTTGAGGGTCAACTCAAAGAGGTTCAGAGTCAACTGATTAGGTCAAAGGAAAAAATGGAGTCAGATCAA
CAGATTTGGAGTAAGAAACAGAAGAGTGTTCTCAAGTCTGTTGATTTGCAGTTCCATACCCTTCATGAGCTAAGGTTTTCATCCACCAGCATCAAAGAAG
AGGTTCTACAAGTGCAGCAAGACTATACAGAGAAGTTCCTCAGCCTAAGAGTGAAGTTCAAGACATTGGTGGATGCAAGTGAGAACTACTACTTGGTTCT
TGCCGAGAACAGGAGGCTATTCAATGAGCTACAAGACTTAAGAGGTAACATCAGAGTGTACTGCAGAGTGAGGCCATTCTTGCCAGGGCACCAGAGCGGT
TCAAATAAGTCAGTTGTGGAGCACATTGGTGACAAAGGGGAGCTGGTAGTTGGGAACCCTGCCAAAGCTGGGAAAGATGCTCGAAAGTTGTTCCGGTTCA
ACAAGGTGTATGGTCCTGGAGCTACTCAAGGTGAGGTTTACTTGGACACACAGCCATTGGTGCAGTCTGTGCTTGATGGGTACAATGTCTGCATATTTGC
TTATGGGCAGACTGGTTCAGGCAAAACTTATACTATGTTGGGTCCTAATGGAGCAAAAGAAGAGGAGTGGGGAGTCAACTACAGAGCTCTAAAGGACCTC
TTCAACATGTCAAAAGGCAGAAGTGACTCCATATCGTATGAAGTCGGTGTTCAGATGGTCGAGATCTACAACGAACAAGGCGGTGAGCTCCACCGCCATG
AACGAGAGGAGCAGTCGGGCCCACCACGGCCTGAGCGTCCACGCGGACAAGCAAAAACCTTAATGTTCGTACAGCTGAACCCAGACGCAAGCTCCTACTC
GGAAAGCCTAAGCACGCTGAAATTCGCCGAGAGAGTCTCGGGAATCGAGCTCGGAGCAGCGAAGAGCAACCCGGGGAGGGACGTCAAGGAGCTAATGGAG
CAGCTTGCATCGCTCAAGGACACTATAGCCAACAAGGACGAGGAGATCGAGAGGCTACAAGTGCTCAAGGTCGCCGGCGCCGGCGCACACCCTCCGCCGG
GGAGGCTGCGGACGAGGCAGTCGTTGGATAAGGACTCGGAGATGGGTGGTGGCGATTCCGAGAGTAGGTATAGTGATAATTATATGTCGGACGGGGAAGC
TTCGGTCGGGACGGAGCCAGAGTCCGTGCATGACGACGGCGGAGGAGGGGGGAGAGCTTCGTCCGCCGATAAGCGTACAGGGGCGAAAGGGTTGACTAAG
ACGAGGAGCTTGCAGAGGCTGGGGGGGGGGAAGGCGGCGGGGGGGGGGGGGAGGCTAGGCCGTCGACGGTGTCGAGAGACCATCTCAAAGGATCAACAAG
TGTTAAGAAAGCTACCGCCGGCAGCAGCACCTCCACCTTACAGAAGCCGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10009028 pacid=23173348 polypeptide=Lus10009028 locus=Lus10009028.g ID=Lus10009028.BGIv1.0 annot-version=v1.0
