Lus10009300 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10009300 pacid=23141848 polypeptide=Lus10009300 locus=Lus10009300.g ID=Lus10009300.BGIv1.0 annot-version=v1.0
ATGGCTTCCGATTCCCGCTTAAGCCAAGTCGTAGCCCCTGCCCTAGAGAAGGTCATCAAGAACGCATCCTGGAGGAAACACTCCAAGCTCGCGCATGAGT
GTAAATCCGTTCTCGAGAGGCTCACTTCTCCGCAGAAGCAGCAGCAGCCTTCTACTCCCGATGCTGACCCTGAAGCTTCCATTCCTGGCCCTCTTCATGA
TGGCGGGGCTATTGAGTACTCGCTTGCTGAATCCGAATCTATCCTCAGTCCTCTTATCAATGCTTGCAGCACTGGATTCCTCAAGATCGTGGATCCGGCC
ATTGATTGCGTTCAGAAATTGATCGCTCACGGCTACTTGCGAGGGGAGGCGGACCCGACCGGTGGTGCGGAGGGGAAGCTTCTGGCTAGACTGATCGAAT
CAGTTTGTAAATGCTACGATATAGGTGACGATGCAATTGAGCTTTCCGTTTTGAAAGCGCTTTTGTCCGCTGTGACTTCCATTTCCCTGCGGATACATGG
AGATTGCTTGTTGCAGATAGTGAGGACTTGTTACGATATTTATCTAGGGAGCAAGAACGTTGTAAATCAGACCACGGCTAAAGCTTCCTTGATCCAAATG
TTGGTCATTGTTTTCCGGAGAATGGAAGCGGATTCATCCACGGTACCCATTCAGCCCATTGTAGTGGCTGAACTAATGGAACCAGTTGAGAGATCCGATG
CAGATGGGTCTATGACGATGTTTGTGCAGGGTTTCATAACCAAAATCATGCAAGATTTCGACGGAGTTTTGAACGCTGGAGGTCCTAGTAAGATTACGGT
TGGAGCACACGATGGAGCCTTTGAGACTACAACGGTTGAGACCACAAACCCGGCAGATTTGTTAGATTCCACGGATAAGGATATGCTAGACGCCAAGTAT
TGGGAGATTAGTATGTATAAGACCGCATTGGAAGGTAGGAAAGGCGAATTGGCAGACGGGGAAGTAGAAAGAGATGATGATTTAGAGGTTCAGATTGGTA
ATAAATTAAGGAGAGATGCATTCTTGGTGTTTAGAGCTCTCTGTAAGTTGTCCATGAAAACTCCTCCAAAGGAGGCTGCAGCAGATCCTCAGTTGATGAG
GGGGAAAATTGTGGCGTTGGAGTTGTTGAAGATTTTATTGGAGAATGCTGGAGCTGTTTTTAGGACCAGCGACAGGTTTTTGGGTGCCATCAAGCAGTAC
TTATGCTTGTCTCTGCTGAAGAACAGTGCTTCAAGTCTCATGATCGTTTTTCAACTTTCTTGCTCTATTTTTGTGAGTCTCGTGTCAAGATTTAGAGCTG
GATTGAAAGCAGAAATTGGAGTATTTTTCCCTATGATTGTGCTCAGAGTTCTGGAAAGTGTTGCTCAACCTAATTTCCAACAGAAGATGATTGTTCTACG
GTTCATTGAAAAGCTCTGTGCGGATTCGCAGATCCAGGTTGATATCTTTATTAACTATGATTGTGATGTCAATTCATCGAACATATTTGAGAGAATGATC
AATGGACTCCTGAAAACTGCACAAGGTGTCCCCCCTGGTACCGTAACATCGCTCATGCCACCTCAGGAGGTGACTCTGAAACTTGAAGCCATGAAGTGCC
TAGTGGCCATTTTGAAATCAATGGGAGATTGGGTAAACAAACAGTTAAGAATTCCAGATCCTAGTTCTGCCAAAAAGGTTGATGCACCTGAGAACATTCC
