Lus10009366 [FLAX]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10009366 pacid=23148915 polypeptide=Lus10009366 locus=Lus10009366.g ID=Lus10009366.BGIv1.0 annot-version=v1.0
ATGGAAGGCGAAGAGCAGCACCCAGTAGTCGCCGGAAGGCGGAGGAGCAGGGGCGGAGGCGACACGGCAAGGGCGGAAGCGTTGGAACGGCTCAAAGCCC
TCCGCCGTGACGGCCGGAGATCCGACAGAGGATATGAGATCAAGGCTGAGAATCCGATCTACGACACCATGCCTGAGGAGGAGTACGCCAAGCTGGTCGC
TCGTCGGAAAGAGGAAGCACACGAGTTTATAGTCGACGACGAAGGGATCGGCTACGGCGACGAGGGAGAGGAAGAGGATTGGTCAAAGGCCGGTCTCCCA
ACTTCCTCCGACGAATCCGACGGCGGCGGTGGAGAACGCAGCAGCAGGCGGAAGAGAAAGATGGCGGAGAGGAAGGAGAAAGAGCCTCAGCAGAAGAAGG
TTCATCCTTCTTCTCTAACGGCTGCTGCAGCGATGATGGGTAAACAGAAGCTTTCGACCTACTTTAATTCGACCGCCATGTTCAGGAATCGTGATTCGGA
TAAATCAAAGAGTCTTGATTGCGATAGTATTGTGGATGACGTTATTGCCGAATTCGCACCTGATGAAGCTGATAGAGAAAAACGAAGGAGGGGACATCCA
GTAGTGGTGAGTAATGCTTCGCAAGTTAAGAGCGAGTATCTGGTTAGTGTAGAGAATAAGGTTTCTGTCAATGTCGGCGATCCTGTTATTGATTCTTTAG
AGAATAATGGCGGTGGTGATGAGGAAGAGAGACCGGATGTCGATGTAAAGGAGGTGGAGGAGGTACACGGAGTGAAAGATAAGGGATCTCTTGTCGGAGT
TGAATTGGAATCGGAGAATGTTCGTGAGGTGAAGTTGGAACCAGTGGTGAAGGTGGAGGAGGCAAGGAGAACTCTAAATGCGAAGATGAGTACAGAGAAA
AGAGATCCTAAATTAAGTGCCACAGCTGGATGGGAGGAAGTGAAAAGGAGTGGTGGAAATGGTGGCGTTGTGGAGGAGAATAACAGTGTGTCACGTAGTG
AAGAGCAACCAGAGTTTGAAGTGGATTCAGACGGGTCTTTACCATTTTACATACTTGACGCTCATGAAGAGGCGTTTGGAGCTAACCGCGGCACTGTTTT
TCTCTTTGGAAAGGTCAAAGCTGGGAATGTATACCATAGTTGCTGCGTGTTAGTAAAGAACATGCAGAGATGTGTTTATGCGATTCCCCACAATTCAATA
TTTCAAACTGATGAGGTGCTTCGGCTAGAGAAAAACGTTGAAGAGTCACGGATTTCCCTTGCAGAATTCCGTAAAAAGCTTCAGGATGTGGCATATGGAT
TAAAGGAGGAGGTGGCGAACCAACTAGTAAATCTCAAAGTGTCGAGTTTTAGCATGACTCCAGTCAAGAGAAAGTATGCGTTTGAGCGATCTGATGTTCC
TCTTGGGGAGAATTATGCACTTAAGATTAATTACTCATTCAAGGATTCGCCACTTCCTACAGACCTAAGAGGAGAGACCTTTACTACTTTGCTTGGAACT