MSTSKSKSEFLARQYGRVNSAKHLSTDWDCSQSASSLCDSDETEERKLAVLIEWLNTILPDLKLPLKASIDELRDCLADGTVLLQILRKVKPRSFTKDLE
GVLEAEIVSPSQNVHLFLDNIDERGIPKFEMGDLEKGSMKRVVDCLLSIRVRYPTGRNNFSSVNGRMTRSGSLREFSPYQSLHLGTFSSSLSSPGAERQP
LYSESKFQQFLRTSPALSESSAAWMHHAGHRFHEVFQVRQGRYADLSPSKISEMMKSTSLDNAPTQSLLSVVNGILDESIDRKNGEIPHRVACLLRKVVQ
EIERRISTQAEHLRTQNNLFKAREEKYQSRIRVLETLASGTGEEPAGVKVKMEDKSKVNEEISKLMKEKEQESIELARVRQELETTKTVYDDMVKLMRQK
DETTTSFKLSSSNQQQRTNAEISALQQEVAAGKKELNHIVDSVRETDRTKNEQTNAEVLALRKELASTRKELDGIFGSMKEKDQMNKVGGDNVVRLLKEK
EEMSLQVVALKLELERVLRQGNVGNDVDVVRLVKEKDDLSVQVMALKQELERVSRQSINVGAADVVRLVKEKEEMSLQIATLKQEHERKKQLGHGNVENG
DVVRLVKEKEDMNGQIATLKQELERVKGSQGDVDNDVDVARLVKEKEEMSLRLSKLQQEMETLTQGNNNDADVARLMKENEEMSLKISSLQQEGDLVSKQ
LTTLQQEHEEMVRSMKEKDQTSHDQLSKLQQEMDSAKRELDGMARLVKEKDQANLRLAMSIQELEVANKAKEQNYLQLEGQLKEVQSQLIRSKEKMESDQ
QIWSKKQKSVLKSVDLQFHTLHELRFSSTSIKEEVLQVQQDYTEKFLSLRVKFKTLVDASENYYLVLAENRRLFNELQDLRGNIRVYCRVRPFLPGHQSG
SNKSVVEHIGDKGELVVGNPAKAGKDARKLFRFNKVYGPGATQGEVYLDTQPLVQSVLDGYNVCIFAYGQTGSGKTYTMLGPNGAKEEEWGVNYRALKDL
FNMSKGRSDSISYEVGVQMVEIYNEQGGELHRHEREEQSGPPRPERPRGQAKTLMFVQLNPDASSYSESLSTLKFAERVSGIELGAAKSNPGRDVKELME
QLASLKDTIANKDEEIERLQVLKVAGAGAHPPPGRLRTRQSLDKDSEMGGGDSESRYSDNYMSDGEASVGTEPESVHDDGGGGGRASSADKRTGAKGLTK
TRSLQRLGGGKAAGGGGRLGRRRCRETISKDQQVLRKLPPAAAPPPYRSR
|
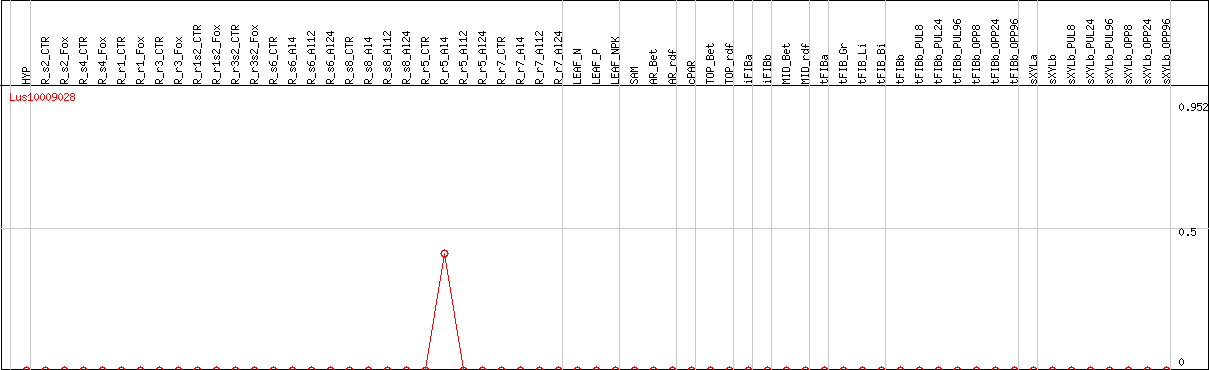
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009028 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
