External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G34430 310 / 2e-105
EMB3003
embryo defective 3003, 2-oxoacid dehydrogenases acyltransferase family protein (.1)
AT3G25860 267 / 1e-88
PLE2, LTA2
PLASTID E2 SUBUNIT OF PYRUVATE DECARBOXYLASE, 2-oxoacid dehydrogenases acyltransferase family protein (.1)
AT3G13930 101 / 3e-25
Dihydrolipoamide acetyltransferase, long form protein (.1)
AT1G54220 99 / 2e-24
Dihydrolipoamide acetyltransferase, long form protein (.1.2)
AT3G52200 97 / 1e-23
LTA3
Dihydrolipoamide acetyltransferase, long form protein (.1.2)
AT4G26910 82 / 9e-19
Dihydrolipoamide succinyltransferase (.1.2.3)
AT5G55070 82 / 1e-18
Dihydrolipoamide succinyltransferase (.1)
AT3G06850 77 / 9e-17
DIN3, LTA1, BCE2
DARK INDUCIBLE 3, 2-oxoacid dehydrogenases acyltransferase family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006069
323 / 1e-114
AT1G34430 377 / 3e-131
embryo defective 3003, 2-oxoacid dehydrogenases acyltransferase family protein (.1)
Lus10041467
273 / 1e-91
AT3G25860 595 / 0.0
PLASTID E2 SUBUNIT OF PYRUVATE DECARBOXYLASE, 2-oxoacid dehydrogenases acyltransferase family protein (.1)
Lus10034304
259 / 1e-85
AT3G25860 592 / 0.0
PLASTID E2 SUBUNIT OF PYRUVATE DECARBOXYLASE, 2-oxoacid dehydrogenases acyltransferase family protein (.1)
Lus10006877
106 / 3e-27
AT1G54220 704 / 0.0
Dihydrolipoamide acetyltransferase, long form protein (.1.2)
Lus10037617
103 / 5e-26
AT1G54220 716 / 0.0
Dihydrolipoamide acetyltransferase, long form protein (.1.2)
Lus10008412
99 / 3e-24
AT3G52200 725 / 0.0
Dihydrolipoamide acetyltransferase, long form protein (.1.2)
Lus10001967
98 / 5e-24
AT3G52200 717 / 0.0
Dihydrolipoamide acetyltransferase, long form protein (.1.2)
Lus10043116
83 / 9e-19
AT4G26910 638 / 0.0
Dihydrolipoamide succinyltransferase (.1.2.3)
Lus10032632
83 / 1e-18
AT4G26910 560 / 0.0
Dihydrolipoamide succinyltransferase (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G114300
319 / 5e-109
AT1G34430 549 / 0.0
embryo defective 3003, 2-oxoacid dehydrogenases acyltransferase family protein (.1)
Potri.019G084900
316 / 9e-108
AT1G34430 473 / 7e-165
embryo defective 3003, 2-oxoacid dehydrogenases acyltransferase family protein (.1)
