External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G07020 228 / 2e-70
UGT80A2, SGT
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
AT1G43620 182 / 6e-53
UGT80B1, TT15
TRANSPARENT TESTA 15, UDP-Glycosyltransferase superfamily protein (.1.2.3)
AT5G24750 56 / 7e-09
UDP-Glycosyltransferase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10020382
305 / 5e-100
AT3G07020 790 / 0.0
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10009556
248 / 2e-78
AT3G07020 707 / 0.0
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10020381
246 / 7e-78
AT3G07020 568 / 0.0
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10011882
247 / 2e-77
AT3G07020 885 / 0.0
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10022815
239 / 1e-75
AT3G07020 742 / 0.0
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10024544
235 / 7e-73
AT3G07020 714 / 0.0
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
Lus10018878
186 / 2e-54
AT1G43620 849 / 0.0
TRANSPARENT TESTA 15, UDP-Glycosyltransferase superfamily protein (.1.2.3)
Lus10028572
183 / 3e-53
AT1G43620 840 / 0.0
TRANSPARENT TESTA 15, UDP-Glycosyltransferase superfamily protein (.1.2.3)
Lus10011883
181 / 5e-53
AT3G07020 648 / 0.0
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G178300
254 / 2e-80
AT3G07020 817 / 0.0
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
Potri.002G238600
253 / 3e-80
AT3G07020 839 / 0.0
UDP-glucosyl transferase 80A2, sterol glucosyltransferase, UDP-Glycosyltransferase superfamily protein (.1.2)
Potri.002G067100
192 / 1e-56
AT1G43620 800 / 0.0
TRANSPARENT TESTA 15, UDP-Glycosyltransferase superfamily protein (.1.2.3)
Potri.005G193100
191 / 4e-56
AT1G43620 805 / 0.0
TRANSPARENT TESTA 15, UDP-Glycosyltransferase superfamily protein (.1.2.3)
Potri.003G021100
64 / 3e-11
AT5G24750 606 / 0.0
UDP-Glycosyltransferase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0113
GT-B
PF00201
UDPGT
UDP-glucoronosyl and UDP-glucosyl transferase
CL0113
GT-B
PF03033
Glyco_transf_28
Glycosyltransferase family 28 N-terminal domain
Representative CDS sequence
>Lus10009557 pacid=23145693 polypeptide=Lus10009557 locus=Lus10009557.g ID=Lus10009557.BGIv1.0 annot-version=v1.0
ATGTCACGTAGTTGGCTATTGACTATTAGTCGTGTTGGATCAAAAGAGTACCTGTTATCCCAGAACATACCTGTTCAGAGGCAACAAATGAGAGAAATCA
TATTCTCTTTACTTCCTGCGTGTCAGGATCCTGATCCTGAAACAAATATTCCGTTCAGAGCAGAAGCGATCATTGCCAATCCTCCAGCATATGGCCATAC
ACATGTTGCAGAGGCACTAAAGGTGCCTTTACATGTGATCTTCACAATGCCTTGGACGCCTACTACTACTGAATTCCCACATCCTATGTCTCGTGTTAAA
CAACCAGTGGCAAATAGAGTGAGCACTCTTTATTTGCAACAGCTTGTTCAAGAACCCGAGAAGATGACACAGATGATTGTAAAGGCTTTGGAACTTACTG
GTCAAAGAGGCATCATCAACAAAGGCTGGGGTGGGTTAGGAAATTTGGCAGAACCAAAGGACTCCATCTACTTGCTGGACAATTGCCCTCATGACTGGCT
CTTCCTAAGATGCAAGGCTGTGGTGCACCACGGAGGTGCTGGAACAACTGCTGCTGGTCTTAAAGCTGCGTGTCCAACCACTATAGTGCCTTTCTTCGGT
GACCAGCCGTTTTGGGGAGATCGAGTCCATGCTAGAGGAGTCGGCCCTGCACCGATACCAGTGGACGAGTTCTCGACTGAAAAACTGGTCGAAGCAATCA
AATACATGGTGGAGCCTGAGGTGAAACAGCGAGCAGTAGAAATCGGGAAGGCGATGGAGAATGAAGACGGGGTGGCAGGTGCGGTGAGAGGCTTCTACAA
GCATTTCCCCTTGAAGAAGACAGAGGAGGAAGAAGCATGCTAA
AA sequence
>Lus10009557 pacid=23145693 polypeptide=Lus10009557 locus=Lus10009557.g ID=Lus10009557.BGIv1.0 annot-version=v1.0
MSRSWLLTISRVGSKEYLLSQNIPVQRQQMREIIFSLLPACQDPDPETNIPFRAEAIIANPPAYGHTHVAEALKVPLHVIFTMPWTPTTTEFPHPMSRVK
QPVANRVSTLYLQQLVQEPEKMTQMIVKALELTGQRGIINKGWGGLGNLAEPKDSIYLLDNCPHDWLFLRCKAVVHHGGAGTTAAGLKAACPTTIVPFFG
DQPFWGDRVHARGVGPAPIPVDEFSTEKLVEAIKYMVEPEVKQRAVEIGKAMENEDGVAGAVRGFYKHFPLKKTEEEEAC
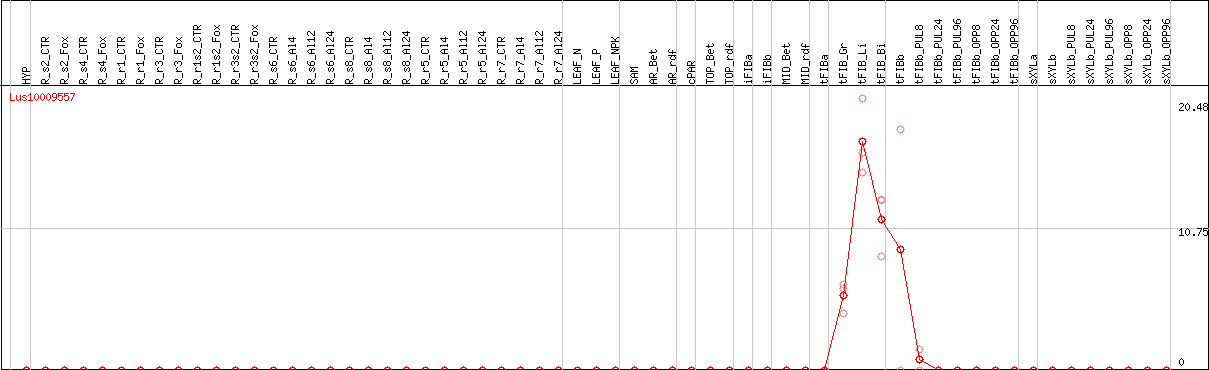
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009557 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.