TGAAAGTAGAAGTCTTTCCTTGGTCAATGGAAATGGGGAAGAGCCAGCTGAAGGATCAGATTCTCAGTCTGAAGCCTCCACCGAAGCTTCTGATGTCTCA
TCCATTGAGCAACGCAGGGCTTACAAACTGGAACTTCAGGAAGGTATTTCTCTTTTCAACCAGAAACCCAAGAAAGGAATTGAGTTTCTTATCAATGCAA
ATAAAGTTGGAAATTCACCCGAAGAGATTGCTGAATTTCTAAAAAGTGCTTCTGGCTTGAACAAGACTCTTATTGGCGATTATCTAGGGGAGAGAGAAGA
TTTACCGCTGAAAGTCATGCATGCATATGTGAATTCATTTGATTTTCAAGGCATGGAGTTCGATGAAGCTATAAGAGGCTTCCTCCTGGGCTTTAGGCTA
CCTGGTGAGGCTCAGAAGATTGACAGAATTATGGAGAAGTTTGCTGAGCGTTACTGCAAATGCAACCCCAAAGTTTTTATCAGTGCAGACACAGCTTATG
TTCTTGCTTACTCTGTGATATTGCTCAATACCGACGCCCACAATCCTATGGTGAAAAACAAGATGTCACCAGATGATTTTATTAGGAACAATCGTGGCAT
TGATGATGGAAAAGATCTTCCCGAGGAATACTTAAGGTCATTGTATGAAAGGATTTCAACAAATGAGATCAAAATGAAAGAAGATGATTTTAATGTCCAA
CAAAGACAGTCTACAAACTCTAACAGACTTTTGGGCTTGGATAATATTTTGAATATTGTCGTTCGTAAAAAAGGGGAGGACAGACATATGGAGACAAGTG
ATGATCTCATTAGGCATATGCAAGAGCAATTTAAAGAAAAAGCTCGCAAATCTGAGTCAGTTTATTATGCAGCTACCGATGTGGTTATACTTCGTTTCAT
GATTGAGGTATGCTGGGCTCCGATGTTAGCTGCCTTCAGTGTGCCTCTGGATCAAAGTGATGATGAAGTGGTGATAGGTCTTTGTCTCGAAGGTTTTCGG
TGTGCTATCCATGTTACTGCTGTAATGTCCATGAAGACGCATAGAGATGCTTTTGTGACTTCATTAGCAAAGTTCACATCCCTTCACTCTCCTGCAGACA
TCAAACAGAAAAATGTTGATGCAATCAAGGCAATAGTTACGATTGCAGATGAAGACGGGAATTATTTACAGGAAGCCTGGGAACATATTTTGACATGTGT
CTCTCGCTTTGAACATCTCCATCTATTGGGAGAGGGCGCTCCTCCAGATGCCACATTTTTCGCCTTCCCCCAAAATGAATCTGAAAAGTCAAAACCAGTC
AAGTCAAGTATCCTTCCCGTTCTGAAGAAGAAGGGTGCAGGAAGAATGCAGTATGCAGCTGCTGCTGTCCTGAGAGGTTCATATGACAGTGCTGGAATTG
GAGGCAATGTTTCTGGGAGTGTCACAAATGAGCAAATGAACAGTTTGGCCTCTAATCTGAACATGTTGGAACAAGTTGGGAGCTCTGAAATGAACCGCAT
ATTCACACGTAGCCAAAAGTTGAACAGTGAGGCCATCATTGACTTTGTTAAGGCTCTTTGCAAGGTATCCATGGAGGAGTTGCGTTCTGCATCTGATCCA
CGAGTGTTCAGCCTTACAAAGATTGTTGAGATTGCGCATTATAACATGAACCGCATCAGGCTAGTTTGGTCAAGCATCTGGCACGTGCTCTCAGATTTCT
TTGTAACCATTGGCTGTTCTGAGAACCTCTCCATTTCCATATTTGCAATGGACTCGTTGCGCCAGTTATCAATGAAGTTCTTGGAGCGTGAAGAGTTGGC
TAACTATAATTTCCAGAATGAATTCATGAAGCCATTTGTCATTGTCATGCGCAAGAGCAGTGCTGTAGAGATCAGAGAACTGATTATCAGATGCGTATCT