Potri.010G126600
273 / 4e-91
AT3G25860 470 / 2e-163
PLASTID E2 SUBUNIT OF PYRUVATE DECARBOXYLASE, 2-oxoacid dehydrogenases acyltransferase family protein (.1)
Potri.008G119500
270 / 6e-90
AT3G25860 464 / 5e-161
PLASTID E2 SUBUNIT OF PYRUVATE DECARBOXYLASE, 2-oxoacid dehydrogenases acyltransferase family protein (.1)
Potri.001G198000
105 / 1e-26
AT3G13930 724 / 0.0
Dihydrolipoamide acetyltransferase, long form protein (.1)
Potri.003G043900
103 / 3e-26
AT3G13930 771 / 0.0
Dihydrolipoamide acetyltransferase, long form protein (.1)
Potri.008G027400
98 / 5e-24
AT3G52200 754 / 0.0
Dihydrolipoamide acetyltransferase, long form protein (.1.2)
Potri.014G154700
83 / 6e-19
AT4G26910 520 / 0.0
Dihydrolipoamide succinyltransferase (.1.2.3)
Potri.011G089200
83 / 8e-19
AT5G55070 562 / 0.0
Dihydrolipoamide succinyltransferase (.1)
Potri.001G357000
82 / 2e-18
AT4G26910 624 / 0.0
Dihydrolipoamide succinyltransferase (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0149
CoA-acyltrans
PF00198
2-oxoacid_dh
2-oxoacid dehydrogenases acyltransferase (catalytic domain)
Representative CDS sequence
>Lus10009432 pacid=23149680 polypeptide=Lus10009432 locus=Lus10009432.g ID=Lus10009432.BGIv1.0 annot-version=v1.0
ATGACAGCCCTGCTAGCCAAGGCGACAGCTCTTGCATTAGCCAAACACCCCGTGGTGAATTCCAGCTGCAGAGACGGCAATAGCTTTACGTACAATAGCA
ACATCAACATTGCTGTCGCTGTAGCCATCGATGGCGGGTTGATCACCCCGGTTCTTCAGGATGCCAATAAGGTCGATCTTTATTCATTGTCGAGAAAGTG
GAAGGAGTTGGTCGATAAGGCCAGAGCCAAGCAGTTGCAGCCTCAGGAGTACAACACAGGTACCTTTACCCTGTCGAACCTAGGAATGTTCGGAGTGGAC
CGATTTGATGCCATTCTTCCACCCGGCACGGGAGCTATCATGGCTGTTGGGGCATCTCAGCCGACAGCAGTTGGTACGAAGGATGGCCGGATCGGCATCA
AGAACCAGATGCAGGTGAACGTGACAGCGGACCATCGAGTCATTTACGGAGCCGATCTAGCTGCCTTCCTGCAAACCTTGGCACAAATCATCGAGGATCC
GAAAGATCTTACCTTCTAG
AA sequence
>Lus10009432 pacid=23149680 polypeptide=Lus10009432 locus=Lus10009432.g ID=Lus10009432.BGIv1.0 annot-version=v1.0
MTALLAKATALALAKHPVVNSSCRDGNSFTYNSNINIAVAVAIDGGLITPVLQDANKVDLYSLSRKWKELVDKARAKQLQPQEYNTGTFTLSNLGMFGVD
RFDAILPPGTGAIMAVGASQPTAVGTKDGRIGIKNQMQVNVTADHRVIYGADLAAFLQTLAQIIEDPKDLTF
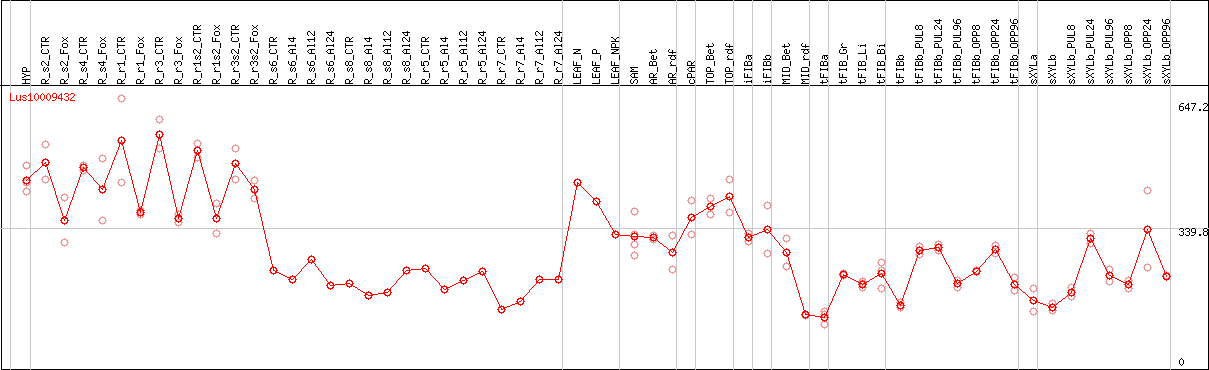
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009432 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.