External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G02500 560 / 0
AtHsp70-1, AT-HSC70-1, HSP70-1, HSC70-1
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
AT3G09440 559 / 0
Heat shock protein 70 (Hsp 70) family protein (.1), Heat shock protein 70 (Hsp 70) family protein (.2)
AT5G02490 557 / 0
AtHsp70-2
Heat shock protein 70 (Hsp 70) family protein (.1)
AT1G56410 555 / 0
HSP70T-1, ERD2
HEAT SHOCK PROTEIN 70T-1, EARLY-RESPONSIVE TO DEHYDRATION 2, heat shock protein 70 (Hsp 70) family protein (.1)
AT3G12580 545 / 0
ATHSP70, HSP70
ARABIDOPSIS HEAT SHOCK PROTEIN 70, heat shock protein 70 (.1)
AT1G16030 503 / 2e-176
HSP70B
heat shock protein 70B (.1)
AT5G42020 387 / 2e-131
BIP2, BIP
luminal binding protein, Heat shock protein 70 (Hsp 70) family protein (.1), Heat shock protein 70 (Hsp 70) family protein (.2)
AT5G28540 387 / 1e-130
BIP1
heat shock protein 70 (Hsp 70) family protein (.1)
AT1G09080 382 / 6e-129
BIP3
binding protein 3, Heat shock protein 70 (Hsp 70) family protein (.1), Heat shock protein 70 (Hsp 70) family protein (.2)
AT4G37910 285 / 5e-91
MTHSC70-1
mitochondrial heat shock protein 70-1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024393
588 / 0
AT5G02310 1770 / 0.0
proteolysis 6 (.1)
Lus10025276
587 / 0
AT5G02500 1233 / 0.0
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
Lus10024407
586 / 0
AT5G02500 1236 / 0.0
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
Lus10025333
581 / 0
AT5G02500 1236 / 0.0
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
Lus10005953
547 / 0
AT5G02500 1207 / 0.0
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
Lus10002319
518 / 0
AT3G12580 1160 / 0.0
ARABIDOPSIS HEAT SHOCK PROTEIN 70, heat shock protein 70 (.1)
Lus10026099
500 / 6e-180
AT1G16030 504 / 1e-176
heat shock protein 70B (.1)
Lus10021345
375 / 1e-127
AT5G28540 843 / 0.0
heat shock protein 70 (Hsp 70) family protein (.1)
Lus10017306
380 / 4e-127
AT5G28540 1169 / 0.0
heat shock protein 70 (Hsp 70) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G054600
568 / 0
AT5G02500 1149 / 0.0
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
Potri.008G054000
563 / 0
AT5G02500 1160 / 0.0
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
Potri.010G205700
563 / 0
AT5G02500 1119 / 0.0
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
Potri.010G206600
562 / 0
AT3G12580 1181 / 0.0
ARABIDOPSIS HEAT SHOCK PROTEIN 70, heat shock protein 70 (.1)
Potri.010G205800
560 / 0
AT5G02500 1142 / 0.0
