External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G23580 194 / 2e-61
ATMES4, ABE4
ARABIDOPSIS THALIANA METHYL ESTERASE 4, ALPHA/BETA FOLD HYDROLASE/ESTERASE 4, methyl esterase 4 (.1.2)
AT3G50440 193 / 1e-60
ATMES10
ARABIDOPSIS THALIANA METHYL ESTERASE 10, methyl esterase 10 (.1)
AT2G23600 192 / 1e-60
ATMES2, ACL, ATME8
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
AT2G23620 191 / 4e-60
ATMES1
ARABIDOPSIS THALIANA METHYL ESTERASE 1, methyl esterase 1 (.1)
AT2G23610 186 / 3e-58
ATMES3
ARABIDOPSIS THALIANA METHYL ESTERASE 3, methyl esterase 3 (.1)
AT2G23560 181 / 4e-56
ATMES7
ARABIDOPSIS THALIANA METHYL ESTERASE 7, methyl esterase 7 (.1)
AT4G37150 179 / 8e-56
ATMES9
ARABIDOPSIS THALIANA METHYL ESTERASE 9, methyl esterase 9 (.1)
AT2G23550 179 / 1e-55
ATMES6, ABE1
ALPHA/BETA FOLD HYDROLASE/ESTERASE 1, methyl esterase 6 (.1.2)
AT2G23590 172 / 1e-52
ATMES8
methyl esterase 8 (.1)
AT5G58310 164 / 1e-49
ATMES18
ARABIDOPSIS THALIANA METHYL ESTERASE 18, methyl esterase 18 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005401
505 / 0
AT2G23600 184 / 5e-56
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
Lus10009203
389 / 3e-137
AT2G23620 205 / 6e-65
ARABIDOPSIS THALIANA METHYL ESTERASE 1, methyl esterase 1 (.1)
Lus10005402
386 / 1e-136
AT2G23580 204 / 3e-65
ARABIDOPSIS THALIANA METHYL ESTERASE 4, ALPHA/BETA FOLD HYDROLASE/ESTERASE 4, methyl esterase 4 (.1.2)
Lus10005400
357 / 2e-125
AT2G23610 206 / 8e-66
ARABIDOPSIS THALIANA METHYL ESTERASE 3, methyl esterase 3 (.1)
Lus10015532
232 / 3e-76
AT2G23600 240 / 2e-79
ARABIDOPSIS THALIANA METHYL ESTERASE 2, ARABIDOPSIS METHYL ESTERASE 8, acetone-cyanohydrin lyase (.1)
Lus10022467
226 / 9e-74
AT3G50440 238 / 2e-78
ARABIDOPSIS THALIANA METHYL ESTERASE 10, methyl esterase 10 (.1)
Lus10038592
186 / 4e-58
AT2G23620 306 / 4e-105
ARABIDOPSIS THALIANA METHYL ESTERASE 1, methyl esterase 1 (.1)
Lus10023326
177 / 1e-54
AT3G50440 263 / 3e-88
ARABIDOPSIS THALIANA METHYL ESTERASE 10, methyl esterase 10 (.1)
Lus10003511
148 / 2e-42
AT4G09900 526 / 0.0
ARABIDOPSIS THALIANA METHYL ESTERASE 12, methyl esterase 12 (.1)
PFAM info
Representative CDS sequence
>Lus10009205 pacid=23145606 polypeptide=Lus10009205 locus=Lus10009205.g ID=Lus10009205.BGIv1.0 annot-version=v1.0
ATGACGAGTAGGCATTTCGTACTACTTCATGGAGCCTGCAACGGAGCGTGGATCTGGTACAAAGTGAGTGCGCTGCTGAAAGAGGCTGGTCATAAGGTCA
CCGCCATTGACCTGGCTTCTTCTGGGATCCACCCCAAGAAATGGGAAGAAGTTAATTCTTTCCTGGAATACTCTGAGCCCTTGCTTACTTTCATGGAATC
TCTTCCTAGTGACGGCGAAAAGGTTGTCATTGTAGCTCACAGCCTCGGCGGCTATAGCGCCTCCATTGCCATGGAAAGGTTCCCTGAGAAGATCTCTCTT
GGTGTCTTTGTCACCGCCAGCATGCTCGGCCCTGATCTTAACTACGAACTTATGGCTGAGAAGTATGTTGAATTTGCTGCGCCCATGATGGACACTGTAT
TTACATATGGGAATGGCCCAGACAAAACACCAACCGTGGCGATGGTTGGTCCCAAGTACATGGAGGCTGTGAACTATCGGAAATCTCCCCGTGCGGACCT
GGAGCTGGGATTAACATTGGTAAGACCGACTCCGGTGTTCGATCAAGAACAAGCTGCAATAGACACAAGAGTGACAAAGGAGAAATACGGAACTATTCCT
CGGGTGTTTGTGCTTTGCGATGCTGAGTATAAGGGAGCATTCCAGTGGTGGCAAGTTAAGAACAACCCTCCTGATGAACACATACTAATTGAAGAGTCTG
ATCACATGGTCATGCTGTCACAACCTGGTCCCCTCGTTGATTATCTCCTCAAAGTTGGTGCCAAATACCTCTGA
AA sequence
>Lus10009205 pacid=23145606 polypeptide=Lus10009205 locus=Lus10009205.g ID=Lus10009205.BGIv1.0 annot-version=v1.0
MTSRHFVLLHGACNGAWIWYKVSALLKEAGHKVTAIDLASSGIHPKKWEEVNSFLEYSEPLLTFMESLPSDGEKVVIVAHSLGGYSASIAMERFPEKISL
GVFVTASMLGPDLNYELMAEKYVEFAAPMMDTVFTYGNGPDKTPTVAMVGPKYMEAVNYRKSPRADLELGLTLVRPTPVFDQEQAAIDTRVTKEKYGTIP
RVFVLCDAEYKGAFQWWQVKNNPPDEHILIEESDHMVMLSQPGPLVDYLLKVGAKYL
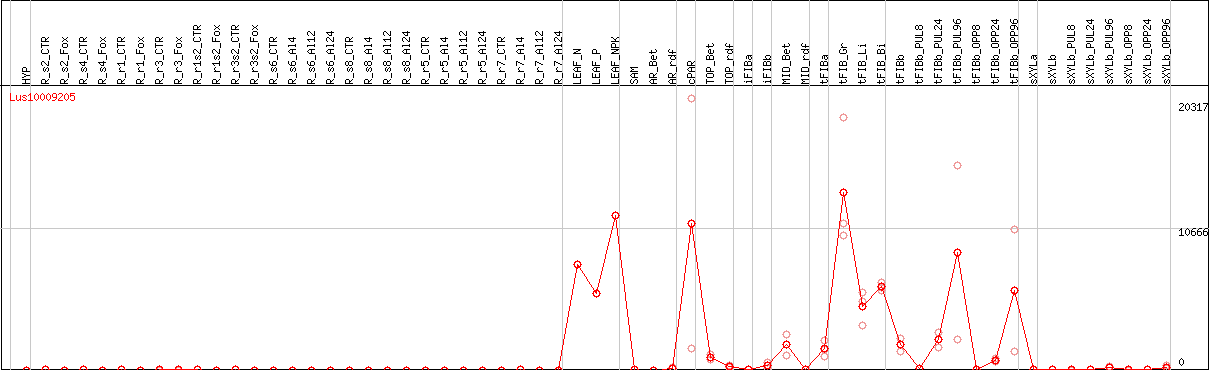
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009205 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.