Lus10009474 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10009474 pacid=23149690 polypeptide=Lus10009474 locus=Lus10009474.g ID=Lus10009474.BGIv1.0 annot-version=v1.0
ATGGCGGCGGCAGAATCTGCTGCGGCTGCTCTTGGGCCAGCTTATGCACCAGAAGACCCTACACTGCCTAAGCCATGGACAGGATTGATTGATAGGAGCA
CTGGTATGTTATACTACTGGAATCCTGAGACCAATGTTACGCAATATGAAAAGCCAGCTTCTGCTTTGCCTCCTCCTGCAACTGGCTCGACACCTAAGTT
GGCTCAGATACCAATGCCTCAGTCTGCTCAACAAACTAATGGGATGGTGGGAGGTCATGGTGCTCAACAACAGCAGGGGCAACAATTCAGGCAAATGCAT
TATCAGGGCCAACAAACACAAGGGCAGCAGCAGATGAGTGCCTTGTCTCAACAGCCTCAGCAAGGGCAAGGCTTAGGACAGCCACAACAACATTCTTCGG
TGCAGCATGTATCTAGCCAGCCGGGTTCACAACAACAGGGAATGCAAATGGGGCAGGTTGGTCAACAACAAGGGCAGTTTCGACCCCAAATGATGCAGCA
GAATCCCGGTCAGCATATGTTTTCACATACGGGACAGCAAATGCATCAACCTTCAGGCCAGCCTATGCCACAGCAGTCAAGCCAGCCGATGCCGCAACAA
CCAGGGCAGCCCATGCAGCAGCAGCCAGGGCCAGGCCAGCCTATGGCGCAGATGACACAAACACAAGGACAGTTGTCCTATCAGCAAATGCAGTACATGC
CTTACCAACAAAATGTGGGTCCCCCAGTGCACCAGCAGTATGCTACCCAAATGAAATCTGCAATGGCTCGGCCAGAGGATGGTGATCTGCCACAGGGTGG
CCAGGCTGGGTACCCTACATCTCAGTTTCATCATCCCACTGGTGGATCCTCAGCTCAAAACATGCCAATTGGGAACAAATCCCCGTTGATGCCTCCTGCA
AGTGGTCACTTCAATCGAGGTCAGCAAGTCACTGGACCCTCCGGGCCTGATTTAGGACATAAACCACCGATTCAAAGGCTTCAGAATCTCACAGGTCCGC
CTGGTGCACATAATCAACAACCCAATGTGCCTCCTGTTGGATTGAAGATGGGCCACGAAAATGCTCCTCCTCCTAGGTCGGGCAATGACTACTACCCGAA
TCCAAAAATTGGAGGCTCCATGATGAGTCCAAACCAACCTAATGGTGCTCCTATGCCTATGGCAAGGAATCAGCAGGATGGAAGAATGGGTGGAGCCCCA
TTTCCAAATTCCGGACCTGCTTATTCAGGTGGGCTTAGCTCACCTGGGCAGGGAATGCACAATGCGTATAACCATGGAAATATCAGCTCACAGTTTCCTG
CTAGTGGGATGATGATGCCTCCATTCAGCGGCCCTGCAGACATGTCTAACCTCTCGCCTGTTGATGCTTATCGCCAAAAGCATGAAGTTTCAACAATGGG
TGACAACGTTCCTGCTCCTTTCATGACATTTGAAGACACTGGTTTCCCTCCAGAGATATTGAGAGATATGGGTGGTTGGCCTTTGACCAATTCACACCTG
ATATGCTCCTCATTTGTCTTTATTGGCCTCTCTTTAGTTGCTTCTATACATTCTGCTGGTTTCTTGTTTCCCACGCCAATCCAGGCACAAACATGGCCGA
