Lus10009218 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10009218 pacid=23160197 polypeptide=Lus10009218 locus=Lus10009218.g ID=Lus10009218.BGIv1.0 annot-version=v1.0
ATGGGAAACGTGGAAATACCAAAATGGCTGCAAGGGTTGCCACTGGCTCCCGAGTTCCGGCCAACCGATACGGAATTCGCTGACCCGATTGCTTATATTT
CGAAAATTGAGAAGGAAGCCGGTGCTTTTGGGATGTGCAAGATTATCCCACCTCTACCTAAACCTTCCAAGAAATATGTGTTTTCCAATCTGAATAAGAC
CCTCGCAAAATCGCCTGAATTGGGGGATAATTGTTCCTCTTTGAAATCGGGTTCTGGAGTTGATGGTAAAGATGGAGATATGAGGGCGGTGTTTACGACA
AGGCACCAAGAACTAGGACAGAGTATCAAGAAGAAGGGTGTCGAAAAGGAGACTCCGCAAATGGTTTTTCATAAACAAGTGTGGCAAAGTGGAGAAGTCT
ATACTTTGGAACAGTTCGAATCTAAGTCCAAGGCCTTTGCAAAGAGTATGTTGGGTTCAGTTAAGGAGGTTAGTCCTTTAGTCATAGAGGCCCTGTTCTG
GAAGGCAGCTTTTGAGAAACCAATCTATGTGGAATATGCAAATGATGTGCCTGGATCAGCTTTCGGAGAACCAGAAGGTCAATTTCGGTATTTCAATCGA
CGGAGAAGGAGGAGGGCATCATATCAATCCTACCGAAGGCATGGGGAAGGCTACAATTGCCAGAACAATGAATTGATAGCTGTTGATAATAATGCAGCTG
ATGAGGTGGTTGATGCTTCTATTAAAAGTGATCCAAGTCCATCATCTCTGTCTACTTCATCAACTGCACTTTCATCCAAATTTGTGAAATCTTATAACCA
AAAAGGTGAAAGTTCTAGTTGCGATGTGGAGGGCACAGCAGGCTGGAAGCTTTCTAACAGCCCTTGGAATCTCCAAATGATTGCCCGTTCACCTGGGTCC
CTCACACGCTATATGCCGGATGATATCCCAGGGGTTACTTCTCCTATGGTTTATATTGGTATGTTATACAGTTGGTTTGCTTGGCATGTTGAGGATCATG
AGCTTCACAGCATGAATTTTCTCCATACTGGCTCTCCAAAAACATGGTATTCTGTTCCTGGAGATTATGCTTTTGCTTTTGAGGAGATTATCCGCAGACA
GGCTTATGGTGGTAAATTGGATCGTTTAGCTTCTTTGTCAAGGTTGGGTGAGAAGACAACCCTTCTATCACCTGAGGTAGTTGTTTCATCTGGCATTCCT
TGTTGCAGGGTTATCCAGAACCCGGGGGAGGTTGTTGTGACTTTTCCAAGGGCTTATCATGTAGGATTCAGCCACGGTTTTAATTGTGGAGAAGCTGCTA
ATTTTAGTACTCCGCAATGGCTTGAAGTAGCTAAGGAAGCTGCAGTACGTAGAGCTGCTATGAATTACCTTCCAATGCTTTCGCATCAGCAACTACTATA
CCTGTTGACAATGTCTTTTGTCCTAAGAGTCCCAAGGTCACTCCTTCCTGGTGCTAGGAGTTCTCGCCTAAGAGATCGCCAGAAAGAAGAAAGAGAGTTG