AATACCAGTGCTTTGGAGCTTTTTCTTGTGAAAAGAAAGATTAAGGGACCCTCGTGGTTATTGATTCAAAAGTTCTCAACCCGCCAAACCTCTCAGAAGG
TGAGCTGGTGCAAGTTTGAAATCGTTGTTGACTTCCCAAAAGACATTCACACTGCATCATCATCAAAAATTACAGCAGAGATCCCCCCAGTGGTGGTTAC
AGCAATAAATTTGAAAACTATCTTAAATGAAAAGAAGAATGTGAACGAAATAGTTTCTGCATCTGTCGTCTGCTGTCACAAGGCCAAGCTTTTAGCCGGA
GCTGAAGGCATGCTACTAGTTTCTAAACCTCGGATTGAGTTGAATCCATTTTTTGGTTTTACTGTTCAGATTGATACTCCTATGCTGGCTCCAGAACGGG
AGAAACCTGGAATGGTCAGTCATTTTACTGTTGTCCGCAAACTTGATGGAGGTATATTTCCGATGGGTTTCTCGAAGGAGCTGGCTGATAGGAACATGAA
GGCTGGGTCTCGGTCCACTATCTTATCCATGGAGCCCAGTGAAAGGTCTTTGCTGAATCGTCTGATGATCGAGTTGCATAAGTTAGATAGCGACATTCTT
GTTGGACATAACATATCTGGGTTCGACTTGGACGTTCTCCTCCATAGAGCACAGGCTTGTCAAGTTCCAAGCAGCATGTGGTCCAAACTAGGCCGTCTGA
AGCGTTCTACGATGCCCAAGCTTTCAAAACCAAATGGTACCAGAGCTAGCCAGGGAATTTTGTCTTGTTTATCTGGTCGGCTTTTATGCGATACATATAT
ATGCTCCCGCGATCTACTGAAAGAGGTTAGCTATTCTTTGACACAACTTGCAAATACTCGACTGGGTAAACATCGGAAAGAGGTTGCTCCTCATGACATT
CCCAAAATGTTCCAGACCTCTGAATCTCTTATCGAACTAATTCAATTGGGTGAGACAGATGCATGGCTTTCTATGGAACTTATGTTTCACCTAAGTGTTC
TTCCACTCACACGTCAACTAACCAATATCAGTGGTAATCTTTGGGGGCAAACTTTCCAAGGTGCTAGGGCATTGAGAGTGGAGTATCTTCTATTGCATGC
ATTTCATTCAAAGAAATATATTGTGCCAGACAAACAATATTCCTCTCATGCAAAGGAAGCATACATTACTAAGCGAAAAATGCATCACAACACTGACAAT
ACGGTTACTGAAGAGTTGGAGGTTGAAGACATCGATTTTGAAAATGAACATCCAGAAAATTCGCATAGAAAAGGGAAGAAGGTTCCTGCCTATGCAGGTG
GCTTAGTTCTGGAGCCAAAGAAAGGCTTGTATGACAAATATGTATTGCTTTTGGATTTCAATAGTCTGTATCCTTCTATCATTCAGGAATACAATATCTG
CTTCACAACTGTGGAAAGATCTTCAGAGGGGGGAGTTTTTTCACGTCTACCTTCCAGCAAGACAACTGGAGTTTTGCCTGAGCTTTTGAAAAATTTAGTT
GAGAGAAGGAGAATGGTAAAGAAATGGATGAAGAGCGCATCTGGCCTTAAAGTCCAACAGCTTGACATTCAACAACAAGCACTGAAGCTAACTGCTAACA
GCATGTACGGATGTCTTGGATTTTCAAACTCAAGATTTTATGCTAAGCCTCTAGCAGAGCTTATTACTCTACAAGGAAGGGAGATTCTACAGAGTACTGT
