External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G72940 107 / 6e-28
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT1G72920 104 / 2e-27
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
AT5G17680 106 / 7e-27
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT1G72930 99 / 1e-26
TIR
toll/interleukin-1 receptor-like (.1.2)
AT5G36930 105 / 2e-26
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT1G72900 100 / 2e-25
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT1G72910 100 / 4e-25
Toll-Interleukin-Resistance (TIR) domain-containing protein (.1)
AT1G17600 100 / 1e-24
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G17615 98 / 2e-24
Disease resistance protein (TIR-NBS class) (.1)
AT5G18360 97 / 1e-23
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030498
155 / 8e-48
AT5G36930 115 / 2e-29
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10012852
136 / 9e-40
AT5G17680 100 / 1e-23
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10012063
126 / 6e-36
AT5G36930 124 / 5e-34
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10026845
124 / 7e-35
AT5G36930 145 / 8e-39
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10029921
125 / 1e-34
AT5G36930 105 / 7e-26
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10041606
122 / 4e-34
AT5G36930 140 / 5e-37
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10006929
107 / 8e-29
AT1G72890 146 / 1e-39
Disease resistance protein (TIR-NBS class) (.1), Disease resistance protein (TIR-NBS class) (.2)
Lus10027920
109 / 9e-29
AT5G36930 139 / 3e-36
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10029919
106 / 1e-28
AT5G36930 105 / 5e-26
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G004233
112 / 1e-30
AT1G27170 153 / 8e-42
transmembrane receptors;ATP binding (.1.2)
Potri.005G003900
108 / 2e-29
AT5G36930 146 / 8e-40
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.004G230000
108 / 4e-29
AT5G36930 169 / 4e-47
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.005G004500
105 / 2e-28
AT5G36930 152 / 3e-42
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G070436
104 / 2e-28
AT2G20142 160 / 7e-49
Toll-Interleukin-Resistance (TIR) domain family protein (.1)
Potri.019G016425
106 / 1e-26
AT5G36930 650 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G007884
105 / 2e-26
AT5G36930 654 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.006G269950
100 / 3e-26
AT5G36930 147 / 3e-40
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G069866
99 / 4e-26
AT4G12010 179 / 4e-52
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Potri.019G068300
104 / 5e-26
AT5G17680 580 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0173
STIR
PF01582
TIR
TIR domain
Representative CDS sequence
>Lus10009500 pacid=23147307 polypeptide=Lus10009500 locus=Lus10009500.g ID=Lus10009500.BGIv1.0 annot-version=v1.0
ATGAAAATTAAAATCAAACAACATGAATTAACAGTGGGGAACCACCGCCGCCGCCGCCATGACGTATTCGTAAGCTTCCGAGGCATAGACGTCCGCTACA
AATTTGTGGATCATCTCTTCTCCGCCTTCCGGAGGAAGAAAGTGAAGGCGTTCAGAGATACTAGCGACTCAAGCAGAGGACAACTCATCGACGAGGAGAT
CCCACTGGCAATTGACGAGTCGAGCTTTTACGTTGTGGTTCTATCCGAAAACTACGATTCGTCTCCATGGTGCTTGGATGAGCTCGTCATGATCATGGAC
AACTCCCGCGGTGGGAAGATGGTATTCCCCATATTCTACCATGTGTTGCCGGACGATGTCTCCTCCGTGGACGCCGTTCACCGGATCAAAGGTCAGTACA
CGTACGAAAGAGTGGAGAGATGGATTGAGGCGCTGGATTGGATCGCGAGGATTGCTGGCTGGGTCGTCACTGTCAGAGGCGTCGGTGGTGGAGAGTATTG
CCCGAATCATCCAACGCAGGGTTCGAAGGAAGGGGAAGCGCATCCGGATGAATCGAGGCACTAA
AA sequence
>Lus10009500 pacid=23147307 polypeptide=Lus10009500 locus=Lus10009500.g ID=Lus10009500.BGIv1.0 annot-version=v1.0
MKIKIKQHELTVGNHRRRRHDVFVSFRGIDVRYKFVDHLFSAFRRKKVKAFRDTSDSSRGQLIDEEIPLAIDESSFYVVVLSENYDSSPWCLDELVMIMD
NSRGGKMVFPIFYHVLPDDVSSVDAVHRIKGQYTYERVERWIEALDWIARIAGWVVTVRGVGGGEYCPNHPTQGSKEGEAHPDESRH
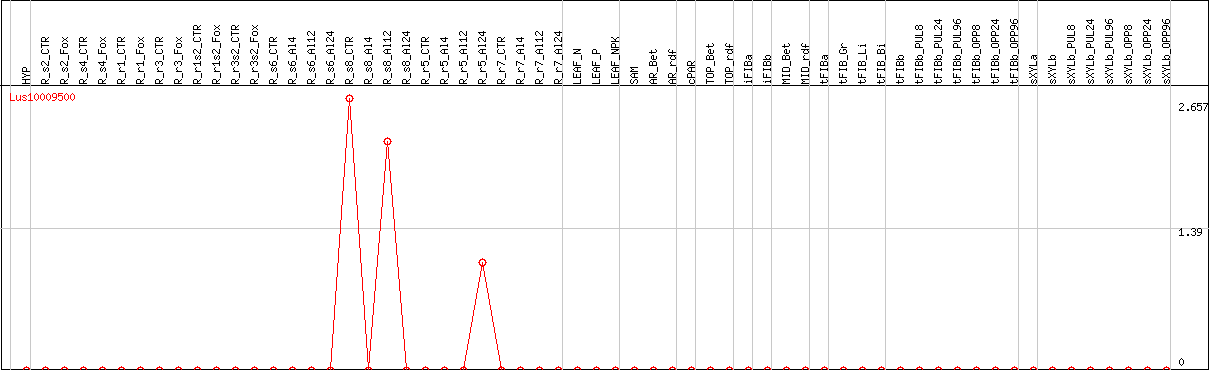
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.