TCTGTGAAGAGGGCTTTTATAGAAGATCTATTACGCGAGAATAATACTTTACGTGCTATTCTCAGAAAAGGTCCTGGCTGTAAAGCTGTGATATGGAATC
CTGATTTGCTACCTCGTGCAAGTAAACAGTCTCAATTGATTGATACCTGTACTGCTGCTACCAGAATGGAAGAAGATTTATGTCATACACGGCTTGAACA
TGAGAATAACAGTACAAGCAATCTGTTTACAGAAATGAGCAAGTATGTTGAGTCTTTGAATGATTTATATGCTGATGGAGGTGATATCTCATGTGATTTT
CAAGTCGATTCGGGTTCACTGGTTTGTATAGCTTGTGGAATCCTTGGTTTTCCATTCATGTCTGTGGTGCAACCATCTGAGAAAGCTGCTGAAGAATCAT
TATATGTGGATAATACATTCAGAAGGGTCCATTCTTCTGCAGATTGCAATGATGATCCTCATTCTGTGCCTGACCAAACTCCAGCTCTAAAAGATCTTAA
AGTGCCTACTGGATGGGATATTTCCAATAAATTTCTCAGACCCCGGATTTTCTGCCTGGAGCATGGCCTTCAAATTTGTGAGCTTTTACAGTCTAGAGGT
GGAGCAAACATGCTTGTAATGTGCCATTCAGATTTTCAGAAAATGAAGGCCTATGCTACTTCTATATCGGAGGAGATCAATGTTCCTTTTGACTACACCC
ATATTCCGCTAGAAAGTGCGTTGAATGAAGACTTGAACTTGATTGACCTTGCGATTGATGGCCATGACTGGGATGGACGTGGGGAGGATTGGACATCCAA
AATGGGTGTCAACTTATTACACTGTGTGAAGATCAGGAAAAAGTCCCCATCAACGCACGTGCATCATGCATTGGCTTTAGCAGGACTCTTCACTGAAGGA
AGTCTCAGTTCGGAGTTCTCTACACTTAAGTGGGAATCCCGAAGATCTCGGTCAAGAAACAAGTTGAATCATTCTGCCAGTTTTTCAGGTGGAAAATTTG
AGTCTAAGAAAGATGATATGCCGGAGAAAGTTTCAGATGGTGTAGCCAGAAAAGAAGAAAAGCTTCTATGTTATACAAGACGGAAGTCTAAAAATAAAGA
AGATTGTTCTATAAATGGTGGTCAAGATCAGCTATATCTCATCCGGCAAACCAGCAAGGTTTCTCGAAACAGTAAGGTAGGAATAAATAGTTTGGAGGAC
CTTGGGGTCAATCCTGACGGGGTTTCTGAAAACCACAATGAAGTTCAATTTCATGAAGCTACTGCTACCACGACTGCAGTTGTCAACTCTGGGCCTTCAG
TAGATACGACATGTTCTGCTGTCATGTCTGATCCAGAAGCCTTCTCGAATTCTGAAATGCGGAATGGCATAATTGCTGCATTAGAAAGCAGTGAGCGTGG
GCTCAATTCTAATGTAGCTACAGATAATGGGTTATTGGTGATTTGTAAAGATGATACCGAAAATGTGATCACTTATGATAGAGCTTCTAAAAGGGGCCTC
AATCTCCAGGTGGATAGAGATGGTCTTGTCAGTGAGACTTCAGATGTTGCTAATTCACCTTCAGCCGAAGCTGAGCTTGTAGTAGCAGAGACTTGTTGCA
