Lus10009530 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10009530 pacid=23145683 polypeptide=Lus10009530 locus=Lus10009530.g ID=Lus10009530.BGIv1.0 annot-version=v1.0
ATGCATCCGAAGGCTCAAACCAGAGTGGAATCTGGAACGTGTAACGTGTGCTCCACTCCTTGTTCATCTTGTATGCACCGGAAGCGAGCTTTTATGGGAT
CAAAGGGTGATGAGGGTGATGAATTTTCCGACGAAACGTGCCACGTAGCTGAAACAAGTCAGAATTCCATTAACAAGGATGATTCCTCACCTATCAAAGG
CGGCATGGCTGCCATGTTACGACATGCCACAAGTGCAGCTAGTAATGTGCCAACTGTTGATTCATGTCAGGATTCAGTGTCTGAAAGTGCAGAAATTAAA
GGTAATCTGAAGATTTCGGATAGAGTGGACGTCTCAATGAAGTCTGAGGCTCTCCCAGAGTCCTCACACGTTGGATCTGTCTCGGTGGTGGACCGGATTT
CTCCTAAGCCAGAGCTTATTGCATATAAGCGAGCTGCGTCTGTTAATCCTGCTGCTTGTAAAGCTGTAGAAGGCCATGGTGATAATATATCATGTGTGAC
GGGTGCTGATGTGAGCAGTAACGGAGATTCTGCCAGTAAGCTGCCTGTCCAGAATAACAATCTCTCGTCCAACGTAGATGTTGTAAACTTGAACAAAGAT
CCAGTCGATCAAGTTTATAGAGAAAGTTCTAAACCCATAAAGGAACACACAAGCCCAATGTTGTCACAAGAGGCAGCACCAGATATTCTACTCCGGGATA
ACTCGCAAAATGGTGCAGACAACAACCTGCAAAGTAGAAGTGTAACTCCGGGTGATGGTCCAGAGGTTTCAAGTAGACTTGGTCAAGAGATCAAAGAAGA
AACTGGTAATGATAGTCGATCACGTCCAGACAAAGGTAATAGTTCTTCTGTCAAAGGCGAGGAAGATGAGAAGGTTAATGAGTCAGTTGAGCTACCTCAT
ACTCGTGAGTCGCCTGCACAATCTTCATCTGCAAATGATGGTGATGACTCGGAAATTGTGGAACAAGACGTGAAAGTATGTGATATCTGTGGGGACTCAG
GTCAGGAAGAGAAGCTTGCATTTTGTAGTAGGTGCAGCGATGGTGCTGAACACACATATTGCATGAAGGAAATGCTCCTAGAAGTTCCGGATGGTGATTG
GTTGTGTGAAGAATGCAAGATAGCTGAAGATTCCGAAAATCAGAAGCAAGGCGAGGAAGATGAGAAGGTTAACGAGTCAGTTGAGCTACCTCATACTCGT
GAGTTGCCTGAACAATCTTCATCTGCCAATGATGGTGATGAGTCGGAAATTGTGGAACAAGACGTGAAAGTATGTGATATCTGTGGGGACTCAGGTCAGG
AAGAGAAGCTTGCATTTTGTAGTAGGTGCAGCGATGGTGCTGAACACACATATTGCATGAAGGAAATGCTCCTAGAAGTTCCGGATGGTGATTGGTTGTG
TGAAGAATGCAAGATAGCTGAAGATTCCGAAAATCAGAAGCAAGGTGCAGAGGAAAAAGATGCAAGCAAATTAACTTATCAGAGGTTGAAGAAGAGACCT