HEAT SHOCK PROTEIN 70-1, ARABIDOPSIS THALIANA HEAT SHOCK COGNATE PROTEIN 70-1, heat shock cognate protein 70-1 (.1.2)
Potri.001G042600
529 / 0
AT3G12580 1061 / 0.0
ARABIDOPSIS HEAT SHOCK PROTEIN 70, heat shock protein 70 (.1)
Potri.001G042700
515 / 0
AT3G12580 1058 / 0.0
ARABIDOPSIS HEAT SHOCK PROTEIN 70, heat shock protein 70 (.1)
Potri.003G184000
499 / 5e-175
AT3G12580 1040 / 0.0
ARABIDOPSIS HEAT SHOCK PROTEIN 70, heat shock protein 70 (.1)
Potri.008G054900
447 / 1e-154
AT3G12580 947 / 0.0
ARABIDOPSIS HEAT SHOCK PROTEIN 70, heat shock protein 70 (.1)
Potri.001G087500
385 / 4e-130
AT5G42020 1184 / 0.0
luminal binding protein, Heat shock protein 70 (Hsp 70) family protein (.1), Heat shock protein 70 (Hsp 70) family protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0108
Actin_ATPase
PF00012
HSP70
Hsp70 protein
Representative CDS sequence
>Lus10009071 pacid=23172426 polypeptide=Lus10009071 locus=Lus10009071.g ID=Lus10009071.BGIv1.0 annot-version=v1.0
ATGGCCGGTAAAGGAGAGGGTCCTGCTATCGGTATCGATCTCGGTACCACTTATTCTTGCGTCGGTGTCTGGCAGCATGATCGCGTTGAGATCATCGCCA
ATGACCAGGGTAACCGGACTACCCCGTCGTACGTCGGATTCACTGATTCCGAGCGTTTGATCGGTGACGCCGCCAAGAACCAGGTCGCCATGAACCCCAT
CAACACTGTTTTCGATGCTAAGAGGTTGATCGGTAGGCGTTTCAGCGACGCCTCAGTCCAAAGTGACATCAAGTTGTGGCCTTTTAAGCTTATCCCCGGC
GCTGCTGATAAGCCTATGATTGTTGTCAACTACAAGGGCGAGGAGAAGCAGTTTGCTGCCGAGGAAATTTCTTCCATGGTTTTGATCAAGATGCGTGAGA
TCGCTGAGGCTTTCTTAGGTACTACCGTGAAGAATGCCGTTGTTACAGTTCCTGCCTACTTCAACGATTCTCAGCGTCAAGCTACCAAGGATGCTGGTGT
CATTGCTGGTCTTAATGTCATGCGAATCATCAACGAACCCACAGCTGCTGCCATTGCCTACGGTCTTGACAAGAAGGCCACCAGCGTTGGAGAGAAGAAC
GTGCTGATCTTTGATCTTGGCGGTGGTACTTTCGATGTGTCCCTCCTCACCATTGAGGAAGGTATCTTCGAGGTTAAGGCCACAGCTGGAGACACCCATC
TTGGAGGGGAAGACTTCGACAACAGAATGGTCAACCACTTCGTCCAGGAGTTCAAGAGGAAGCACAAGAAGGACATCTCTGGAAATCCCAGAGCCCTCAG
AAGATTGAGGACATCTTGCGAGAGGGCCAAAAGAACTCTCTACTCCACCGCGCAAAAAAAAAAGATTTCTTCAACGGCAAGGAATTGTGCAAGAGCATTA
ACCCTGACGAGGCTGTTGCGTATGGTGCTGCTGTCCAAGCTGCCATCTTGA
AA sequence
>Lus10009071 pacid=23172426 polypeptide=Lus10009071 locus=Lus10009071.g ID=Lus10009071.BGIv1.0 annot-version=v1.0
MAGKGEGPAIGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVGFTDSERLIGDAAKNQVAMNPINTVFDAKRLIGRRFSDASVQSDIKLWPFKLIPG
AADKPMIVVNYKGEEKQFAAEEISSMVLIKMREIAEAFLGTTVKNAVVTVPAYFNDSQRQATKDAGVIAGLNVMRIINEPTAAAIAYGLDKKATSVGEKN
VLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGEDFDNRMVNHFVQEFKRKHKKDISGNPRALRRLRTSCERAKRTLYSTAQKKKISSTARNCARAL
TLTRLLRMVLLSKLPS
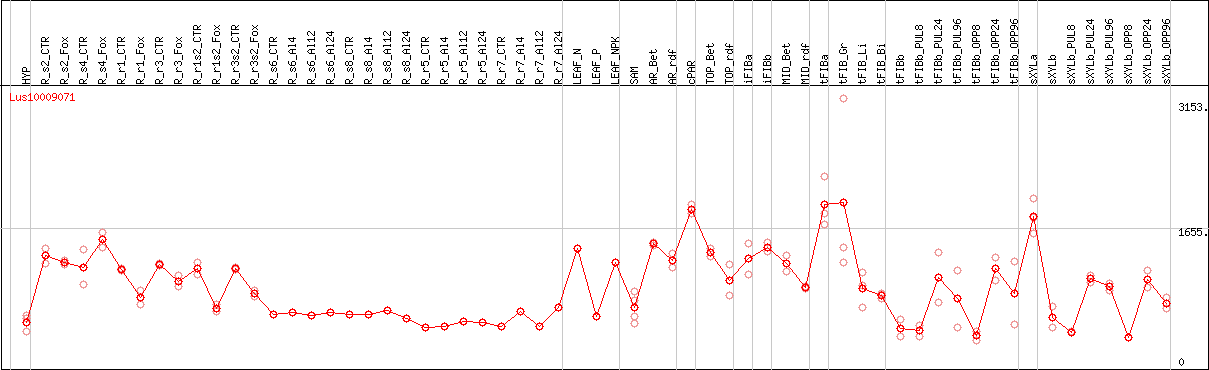
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009071 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.