CAAAGCGTGAAGCTTGTAATCATGTGACTTCACAAAGTGAACAACAGGAAGAAGGCCAAAAATTGCCTAATAGCAATTATGTTATTTCATCTAATGAGAC
AGTGACTACGGAGACCATTACCTTGAATGGTGAAGCTTGCGAGCAGTTGCAAGACATCTCTCTTGTAGAAGACGCCATGCAGAAACAAACAGAATATTGC
GATGATAGGGAGGAAATGATGCAGTCGAAACTTCAATCAGAAGAAGAGCCAGACTTGGGGCCTTTGGATGGAAGTGGAAGCTCTGAGAGTGTTAGGGAGC
AAATTGACCTCCCAGTTACAACCCAAGAGCTAGCTTCCAGTTCCACCTCAGAAATGGAGGATCAGCCTATTCCTGAGACAATGGAAGCATGCTGCGAACC
TTGTGCAACTAGTCGTACTGATGAAGATTCATCTGCTTTTAAGAATTCAGATGGAGAAGTAGAGCCAGTAAATAACTCAGTTGATGGGATGCCAGAGTGT
TTTGAAATTCGGACGGGTTCTGAAATGGATGAAGAGTCTTCTGCTGTAAGAAGTGCAGTTGAAGAAAACAGATCATCTCCACCACACGGACATGGAAGTA
TTGAGTCTGCTAGTTCTGGATCAATCCATGTCAAGGTCAGAAAGAGGAGAAGAGAATTGGACCAGATAACAGAAGATAATCTCAATTTCAAGAGCTTCGT
TAGAAGTCCATGTGAAAGGTTGAGACCCAGAAGAATGGGCAGAGATGACCCAAGCACAAGCGGAGCTGATTTGGAAAAAATTCCTGTGGAGAATGCAGTA
GCAAAGAGGTCGAGGTCGAAGAGGCCTTCAAATGATCCAGTTAATAGTCAGTTGAAGAAAAAGCAGGTGACAAAAAGACCTCACAAGTGTGAAATAGAAG
GTTGCGGCATGAGATTTGAGACGAAGGAAGACCTAGTCTTGCACAGTCGCAATCGCTGCCCATATGAAGGATGCAGGAAGAAGTTTAGTTCGCACAGGTA
TGCATTGATACATCAGCGGGTACATGATGACGATAGACCCTTGAAATGTCCATGGAAGGGTTGTAAAATGGCGTTTAAGTGGGCATGGGCTCGGACTGAG
CACATAAGGGTGCATACAGGTGAGAAGCCCTACCATTGCAAAGTTGATGGATGTGGGCTTGCATTCAGCCCTCTGATCATCATCATCATCATCATCTACC
ATCTCGCAACCATTCTTCACCTTGTCAGCTCAGACTCCATAAGCAGAGCTCATTTCCCTGATGGGTTCATCTTTGGGACAGCCTCTTCTGCTTATCAATT
TGAAGGAGCTGCAACTGAAGGGAATAAAGGAGCTAGCATATGGGATACCTTCACAAGAAAGCCCGGGTCGTCTTCGCCAGGGAGAATTTTGGATTTCAGC
AATGCTGATGTCGCTGTTGATCAATACCACAGGTTTCAGGATGATATAGGGTTGATGAAAGATTTGGGAATGGATACCTACAGATTCTCCATCTCATGGG
CAAGAATCTTTCCAAATGGAACAGGGAATCTAAACCCAGAAGGCATAACATATTACAACAAACTCATTGATGGGTTACTAGAAAAAGGAATCAAGCCTTA
TGTGACTTTATATCACTGGGATCTTCCGCAGATGCTTCAGGACAAGTATGGAGGCTGGTTGGACAAGAAAATTATAGAGGACTTTGAGCATTATGCTAAC