GCTGAGACTGTAGAGGTGGCCGCTATTTCCAAAAGGCAGGCTGTTGACATGGATTCAGGATCGTCGAATCCATCGAGCCCTATCAAAGTCGCTGCCCTGC
AGCGTAACAGCTCCTCCAGGAGTTGGGATAAGGAGAAAGTAAAGCCAAGCCATCCTGTTTATGACATTACAGAAAGTACACGATTTCCTTCAACTGGTTC
ACGACTTCAAAGTCACAAGGGTACATATCTGAAGTCCAATTCCTTCAACGCAATAAGTTCCAAGTCGAAAGTTAAGCTTGTTGATGAAGTGCCACGCAAG
CACAAGAGCTTTAAGGAGAGCTCCTTATCTGATAATAAGGATGGGCCTGGCAAAACAATGGGCAAATCTATGTCATTTCATTACGCAGGGTCTGGAAAAG
CTGAGTCAAAAGTGAAAATGTTACCGTCAAAACTTGATAGTAAAGGATTAAAGCAAGCACGAGACTATAACACCTTTGAGAGAAGAAGTTTGCCTAAGTT
AGACCGTCCTGGTGGTATGGTGGCACCGAGTTCGAAAGTCACAACACCGAAACTTGAACAAAAGCTCACTCGTAACGAAGGTGTCCAATCGTCATCGTCT
GGCAACAACAGAGACTTGAATGCCGCTGCTTCTGAGGGAAGGTTGGGCAGCTTATCCAGATCAAATGCTGGCATTACCCGTAAGGGCGACGGTCCTGGTA
CTTCTGTTCGATCCTCATCTCTTCATGGAGCAGGCAACGTTTCCATTGAAAGAACTACAAATCAAGCTATCCCCAAGGATGAACCCTCACCTAGTACTTC
TTCGTCTGCAGAAAGAACACTGGATACCGGTGATGAAAGCTCATATGATGGCCTTCGTCCACAGAAATCATTGAGCCTGAATGAGAAAAGCAGGGCGAGT
TCTGTTAGTCGCACAAGACCTGCTGGTGCAGTGGGCCTAAAAGAAGCAGGTACTAGTCGGAAAAGGAAGGATATTGGTCTTTCCGGGGAAAACCCCACAT
CTGTTAGTCCTCGTGGTCCTTCTAGTGAAGCACCGGGTACTAGAAGTGTTGGTGATGAAATAAATAGAGACAGTAAACTAAAAGCTGCAATTGAAGTTGC
TATGCTGAAGAGACCGGGAATTTATAGAAAGAAGATTGATCGTGACCGATCTGACGGGCTATCCTCATCAAATGTGGATGTGAACTCTCATAAGGATCCA
CTGGCTAAGTTTTCAGGTTCAAATAGGCTGAAAGTAGCCACTTTTGACGAACGAGCTCATGAAGAGAAACCAGTCCTTGGCCCTTGGTCCACGGACTCTC
CAAAACAATCCAACATGGGAAGTCTTAACCTATCTAATACACACTCTGTTAGTACTGCATTCCCGTTGAAAGGGGACTTAGACACAGTAAACACTTCTGC
TGGAAAATCTGGTCCTATCCCACTTACAACCATTGAGAAAGCGTCCCCAATTCCAGAGCATGACTTTATCTGGCAGGGAACATTTGAGGTTAACCGGGGT
GGAAAAGTTACGAATATCCATGGAGGCATTCAAGCACACTTGTCAACTTATGCATCATCCAAAGTTCTTGAAGTGGTAAACAAGTTTCCTGAGAATGTTC
GTTTAGATGAAGTATCTCGTTCAAGTACGTGGCCGAGACAGTTCCAAGAAATTGGCGCTACTGGTGATAATATTGCTCTTTACTTCTTTGCCAAGGACCT