CAAATGGTTTTGTCTCGCGTGAATAATGTCAAATCAGGATGGAAGAGCATGTTCATGGTTTTCACAACAGCAGCATACGATGACCACAGAAACATTGTCC
TCCTGGCTTTTGAAATAATGGAGAAGATTATTCGTGATTACTTCCCGTATATTACTGAAACTGAGACGACAGCGTTTACTGATTGTGTGAATTGCCTCAT
TGCTTTCACCAATAGCCGATTTGACAAAGATATCAGCTTGAATGCAATTTCTTTCCTTCGGTATTGTGCTACAAAACTTGCAGAAGGAGATTTTGGATGT
GCCTCAGGGAACAAAGAAAAGGAAGCGTTGTCAAGACTTTCTCCATCTTCACCTCACACGGGCAAAGATGTAAAGCAAGAAAATGGGTACATAGATGATA
GGGAAGATCATGTATACCTTTGGTTCCCTTTGTTGGCAGGTCTGTCAGAACTTAGTTTTGATCCTAGGCCTGAAATCAGGAAGAGTGCCTTACAAATCCT
GTTTGATACTTTGCGCAATCATGGCCACCTTTTCTCTCTACCTTTATGGGAACGAGTGTTCGAGTCCGTCCTGTTTCCGATATTTGACTATGTACGGCAT
GCAATCGATCCAACAGGGGGTGACTCTCCTGGGCCAGGTAGTGACAGTGATTTGGGTGAGCATGATCAAGATGCATGGCTCTATGAGACTTGCACCTTGG
CTCTCCAGCTGGTTGTAGATCTTTTTGTCAAGTTCTACACCACTGTCAACCCACTATTAAGGAAGGTCTTGATGCTTCTGGTTAGCTTTATTAAACGCCC
TCACCAGAGCCTTGCTGGTATTGGCATTGCTGCTTTTGTTAGGTTGATGAGCAATGCTGGTGACCTTTATTCCGAGGAAAAGTGGTTAGAAGTGGTTTTG
TCACTAAGAGAAGCAGCTAACGCAACTCTTCCTGATTTCACCTTTTTAGTAAGTGGTGAGAATACAGTGCCAAGCCACAGGCATGTTTTAAGTGAAGAAC
CCGTCCAAGAAAATGGTGAATCTTCCGATGATCCAGAGCAGTTGATGGCGCAACGTCTTTATGCTGCAATATCTGATGCAAAGTGTCGTGCTGCTGTTCA
ACTTTTGCTGATTCAGGCAGTGATGGAGATTTACAATATGTACAGGCCTCAACTTTCAGCAAAAATTACACTCGTCCTCTTCACTGTGTTGCATGACATT
GCTTCCCATGCTCACAAGATCAACACGGATACCCAGCTTCGGTCAAGGTTATTGGAGTTCACCTTGATGACTCAAATGCAAGACCCTCCACTATTACGGC
TGGAGAACGAATCTTTCCAAATATGCCTCACATTCTTGCAGAATCTCATAGTAGACAGGCCGTCAAGCTACGACGAGACAGAAGTGGAGTCCTGCTTAGC
CAACCTCTGCCAGGAAGTTCTCAAGTTCTACATCAGCACAGCCAGTCATGGACAGGTGTCCTCTTCCTCCTCTCCAAGCGAGCGCCCTCATCCTCCGTCC
CTAATCCCTTTGGGTTCTGGGAAAAGGAGGGAATTAGCTGCTCGTGCACCACTAGTAGTTGTCACTCTCCATGCCATTTGTAGTTTGGGCGAGACCTCTT
TCCAGAGGAACTTGACCGTTTTCTTTCCTCTACTGTCAAGCTTGATAAGCTGCGAGCATGGGTCGAATGAGGTTCAGGTAGCTCTGAGCGAGATGCTGAG
CTCGTCGGTTGGTCCAGTCTTGCTTCGGTCTTGTTGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10009300 pacid=23141848 polypeptide=Lus10009300 locus=Lus10009300.g ID=Lus10009300.BGIv1.0 annot-version=v1.0