ACCTGCTTCAAGGCATTTGGAGATAGAGTGAAAAACTGGATTACCTTCAATGAACCTCACAACTTTGCCATCCAGAGCTATGATTTTGGCATTCAAGCCC
CTGGGAGATGCTCCATTATTGGTCATCTATTCTGCAAGGAAGGAAATTCGTCGGCTGAACCATATATAGTAGCTCACCACATCCTCTCGTCACATGCAGC
TGCTTATCACAGCTACCAGTCATATTTCAAGGGGAAACAAGGTCGCATTGGCATAGTATTAGATGCCAAATGGTTCGAACCAAAATCTGATGCTGATGAG
GACAAAGATGCTGCAAATAGAGCAATGGACTTCACACTTGGATGGTTTCTTGATCCAATTTTCGTTGGCCGATACCCACTTTCAATGAGAAAGCTTGCTG
GTGAACGACTACCTGAGATCTCTTCAGAGATGGCAAAGCTAATTACGGGGTCAGTGGACTTTGTTGGCCTCAACCATTACACAACATCTTATGTTCGGAA
CGACAGAAGTAGGTTTCAGAAGCTGATACTTCAAGACGCTTCCACTGATTCAGCTGTCATTACTTCCTCTTACAGGAATGGAATTGCTATTGGAGAAAAA
GCAGCATCCAGTTGGCTAAGGATTGTCCCTTGGGGTATCCGGAAGTTGATGAATTATGTAAAAGATAAGTATGGAAACCCACTTGTGATCATAACAGAGA
ATGGAATGGACGATCCCAATAGCCCATTCATACCAAGAGAAAAGGCATTGCAGGATGAGAAAAGGATAGAGTACCACAGAGATTATCTCTCAAACCTGTC
TGCATCCATAAGGGAAGACGGGTGCAATGTAGGAGGGTATTTCATATGGTCATTGCTAGACAACTGGGAATGGAACTCGGGGTACACAGTCCGGTTCGGG
CTATACTATGTTGACTACAGTCATAACCTCACAAGGATACCAAAAGCTTCTGTAAATTGGTTCAAAAGTATGCTGAGATCACAAAGTACTGAACTTGATA
GCCAGAGTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10009218 pacid=23160197 polypeptide=Lus10009218 locus=Lus10009218.g ID=Lus10009218.BGIv1.0 annot-version=v1.0
MGNVEIPKWLQGLPLAPEFRPTDTEFADPIAYISKIEKEAGAFGMCKIIPPLPKPSKKYVFSNLNKTLAKSPELGDNCSSLKSGSGVDGKDGDMRAVFTT
RHQELGQSIKKKGVEKETPQMVFHKQVWQSGEVYTLEQFESKSKAFAKSMLGSVKEVSPLVIEALFWKAAFEKPIYVEYANDVPGSAFGEPEGQFRYFNR
RRRRRASYQSYRRHGEGYNCQNNELIAVDNNAADEVVDASIKSDPSPSSLSTSSTALSSKFVKSYNQKGESSSCDVEGTAGWKLSNSPWNLQMIARSPGS
LTRYMPDDIPGVTSPMVYIGMLYSWFAWHVEDHELHSMNFLHTGSPKTWYSVPGDYAFAFEEIIRRQAYGGKLDRLASLSRLGEKTTLLSPEVVVSSGIP