TGAAAGTTACAACAAGCACTACAAGAGCTTACTGGACAATGCAATGAAACGAGATTTGGCCTTCAAAGGATACTTTGATGGTGTTGAACTCTTGATATTC
CCATCCACTCAGCTTCCAGATGATTCACAACGTTGGAACATGATGTTTTTCCTTTGGGGTGTGTTCAAGGGGAGGAAATCCAGTTCTGATGATTGCCTAA
GAAAGCCAGCTGTCCCCAGTTTGAATTTGGTTGCTCAGGGTGAAGAAATGGCCGTGGCTAATGTATCATCTCCGGCGAAGAGGTTCCGACCTCAATATAT
AGGGAGAGAAATTCCCGCATTAGATGATCACTGCAACACTGCTGTTTCATCAAGCGACTTGAAAAACACGTATGTAGCAGAAAATGGTGCTGTGGATAAG
GGAAAAGTAGACTCTGTGATCAAGGTAGAGCGGCAAGAATGTATACTCGACTGTGGTGGAACTACAAGTGTGGCATCGTCAAATCAAGTTCTGAGCAGCA
GAGCTATGGCTAGCCCGGTCGACGTTGGCCGTCCAGAGTCCAGAACGGATAGAGAAGAGAAACCATGTGCTCCGGTTGTCGAGTCCAAGAACGGCTTCGA
TGACAAGGAAGCACAGAAGCAAGTAGACACTTCAAATCTCCGAGCTGATGTGTCTAAACTCGAGAACCTTCAGATCGAGACTCAAAACGTAGGTGTGAGG
GGCTGCATTTCTGATCAAGAGCCAGCAGTTACCAGGGAAGACGGTAGCAATACAGGTGGAGTTAACATCGTGAGAGACTTGAACGAAGTTGTTGATATGG
AGATCGAACCTTGTGAGGATAGAGACGACATGGCCTCCGAGAAAAGGAAAAGGCCGCTCTTGGATCTCTCCAAGATGCCACCTGAGAGTCCTAGTAGCAT
CATAGATCATGAAATGCCATGGAGTAATGCAGTAGCAAAAACTGGGTCGGTAGATGGACAAGGTGCCAACAAGAAGCGGAGAATGAGCTACGAAGGTTCT
GAGTCTTCCCCACTACCTATTGGTTCTTCAGTAGAGACGAGATACGTTGAGATGTCGGGGAAGGAAGCTGTTCCTGAAAACTCGAAAGCTCCGGAGTGGT
TATCTTTCCCAGCGGTGGATTCACACAGTGTGTCAATGAGGCCGAATACCATACCCTGGGGACGGGGCTCCTCGTCCGACCACCCATCATCGCGTCACGA
AATCCCTAACCTTAATCTCGCTTTGGAGGAAGATGAGATAGAGACCGACACCTTACCACCCCCTCCTTCATCAAGTAAGGGAATGCTGCCTTTCATAATT
TCAACAGTAGACAAGCATGCTGCTAGCCAGAATAACTCATTGCTGCCCCACCCACCCCCACCGTCGGTGGAGGAAGTGACGGAGAGAGAGGAAGAGGAGA
GCGTTTCGGCGTCCCTTTCTCTATCTCTTTCCTTTCCCTTCCCGGATAAGGAACAACCGGCCAAACCACCACCGGAATATGTATCTGCTCCTTTTTGGAA
GGTTCGCAGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10009530 pacid=23145683 polypeptide=Lus10009530 locus=Lus10009530.g ID=Lus10009530.BGIv1.0 annot-version=v1.0