TAGCACTACAGAGCCGAGATATAGTTGCCATTGCAAAAACTGGTTCTGGGAAAACTCTTGGTTACTTAATCCCGGCATTTATTCTTCTAAGGCAGCGTCG
GAACAATCCTCAGAATGGTCCCACAGTATTGGTTTTAGCTCCAACTAGAGAGCTTGCCACACAAATCCAAGAGGAGGTGAACAAATTTGGTCGATCTTTT
AGGGTGTCTTGCACGTTGCTTGAGCATTACATGATCCATCCATTGAACTTGCAGTGCCTCTATGGCGGTGCACCGAAGGGTCCTCAATTGAATGAACTAG
AACGCGGCGCTGATGTGGTTGTGGCAACTCCTGGTCGTCTTAACGACATCCTTGAAATGAAGAGGATTGATTTCCGTCAAATTTCGCTCCTTGTTCTTGA
CGAGGCAGATAGAATGCTTGATATGGGTTTCGAGCCTCAAATACGTAAGATTGTAAATGAGATACCCCCTAGGAGGCAAACACTTATGTACACTGCAACT
TGGCCAAAAGAAGTGAAGAAGATAGCAGGTGATCTTCTTGTCAACCCTGTTCAAGTCAACATTGGAAATGTTGATGAACTTGCTGCAAACAAATCCATCA
CTCAGTTTGTGGAAATGGTACCCCATATGGAGAAGGAGAAGCGTTTAGACCAGATCTTAAGGTCACAAGAGAGAGGTTCAAAAGTCATTATCTTCTGTTC
CACTAAGAGATTGTGTGACCAGCTAGCTAGAGGTATTTTGCGCAATTTTGGGGCCAACGCAATCCACGGGGACAAGTCTCAAGGTGAAAGAGACTGGGTT
TTGAACCAGTTCAGGAGTGGGAAGTCACCAATCTTAGTCGCCACTGATGTTGCTGCTCGTGGCCTCGACATTAAGGACATACGGGTTGTGGTTAATTATG
ATTTCCCTACTGCGATCGAGGACTACGTGCATCGAATTGGCAGAACCGGAAGGGCAGGTGCGACTGGTGTGGCGTACACATTCTTCTCTGAGCAGGACTG
GAAGCATGCACCGGATTTGGTCAAAGTTTTGGAGGGAGCTAACCAGCAGGTGCCTCCAGCAGTTCGGGAGATTGCTTCGCGTGGTGGTCCTGGTTCTGGT
TTCATGAAAGACCGAGGTCCGATGAATCGTTTCGATCCCGGTCCTGGAGCCGTCGGTGGAGGTGGTCGCTGGGACGGTGGAGGCCGCGGTGGTGGGATGA
GGGACGGTGGATACGGTGGCCGCGGTGGAATGTTGAGGGACGGAGGATTCGGTGGCCGCGGTGGAATGGGGAGGGACGGAGGGTTTGGTGGACGAGGTGG
AATGCGCGATGACGGATTCGGTGGACGTGGTGGAATGCGCGATGAAGGAGGATTCGGTGGAAGAGGTGGCCGAAACGACATGTTTGCTGGTCGTGGTGGG
GGGAATAGGGGTCGAGGAGGTTTCGGTGGGCCTGGCGGTTGGGGTAGAAACGAGCGGGGTCCACAAGATCGTTACGACAACCGAGGAGGAGGACGCGGCG
GTCGTGGCGGAGGAAGATTCGATGGGAGGAGAGATATGAGCAGCGGCAGGAGTAGGGGTAGAAGCCCGAGCCGTAGCCGAAGTAGAAGTTGGAGTAGGAG
CAGGAGCCGGAGTAGGAGTTGGTCCAGGGACCGGAGTCGAAGCTACAGTAGGAGTCGAAGCCCGAAAAGAAACCGTAGCTACAGCAGGAGTCCGCGCCTT