CCRVIQNPGEVVVTFPRAYHVGFSHGFNCGEAANFSTPQWLEVAKEAAVRRAAMNYLPMLSHQQLLYLLTMSFVLRVPRSLLPGARSSRLRDRQKEEREL
SVKRAFIEDLLRENNTLRAILRKGPGCKAVIWNPDLLPRASKQSQLIDTCTAATRMEEDLCHTRLEHENNSTSNLFTEMSKYVESLNDLYADGGDISCDF
QVDSGSLVCIACGILGFPFMSVVQPSEKAAEESLYVDNTFRRVHSSADCNDDPHSVPDQTPALKDLKVPTGWDISNKFLRPRIFCLEHGLQICELLQSRG
GANMLVMCHSDFQKMKAYATSISEEINVPFDYTHIPLESALNEDLNLIDLAIDGHDWDGRGEDWTSKMGVNLLHCVKIRKKSPSTHVHHALALAGLFTEG
SLSSEFSTLKWESRRSRSRNKLNHSASFSGGKFESKKDDMPEKVSDGVARKEEKLLCYTRRKSKNKEDCSINGGQDQLYLIRQTSKVSRNSKVGINSLED
LGVNPDGVSENHNEVQFHEATATTTAVVNSGPSVDTTCSAVMSDPEAFSNSEMRNGIIAALESSERGLNSNVATDNGLLVICKDDTENVITYDRASKRGL
NLQVDRDGLVSETSDVANSPSAEAELVVAETCCTKREACNHVTSQSEQQEEGQKLPNSNYVISSNETVTTETITLNGEACEQLQDISLVEDAMQKQTEYC
DDREEMMQSKLQSEEEPDLGPLDGSGSSESVREQIDLPVTTQELASSSTSEMEDQPIPETMEACCEPCATSRTDEDSSAFKNSDGEVEPVNNSVDGMPEC
FEIRTGSEMDEESSAVRSAVEENRSSPPHGHGSIESASSGSIHVKVRKRRRELDQITEDNLNFKSFVRSPCERLRPRRMGRDDPSTSGADLEKIPVENAV
AKRSRSKRPSNDPVNSQLKKKQVTKRPHKCEIEGCGMRFETKEDLVLHSRNRCPYEGCRKKFSSHRYALIHQRVHDDDRPLKCPWKGCKMAFKWAWARTE
HIRVHTGEKPYHCKVDGCGLAFSPLIIIIIIIYHLATILHLVSSDSISRAHFPDGFIFGTASSAYQFEGAATEGNKGASIWDTFTRKPGSSSPGRILDFS
NADVAVDQYHRFQDDIGLMKDLGMDTYRFSISWARIFPNGTGNLNPEGITYYNKLIDGLLEKGIKPYVTLYHWDLPQMLQDKYGGWLDKKIIEDFEHYAN
TCFKAFGDRVKNWITFNEPHNFAIQSYDFGIQAPGRCSIIGHLFCKEGNSSAEPYIVAHHILSSHAAAYHSYQSYFKGKQGRIGIVLDAKWFEPKSDADE
DKDAANRAMDFTLGWFLDPIFVGRYPLSMRKLAGERLPEISSEMAKLITGSVDFVGLNHYTTSYVRNDRSRFQKLILQDASTDSAVITSSYRNGIAIGEK
AASSWLRIVPWGIRKLMNYVKDKYGNPLVIITENGMDDPNSPFIPREKALQDEKRIEYHRDYLSNLSASIREDGCNVGGYFIWSLLDNWEWNSGYTVRFG
LYYVDYSHNLTRIPKASVNWFKSMLRSQSTELDSQS
|
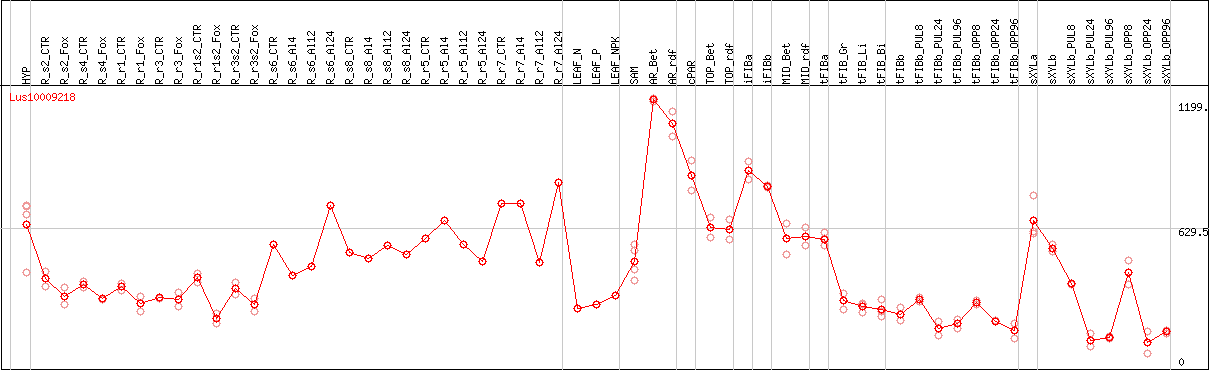
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009218 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