MHPKAQTRVESGTCNVCSTPCSSCMHRKRAFMGSKGDEGDEFSDETCHVAETSQNSINKDDSSPIKGGMAAMLRHATSAASNVPTVDSCQDSVSESAEIK
GNLKISDRVDVSMKSEALPESSHVGSVSVVDRISPKPELIAYKRAASVNPAACKAVEGHGDNISCVTGADVSSNGDSASKLPVQNNNLSSNVDVVNLNKD
PVDQVYRESSKPIKEHTSPMLSQEAAPDILLRDNSQNGADNNLQSRSVTPGDGPEVSSRLGQEIKEETGNDSRSRPDKGNSSSVKGEEDEKVNESVELPH
TRESPAQSSSANDGDDSEIVEQDVKVCDICGDSGQEEKLAFCSRCSDGAEHTYCMKEMLLEVPDGDWLCEECKIAEDSENQKQGEEDEKVNESVELPHTR
ELPEQSSSANDGDESEIVEQDVKVCDICGDSGQEEKLAFCSRCSDGAEHTYCMKEMLLEVPDGDWLCEECKIAEDSENQKQGAEEKDASKLTYQRLKKRP
AETVEVAAISKRQAVDMDSGSSNPSSPIKVAALQRNSSSRSWDKEKVKPSHPVYDITESTRFPSTGSRLQSHKGTYLKSNSFNAISSKSKVKLVDEVPRK
HKSFKESSLSDNKDGPGKTMGKSMSFHYAGSGKAESKVKMLPSKLDSKGLKQARDYNTFERRSLPKLDRPGGMVAPSSKVTTPKLEQKLTRNEGVQSSSS
GNNRDLNAAASEGRLGSLSRSNAGITRKGDGPGTSVRSSSLHGAGNVSIERTTNQAIPKDEPSPSTSSSAERTLDTGDESSYDGLRPQKSLSLNEKSRAS
SVSRTRPAGAVGLKEAGTSRKRKDIGLSGENPTSVSPRGPSSEAPGTRSVGDEINRDSKLKAAIEVAMLKRPGIYRKKIDRDRSDGLSSSNVDVNSHKDP
LAKFSGSNRLKVATFDERAHEEKPVLGPWSTDSPKQSNMGSLNLSNTHSVSTAFPLKGDLDTVNTSAGKSGPIPLTTIEKASPIPEHDFIWQGTFEVNRG
GKVTNIHGGIQAHLSTYASSKVLEVVNKFPENVRLDEVSRSSTWPRQFQEIGATGDNIALYFFAKDLESYNKHYKSLLDNAMKRDLAFKGYFDGVELLIF
PSTQLPDDSQRWNMMFFLWGVFKGRKSSSDDCLRKPAVPSLNLVAQGEEMAVANVSSPAKRFRPQYIGREIPALDDHCNTAVSSSDLKNTYVAENGAVDK
GKVDSVIKVERQECILDCGGTTSVASSNQVLSSRAMASPVDVGRPESRTDREEKPCAPVVESKNGFDDKEAQKQVDTSNLRADVSKLENLQIETQNVGVR
GCISDQEPAVTREDGSNTGGVNIVRDLNEVVDMEIEPCEDRDDMASEKRKRPLLDLSKMPPESPSSIIDHEMPWSNAVAKTGSVDGQGANKKRRMSYEGS
ESSPLPIGSSVETRYVEMSGKEAVPENSKAPEWLSFPAVDSHSVSMRPNTIPWGRGSSSDHPSSRHEIPNLNLALEEDEIETDTLPPPPSSSKGMLPFII
STVDKHAASQNNSLLPHPPPPSVEEVTEREEEESVSASLSLSLSFPFPDKEQPAKPPPEYVSAPFWKVRR
|
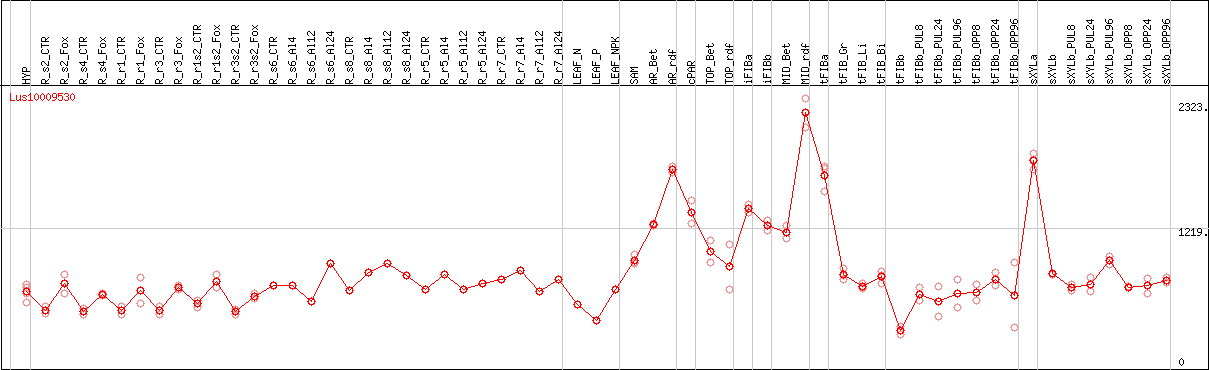
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009530 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