CGCAGTAGTCGTAGCTATAGCCGTAGCCCTGATGGAGGAAGCCGTAGCCCTACCCCCGAAAGATTCGTTAGGCGTCAAAAGCAACGAGAGTCCCTGTTCG
ATGTCAAGGATGTCCCAACGGCAGCTTCTGAACCAATGCTCTCTAACCCAACTACGGATCCTGTCGTGAACTCTGTTTCGGAGATGGAACAAGCAGCCGT
TGAGAATGCTGGACCACCACCGGCTGACGGGGAGGGACCTGAATCGGCAGCCGAAACTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10009474 pacid=23149690 polypeptide=Lus10009474 locus=Lus10009474.g ID=Lus10009474.BGIv1.0 annot-version=v1.0
MAAAESAAAALGPAYAPEDPTLPKPWTGLIDRSTGMLYYWNPETNVTQYEKPASALPPPATGSTPKLAQIPMPQSAQQTNGMVGGHGAQQQQGQQFRQMH
YQGQQTQGQQQMSALSQQPQQGQGLGQPQQHSSVQHVSSQPGSQQQGMQMGQVGQQQGQFRPQMMQQNPGQHMFSHTGQQMHQPSGQPMPQQSSQPMPQQ
PGQPMQQQPGPGQPMAQMTQTQGQLSYQQMQYMPYQQNVGPPVHQQYATQMKSAMARPEDGDLPQGGQAGYPTSQFHHPTGGSSAQNMPIGNKSPLMPPA
SGHFNRGQQVTGPSGPDLGHKPPIQRLQNLTGPPGAHNQQPNVPPVGLKMGHENAPPPRSGNDYYPNPKIGGSMMSPNQPNGAPMPMARNQQDGRMGGAP
FPNSGPAYSGGLSSPGQGMHNAYNHGNISSQFPASGMMMPPFSGPADMSNLSPVDAYRQKHEVSTMGDNVPAPFMTFEDTGFPPEILRDMGGWPLTNSHL
ICSSFVFIGLSLVASIHSAGFLFPTPIQAQTWPIALQSRDIVAIAKTGSGKTLGYLIPAFILLRQRRNNPQNGPTVLVLAPTRELATQIQEEVNKFGRSF
RVSCTLLEHYMIHPLNLQCLYGGAPKGPQLNELERGADVVVATPGRLNDILEMKRIDFRQISLLVLDEADRMLDMGFEPQIRKIVNEIPPRRQTLMYTAT
WPKEVKKIAGDLLVNPVQVNIGNVDELAANKSITQFVEMVPHMEKEKRLDQILRSQERGSKVIIFCSTKRLCDQLARGILRNFGANAIHGDKSQGERDWV
LNQFRSGKSPILVATDVAARGLDIKDIRVVVNYDFPTAIEDYVHRIGRTGRAGATGVAYTFFSEQDWKHAPDLVKVLEGANQQVPPAVREIASRGGPGSG
FMKDRGPMNRFDPGPGAVGGGGRWDGGGRGGGMRDGGYGGRGGMLRDGGFGGRGGMGRDGGFGGRGGMRDDGFGGRGGMRDEGGFGGRGGRNDMFAGRGG
GNRGRGGFGGPGGWGRNERGPQDRYDNRGGGRGGRGGGRFDGRRDMSSGRSRGRSPSRSRSRSWSRSRSRSRSWSRDRSRSYSRSRSPKRNRSYSRSPRL
RSSRSYSRSPDGGSRSPTPERFVRRQKQRESLFDVKDVPTAASEPMLSNPTTDPVVNSVSEMEQAAVENAGPPPADGEGPESAAET
|
DESeq2's median of ratios [FLAX]
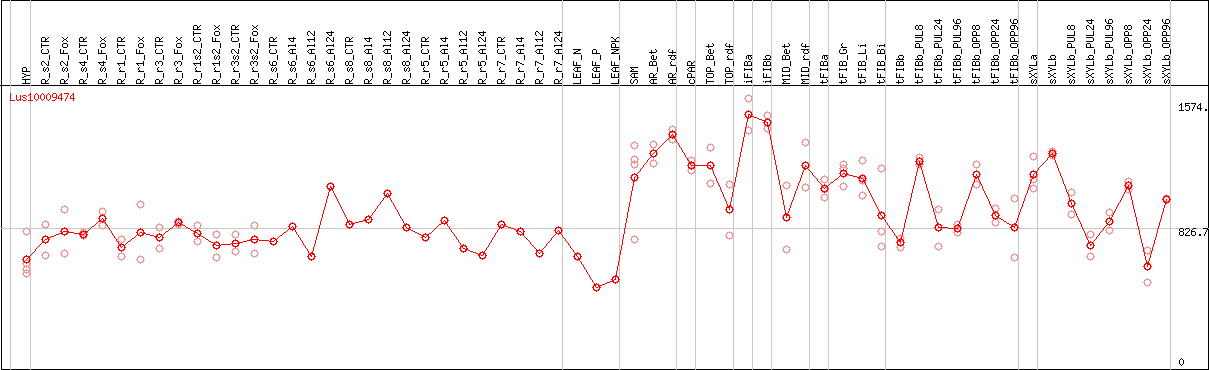
Coexpressed genes
Lus10009474 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