TGATCTAGTTCAACACAATCTGAACTTTGAGGTTATCTATGGTGATACAGATTCGATCATGATTTACAGTGGACTAGATGATATAGCAAAAGCAAAAGCA
ATTGCCGGGAAGGTTATTCAGGAGGTGAACAAGAAGTACAGGTGTTTAGAAATAGATCTTGATGGCCTATACAAGAGAATGCTGCTACTGAAAAAGAAAA
AGTATGCAGCTGTGAAGATTCAGTTCAAGGATGGGACACCATACGAGGTTAGAGAGTGCAAGGGTATCGACATGGTTCGTCGTGACTGGAGCTTGCTATC
CAAGGAAGTGGGAGGTTTCTGTTTGGATCAGATCTTATCCGGGAGGTCGTGTGAAGATGTAGTGGAGTCGATACATAGTTATCTTATGAAGGTGCGGGAT
GAAATGAGGAGTGAACAAGTAGAACTTAAGCAGTATGTTATCACTAAGTCATTAACTAAACCACCTGAAGCATACCCAGATGCTAAAAACCAACCCCATG
TCATGGTGGCACTCAGATTGAAGCAAAGTGGTTACACAGCTGGTTGCTCAGCTCATGACACAGTCCCATACGTGATTTGCATCGAGCAGGGTGCTAGTTC
AGGAGGTTCAGCAGGGATCGCTCAGCGAGCAAGGCACCCTGATGAGTTGGAAAAAGATGATGGGAAATGGATGATTGACATTGATTATTACTTGTCTCAA
CAAATTCATCCAGTAGTTTCTCGTCTATGTGCTCCAATTCAGGGTACAAGTCCAGAGCGCTTGGCTGATTGCTTGGGGCTCGATTCGTCGAGGTTCCGGA
GTAGATCGAACGATGCTGCAGATAGTGATCTTTCCAGCTCACTTGTGTTTGCTGGGAATGATGAGGAGAGATACCAGAGTTGTGAACCATTGACCTTGTC
GTGTCCAAGCTGCTCCTCTACCTTTGACTGTCCTGCTATCTTCAGTTCCATTTACAAATCAATAAATGGAACAACGACACCAGAAGCGCAACCAGAAGAA
TCGACATCCAATTTCTGGCGCACACTGAGGTGCCCGAAATGTCCAGAAGAAGGGAATCTCGGCAGAGTATCTCCAGCCACGATAGCCAACCAGGTTAGAA
GGCAAGCAGAGAGTTATGTTTCCACTTACTACAAGGGCATAATGATGTGTGACGATGAAACGTGCAAGCATACGACGAGGAGTCTGAATCTGAGGCTAGT
TGGAGAATCAGAGAAAGGAACAGTTTGCCCAAACTATCCCAGGTGCAACGGCCGGCTTGTGAGGAAGTATTCGGAAGGAGAACTGTACAGGCAACTCTCG
TACTTCTGCTATTTGCTGGACACACAGCGCTGCCTTGAGAAGGAGCAGCAGCAGATGGAAGTGAGTAAGAGATTGGCACTGGAGGAAGAGCTGAGGAAAA
TTCGGGGAGTGGTGGAGATGGCGGTGTCGACGGTGGAAAAGATGAGAGATCGGTGTGCCTATGGATGGGTGCAGCTTAACGACTTCACTGTTACAGTATG
A
|
|||||||||||||||||||||||||
|
AA sequence
|
>Lus10009366 pacid=23148915 polypeptide=Lus10009366 locus=Lus10009366.g ID=Lus10009366.BGIv1.0 annot-version=v1.0
