External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G22430 97 / 1e-24
GroES-like zinc-binding dehydrogenase family protein (.1.2)
AT1G22440 94 / 1e-23
Zinc-binding alcohol dehydrogenase family protein (.1)
AT5G24760 93 / 2e-23
GroES-like zinc-binding dehydrogenase family protein (.1.2.3)
AT4G22110 89 / 8e-22
GroES-like zinc-binding dehydrogenase family protein (.1.2)
AT5G43940 88 / 1e-21
PAR2, ATGSNOR1, HOT5, GSNOR, ADH2
PARAQUAT RESISTANT 2, sensitive to hot temperatures 5, S-NITROSOGLUTATHIONE REDUCTASE, ALCOHOL DEHYDROGENASE 2, GroES-like zinc-binding dehydrogenase family protein (.1.2)
AT1G32780 88 / 2e-21
GroES-like zinc-binding dehydrogenase family protein (.1)
AT1G77120 82 / 2e-19
ADH1, ATADH, ATADH1
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
AT1G64710 82 / 3e-19
GroES-like zinc-binding dehydrogenase family protein (.1.2)
AT5G42250 79 / 2e-18
Zinc-binding alcohol dehydrogenase family protein (.1)
AT2G21730 58 / 8e-11
CAD2, ATCAD2
cinnamyl alcohol dehydrogenase homolog 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005656
251 / 3e-86
AT1G22430 155 / 4e-45
GroES-like zinc-binding dehydrogenase family protein (.1.2)
Lus10042787
87 / 3e-21
AT1G77120 674 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Lus10029759
87 / 4e-21
AT1G77120 673 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Lus10010506
87 / 5e-21
AT1G77120 673 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Lus10034042
87 / 5e-21
AT1G77120 672 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Lus10022853
87 / 8e-21
AT5G43940 700 / 0.0
PARAQUAT RESISTANT 2, sensitive to hot temperatures 5, S-NITROSOGLUTATHIONE REDUCTASE, ALCOHOL DEHYDROGENASE 2, GroES-like zinc-binding dehydrogenase family protein (.1.2)
Lus10033241
82 / 3e-19
AT1G32780 511 / 0.0
GroES-like zinc-binding dehydrogenase family protein (.1)
Lus10023572
79 / 5e-18
AT5G42250 439 / 1e-152
Zinc-binding alcohol dehydrogenase family protein (.1)
Lus10040457
78 / 1e-17
AT5G42250 488 / 7e-173
Zinc-binding alcohol dehydrogenase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G142700
110 / 5e-30
AT1G22430 295 / 3e-97
GroES-like zinc-binding dehydrogenase family protein (.1.2)
Potri.018G142750
110 / 8e-30
AT1G22430 299 / 4e-99
GroES-like zinc-binding dehydrogenase family protein (.1.2)
Potri.019G041500
97 / 7e-25
AT1G22430 308 / 1e-102
GroES-like zinc-binding dehydrogenase family protein (.1.2)
Potri.019G041600
96 / 2e-24
AT1G22430 300 / 3e-99
GroES-like zinc-binding dehydrogenase family protein (.1.2)
Potri.002G254900
89 / 1e-21
AT5G43940 694 / 0.0
PARAQUAT RESISTANT 2, sensitive to hot temperatures 5, S-NITROSOGLUTATHIONE REDUCTASE, ALCOHOL DEHYDROGENASE 2, GroES-like zinc-binding dehydrogenase family protein (.1.2)
Potri.002G072100
86 / 7e-21
AT1G77120 645 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.014G193800
86 / 1e-20
AT5G43940 684 / 0.0
PARAQUAT RESISTANT 2, sensitive to hot temperatures 5, S-NITROSOGLUTATHIONE REDUCTASE, ALCOHOL DEHYDROGENASE 2, GroES-like zinc-binding dehydrogenase family protein (.1.2)
Potri.007G108500
86 / 1e-20
AT1G77120 626 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.007G108401
86 / 2e-20
AT1G77120 630 / 0.0
ARABIDOPSIS THALIANA ALCOHOL DEHYDROGENASE, alcohol dehydrogenase 1 (.1)
Potri.004G067000
86 / 2e-20
AT5G24760 616 / 0.0
GroES-like zinc-binding dehydrogenase family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0296
GroES
PF08240
ADH_N
Alcohol dehydrogenase GroES-like domain
Representative CDS sequence
>Lus10009516 pacid=23145708 polypeptide=Lus10009516 locus=Lus10009516.g ID=Lus10009516.BGIv1.0 annot-version=v1.0
ATGGCGTCTGTCAGCTTTGCTCGCCCTGACAAGAATGGAGTTATCACTTGCAAGGCAATAGTGCTTAAGGAGGCGAAGAAGCCAGGGATGTCATATGCTG
ATACCGTCGAGATAAAAGACATCCAAGTCGACCCGCCGCAAAATGTCGAGCTTAGGGTTATGATGCTATGTGCTAGTGTGTGCCGCACCGATATTTTAAC
CATTGAAGGCTTCATGGCTCCGACTCAATTCCCTAAAGTCAATGGGCATGAAGGTGTTGGGATAATCGAGAGCATGGGCCCGGACACGAAGAACTTCAAA
GTGGGTGACGTCATCGTGGCTCCGACATTAGGAGAGTGCCAGACCTGCAGCAGCTGCAGGTCCGGCCGAACCAACTTCTGCCAGAACTAA
AA sequence
>Lus10009516 pacid=23145708 polypeptide=Lus10009516 locus=Lus10009516.g ID=Lus10009516.BGIv1.0 annot-version=v1.0
MASVSFARPDKNGVITCKAIVLKEAKKPGMSYADTVEIKDIQVDPPQNVELRVMMLCASVCRTDILTIEGFMAPTQFPKVNGHEGVGIIESMGPDTKNFK
VGDVIVAPTLGECQTCSSCRSGRTNFCQN
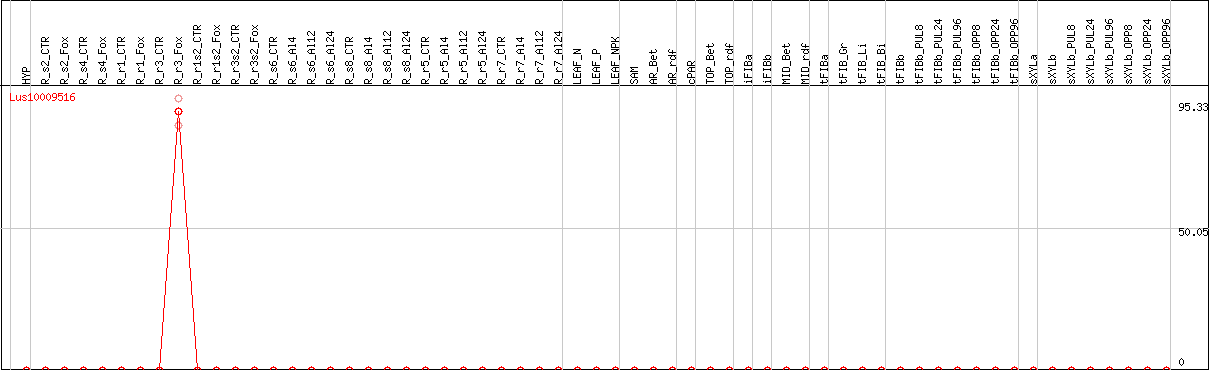
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009516 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.