External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G17980 254 / 2e-86
AtC2
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G48590 243 / 6e-82
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2), Calcium-dependent lipid-binding (CaLB domain) family protein (.3)
AT5G37740 240 / 6e-81
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT1G73580 229 / 1e-76
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G70810 226 / 3e-75
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G66360 223 / 6e-74
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G70800 204 / 1e-66
EHB1
ENHANCED BENDING 1, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT1G70790 204 / 2e-66
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
AT1G23140 202 / 6e-66
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
AT2G01540 199 / 2e-64
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009684
428 / 8e-154
AT3G17980 254 / 1e-86
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10042002
260 / 2e-88
AT1G48590 240 / 6e-82
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2), Calcium-dependent lipid-binding (CaLB domain) family protein (.3)
Lus10018005
245 / 1e-82
AT3G17980 222 / 1e-74
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10033585
232 / 1e-77
AT3G17980 244 / 1e-83
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10017629
228 / 4e-76
AT3G17980 241 / 2e-82
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10021775
194 / 1e-62
AT1G70810 209 / 6e-70
Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10001594
181 / 8e-57
AT3G17980 187 / 7e-61
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Lus10034591
163 / 9e-51
AT1G70790 181 / 3e-59
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Lus10003710
155 / 1e-47
AT1G48590 158 / 2e-50
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2), Calcium-dependent lipid-binding (CaLB domain) family protein (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G047100
260 / 1e-88
AT3G17980 251 / 2e-86
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.017G125500
244 / 1e-82
AT3G17980 249 / 6e-86
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.004G089500
243 / 5e-82
AT3G17980 245 / 4e-84
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.010G110900
220 / 1e-72
AT1G70790 233 / 3e-79
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Potri.008G131500
219 / 2e-72
AT3G17980 202 / 5e-67
Arabidopsis thaliana C2 domain, Calcium-dependent lipid-binding (CaLB domain) family protein (.1)
Potri.011G098500
153 / 1e-44
AT4G21160 506 / 0.0
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Potri.001G372000
142 / 1e-40
AT4G21160 506 / 0.0
ARF-GAP domain 12, Calcium-dependent ARF-type GTPase activating protein family (.1.2.3.4)
Potri.016G005300
134 / 3e-39
AT5G47710 264 / 1e-91
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Potri.006G004600
132 / 1e-38
AT5G47710 270 / 3e-94
Calcium-dependent lipid-binding (CaLB domain) family protein (.1), Calcium-dependent lipid-binding (CaLB domain) family protein (.2)
Potri.003G198301
135 / 7e-37
AT3G07940 503 / 7e-178
Calcium-dependent ARF-type GTPase activating protein family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0154
C2
PF00168
C2
C2 domain
Representative CDS sequence
>Lus10009053 pacid=23173318 polypeptide=Lus10009053 locus=Lus10009053.g ID=Lus10009053.BGIv1.0 annot-version=v1.0
ATGAGGAACAGGAATTCAAAGCTGAAGAAGGCCCAGACGACGGTGGCTCAAATGCTGAGCTTCCCGCCGGCCAATCATATCGAACGGCTATTCCGATTCA
GCTCGTCTAAGAAGCCACCTACCCATCGTCGCACTAAAAGCGACCCAGTGGAACTTGACCAAGAGCAGTATAAAATGAACAAGATGACGGACTCGCCAAC
TGGAGTAGCAAAATCAACGCCGCCGCCGTCATCGTCGTCGCCGTCAGCAGAAATGACTGGAACACCTTCTTCGATGGAAAGCCTCCTAGGTCTCCTCCGC
ATCCGAGTCAAGCGGGGCATCAACCTCGCCGTCCGTGACGTTCTCAGCAGCGATCCTTACTGCGTCGTCCGGATGGGAAAACAGAAACTGAAGACTAGAT
TTGTAAAGAAGGATGTAAATCCAGAATGGAATGAAGATTTAACTCTGTCGATTATGGAACCGGAATTGCCCATCAAGCTGACTGTGTACGACCACGACAC
ATTCAGCCTAGACGACAAAATGGGTGACGCAGAGTTCGACATCATACCATTCATGGAGGTACTACGCACAAACATCAAAGGCCTCCCAAACGGGACCGTA
ATAAAAAAGCTACAGCCAAGCAGGCAAAACTGCTTGTCCGAGGAGTCCAGCATCACATGCACTGATGGAAAGATCCTCCAGGACATGTGCCTCAGACTTA
GAAACGTGGAATGCGGAGAAGTCGAGCTCCAATTGCAGTGGATCGACCTCCCCGGTGCCAGGGGCTTTTAG
AA sequence
>Lus10009053 pacid=23173318 polypeptide=Lus10009053 locus=Lus10009053.g ID=Lus10009053.BGIv1.0 annot-version=v1.0
MRNRNSKLKKAQTTVAQMLSFPPANHIERLFRFSSSKKPPTHRRTKSDPVELDQEQYKMNKMTDSPTGVAKSTPPPSSSSPSAEMTGTPSSMESLLGLLR
IRVKRGINLAVRDVLSSDPYCVVRMGKQKLKTRFVKKDVNPEWNEDLTLSIMEPELPIKLTVYDHDTFSLDDKMGDAEFDIIPFMEVLRTNIKGLPNGTV
IKKLQPSRQNCLSEESSITCTDGKILQDMCLRLRNVECGEVELQLQWIDLPGARGF
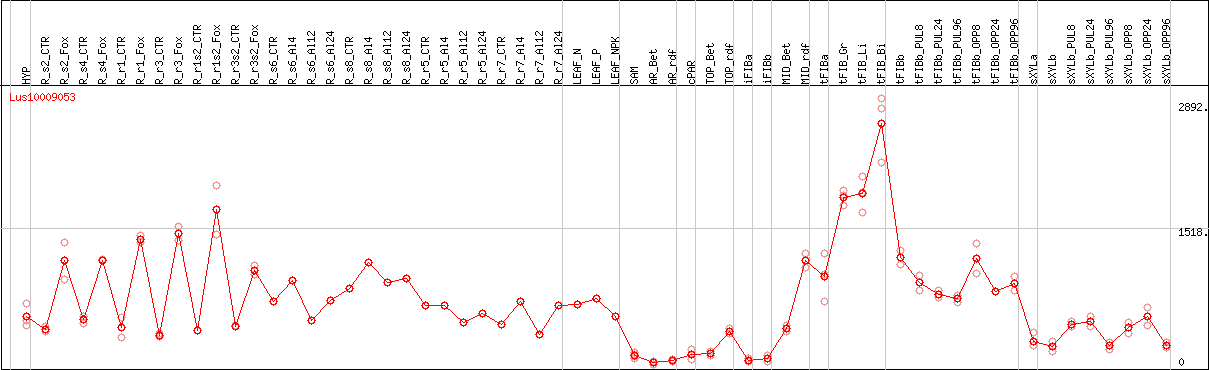
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009053 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.