MEGEEQHPVVAGRRRSRGGGDTARAEALERLKALRRDGRRSDRGYEIKAENPIYDTMPEEEYAKLVARRKEEAHEFIVDDEGIGYGDEGEEEDWSKAGLP
TSSDESDGGGGERSSRRKRKMAERKEKEPQQKKVHPSSLTAAAAMMGKQKLSTYFNSTAMFRNRDSDKSKSLDCDSIVDDVIAEFAPDEADREKRRRGHP
VVVSNASQVKSEYLVSVENKVSVNVGDPVIDSLENNGGGDEEERPDVDVKEVEEVHGVKDKGSLVGVELESENVREVKLEPVVKVEEARRTLNAKMSTEK
RDPKLSATAGWEEVKRSGGNGGVVEENNSVSRSEEQPEFEVDSDGSLPFYILDAHEEAFGANRGTVFLFGKVKAGNVYHSCCVLVKNMQRCVYAIPHNSI
FQTDEVLRLEKNVEESRISLAEFRKKLQDVAYGLKEEVANQLVNLKVSSFSMTPVKRKYAFERSDVPLGENYALKINYSFKDSPLPTDLRGETFTTLLGT
NTSALELFLVKRKIKGPSWLLIQKFSTRQTSQKVSWCKFEIVVDFPKDIHTASSSKITAEIPPVVVTAINLKTILNEKKNVNEIVSASVVCCHKAKLLAG
AEGMLLVSKPRIELNPFFGFTVQIDTPMLAPEREKPGMVSHFTVVRKLDGGIFPMGFSKELADRNMKAGSRSTILSMEPSERSLLNRLMIELHKLDSDIL
VGHNISGFDLDVLLHRAQACQVPSSMWSKLGRLKRSTMPKLSKPNGTRASQGILSCLSGRLLCDTYICSRDLLKEVSYSLTQLANTRLGKHRKEVAPHDI
PKMFQTSESLIELIQLGETDAWLSMELMFHLSVLPLTRQLTNISGNLWGQTFQGARALRVEYLLLHAFHSKKYIVPDKQYSSHAKEAYITKRKMHHNTDN
TVTEELEVEDIDFENEHPENSHRKGKKVPAYAGGLVLEPKKGLYDKYVLLLDFNSLYPSIIQEYNICFTTVERSSEGGVFSRLPSSKTTGVLPELLKNLV
ERRRMVKKWMKSASGLKVQQLDIQQQALKLTANSMYGCLGFSNSRFYAKPLAELITLQGREILQSTVDLVQHNLNFEVIYGDTDSIMIYSGLDDIAKAKA
IAGKVIQEVNKKYRCLEIDLDGLYKRMLLLKKKKYAAVKIQFKDGTPYEVRECKGIDMVRRDWSLLSKEVGGFCLDQILSGRSCEDVVESIHSYLMKVRD
EMRSEQVELKQYVITKSLTKPPEAYPDAKNQPHVMVALRLKQSGYTAGCSAHDTVPYVICIEQGASSGGSAGIAQRARHPDELEKDDGKWMIDIDYYLSQ
QIHPVVSRLCAPIQGTSPERLADCLGLDSSRFRSRSNDAADSDLSSSLVFAGNDEERYQSCEPLTLSCPSCSSTFDCPAIFSSIYKSINGTTTPEAQPEE
STSNFWRTLRCPKCPEEGNLGRVSPATIANQVRRQAESYVSTYYKGIMMCDDETCKHTTRSLNLRLVGESEKGTVCPNYPRCNGRLVRKYSEGELYRQLS
YFCYLLDTQRCLEKEQQQMEVSKRLALEEELRKIRGVVEMAVSTVEKMRDRCAYGWVQLNDFTVTV
|
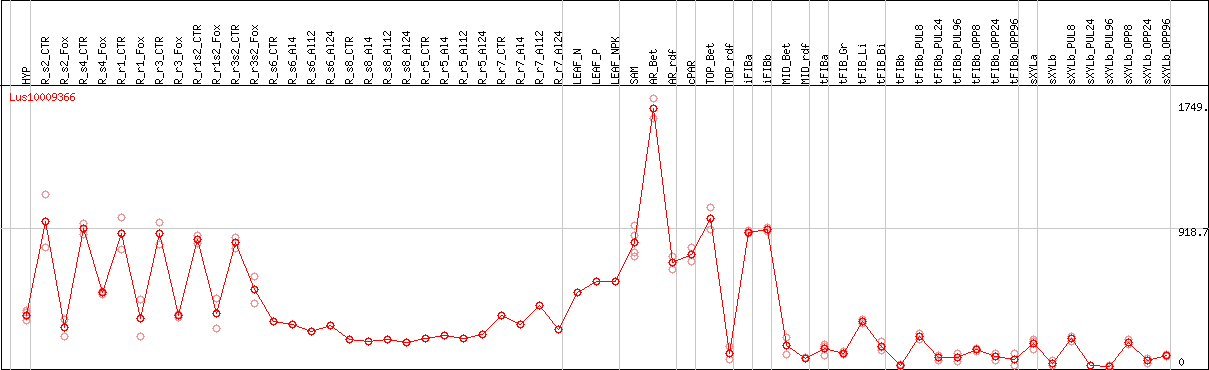
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009366 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