MASDSRLSQVVAPALEKVIKNASWRKHSKLAHECKSVLERLTSPQKQQQPSTPDADPEASIPGPLHDGGAIEYSLAESESILSPLINACSTGFLKIVDPA
IDCVQKLIAHGYLRGEADPTGGAEGKLLARLIESVCKCYDIGDDAIELSVLKALLSAVTSISLRIHGDCLLQIVRTCYDIYLGSKNVVNQTTAKASLIQM
LVIVFRRMEADSSTVPIQPIVVAELMEPVERSDADGSMTMFVQGFITKIMQDFDGVLNAGGPSKITVGAHDGAFETTTVETTNPADLLDSTDKDMLDAKY
WEISMYKTALEGRKGELADGEVERDDDLEVQIGNKLRRDAFLVFRALCKLSMKTPPKEAAADPQLMRGKIVALELLKILLENAGAVFRTSDRFLGAIKQY
LCLSLLKNSASSLMIVFQLSCSIFVSLVSRFRAGLKAEIGVFFPMIVLRVLESVAQPNFQQKMIVLRFIEKLCADSQIQVDIFINYDCDVNSSNIFERMI
NGLLKTAQGVPPGTVTSLMPPQEVTLKLEAMKCLVAILKSMGDWVNKQLRIPDPSSAKKVDAPENIPESRSLSLVNGNGEEPAEGSDSQSEASTEASDVS
SIEQRRAYKLELQEGISLFNQKPKKGIEFLINANKVGNSPEEIAEFLKSASGLNKTLIGDYLGEREDLPLKVMHAYVNSFDFQGMEFDEAIRGFLLGFRL
PGEAQKIDRIMEKFAERYCKCNPKVFISADTAYVLAYSVILLNTDAHNPMVKNKMSPDDFIRNNRGIDDGKDLPEEYLRSLYERISTNEIKMKEDDFNVQ
QRQSTNSNRLLGLDNILNIVVRKKGEDRHMETSDDLIRHMQEQFKEKARKSESVYYAATDVVILRFMIEVCWAPMLAAFSVPLDQSDDEVVIGLCLEGFR
CAIHVTAVMSMKTHRDAFVTSLAKFTSLHSPADIKQKNVDAIKAIVTIADEDGNYLQEAWEHILTCVSRFEHLHLLGEGAPPDATFFAFPQNESEKSKPV
KSSILPVLKKKGAGRMQYAAAAVLRGSYDSAGIGGNVSGSVTNEQMNSLASNLNMLEQVGSSEMNRIFTRSQKLNSEAIIDFVKALCKVSMEELRSASDP
RVFSLTKIVEIAHYNMNRIRLVWSSIWHVLSDFFVTIGCSENLSISIFAMDSLRQLSMKFLEREELANYNFQNEFMKPFVIVMRKSSAVEIRELIIRCVS
QMVLSRVNNVKSGWKSMFMVFTTAAYDDHRNIVLLAFEIMEKIIRDYFPYITETETTAFTDCVNCLIAFTNSRFDKDISLNAISFLRYCATKLAEGDFGC
ASGNKEKEALSRLSPSSPHTGKDVKQENGYIDDREDHVYLWFPLLAGLSELSFDPRPEIRKSALQILFDTLRNHGHLFSLPLWERVFESVLFPIFDYVRH
AIDPTGGDSPGPGSDSDLGEHDQDAWLYETCTLALQLVVDLFVKFYTTVNPLLRKVLMLLVSFIKRPHQSLAGIGIAAFVRLMSNAGDLYSEEKWLEVVL
SLREAANATLPDFTFLVSGENTVPSHRHVLSEEPVQENGESSDDPEQLMAQRLYAAISDAKCRAAVQLLLIQAVMEIYNMYRPQLSAKITLVLFTVLHDI
ASHAHKINTDTQLRSRLLEFTLMTQMQDPPLLRLENESFQICLTFLQNLIVDRPSSYDETEVESCLANLCQEVLKFYISTASHGQVSSSSSPSERPHPPS
LIPLGSGKRRELAARAPLVVVTLHAICSLGETSFQRNLTVFFPLLSSLISCEHGSNEVQVALSEMLSSSVGPVLLRSC
|
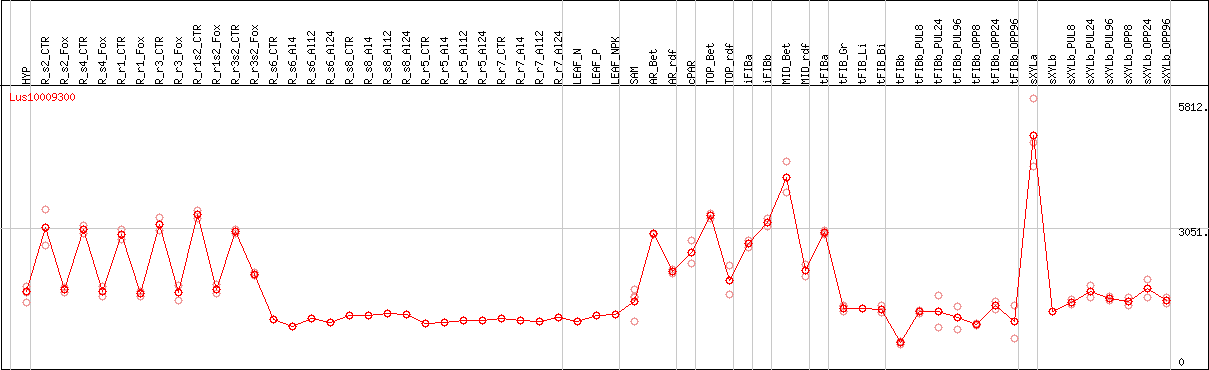
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
