External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G63320 58 / 3e-11
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G63150 58 / 7e-11
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G63130 55 / 1e-09
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G63070 54 / 2e-09
pentatricopeptide (PPR) repeat-containing protein (.1)
AT3G16710 54 / 2e-09
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G22470 54 / 3e-09
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G15930 53 / 5e-09
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G19290 53 / 5e-09
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G62930 52 / 7e-09
RPF3
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G63080 52 / 9e-09
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10003446
153 / 4e-48
AT1G62930 137 / 2e-37
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10003429
152 / 6e-47
AT1G62930 173 / 1e-49
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10014245
157 / 5e-46
AT1G12700 427 / 5e-141
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10014242
152 / 2e-45
AT1G62930 256 / 5e-78
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10014244
154 / 6e-45
AT1G12700 397 / 7e-130
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10014247
153 / 8e-45
AT1G62930 340 / 3e-109
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10003447
153 / 1e-44
AT1G12700 202 / 1e-56
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10003433
153 / 1e-44
AT1G12700 396 / 2e-129
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Lus10021074
145 / 6e-43
AT1G62930 273 / 5e-86
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G117600
80 / 2e-18
AT1G12700 330 / 1e-105
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.016G025600
72 / 1e-15
AT3G22470 426 / 3e-141
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G149800
71 / 2e-15
AT1G12700 482 / 6e-162
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G038400
69 / 2e-14
AT1G12700 478 / 7e-161
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.007G068400
68 / 3e-14
AT3G22470 300 / 8e-96
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.004G074500
67 / 8e-14
AT1G62930 478 / 1e-162
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G050300
66 / 1e-13
AT1G12700 466 / 5e-156
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G045000
65 / 3e-13
AT1G12700 502 / 4e-170
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.006G257300
65 / 3e-13
AT1G62930 481 / 3e-163
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G050500
64 / 5e-13
AT1G12700 523 / 2e-178
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10009331 pacid=23148917 polypeptide=Lus10009331 locus=Lus10009331.g ID=Lus10009331.BGIv1.0 annot-version=v1.0
ATGGTTTTTGGGCTTCATATATTTACTGGAAGGGTTAAAGAAGCAAGAGACATGTTTTCTAGGCTCTCTAAAGATGAGAGTTTACAGCCTAGTGTCTACA
CATATAGCACCGTAATTGATGGACTTTGTAGACATGGGTTGGTAGATGAAGCATATGATTTGTTTAGAACAATGGAGAGGAATCCACTCGAAGCAGTAGA
CCTAATACAAGAGATGGTGTCTAAGGGTTTCTCAGCTGATGCAACCACATTGGAATTGCTGTTAGAATTGTTGCCCAAAAACCAATTGGATCATCCACTT
CTCAGGAAGCTGGTAGGTGATAGTGAAGAGATAAAGCCTAAGGAGGGTTTATCAGCAGATACTAGTACTCCGTTATAA
AA sequence
>Lus10009331 pacid=23148917 polypeptide=Lus10009331 locus=Lus10009331.g ID=Lus10009331.BGIv1.0 annot-version=v1.0
MVFGLHIFTGRVKEARDMFSRLSKDESLQPSVYTYSTVIDGLCRHGLVDEAYDLFRTMERNPLEAVDLIQEMVSKGFSADATTLELLLELLPKNQLDHPL
LRKLVGDSEEIKPKEGLSADTSTPL
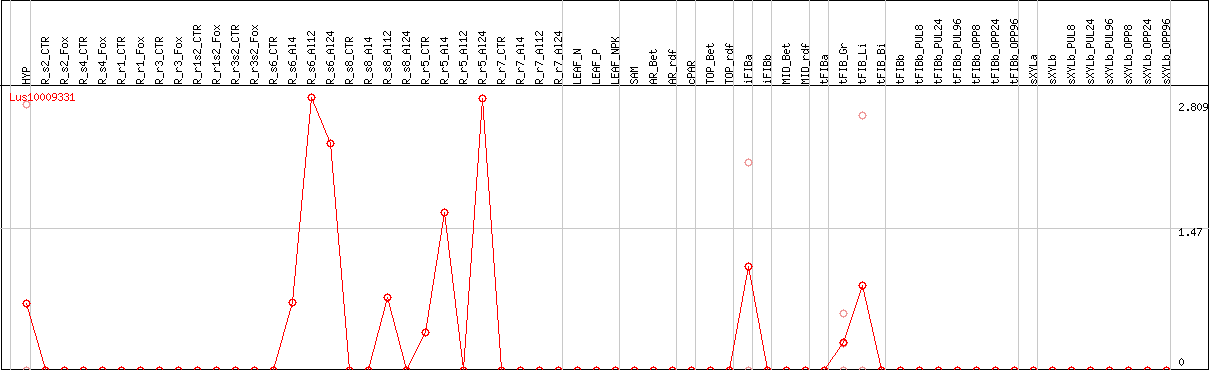
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009331 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.