External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G23340 358 / 6e-124
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
AT5G51310 336 / 2e-115
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G50960 147 / 2e-41
ATGA2OX7
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 7, gibberellin 2-oxidase 7 (.1)
AT1G80330 132 / 1e-35
ATGA3OX4
ARABIDOPSIS THALIANA GIBBERELLIN 3-OXIDASE 4, gibberellin 3-oxidase 4 (.1)
AT4G21200 131 / 2e-35
ATGA2OX8
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
AT2G38240 127 / 1e-33
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G07200 122 / 8e-32
ATGA20OX3, YAP169
ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 3, gibberellin 20-oxidase 3 (.1)
AT1G52820 119 / 4e-31
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G25300 119 / 7e-31
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
AT3G19010 119 / 8e-31
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018836
325 / 7e-111
AT5G51310 348 / 7e-120
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10009025
276 / 1e-92
AT5G51310 131 / 9e-38
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10021002
134 / 3e-36
AT3G19000 389 / 3e-135
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10007640
129 / 2e-34
AT4G21200 429 / 1e-151
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10015016
127 / 1e-33
AT4G25420 494 / 3e-176
GA REQUIRING 5, ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10030567
126 / 2e-33
AT4G21200 370 / 2e-128
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10018372
126 / 2e-33
AT4G21200 419 / 2e-147
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Lus10016659
125 / 2e-33
AT1G52820 445 / 1e-158
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10031671
126 / 8e-33
AT5G07200 508 / 1e-180
ARABIDOPSIS THALIANA GIBBERELLIN 20-OXIDASE 3, gibberellin 20-oxidase 3 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G106900
398 / 1e-139
AT4G23340 392 / 1e-137
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.002G159500
383 / 1e-133
AT4G23340 402 / 3e-141
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.003G128100
370 / 1e-128
AT4G23340 458 / 2e-163
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.001G418200
133 / 4e-36
AT4G21200 369 / 8e-128
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.004G022800
133 / 4e-36
AT4G21200 441 / 7e-156
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.008G145300
133 / 4e-36
AT4G21200 354 / 1e-121
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.010G201000
132 / 1e-35
AT3G21420 290 / 5e-96
LATERAL BRANCHING OXIDOREDUCTASE 1, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.010G096800
131 / 2e-35
AT4G21200 362 / 8e-125
ARABIDOPSIS THALIANA GIBBERELLIN 2-OXIDASE 8, gibberellin 2-oxidase 8 (.1.2.3)
Potri.015G002800
130 / 5e-35
AT5G24530 521 / 0.0
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G176600
130 / 7e-35
AT1G15550 452 / 1e-159
GA REQUIRING 4, ARABIDOPSIS THALIANA GIBBERELLIN 3 BETA-HYDROXYLASE 1, gibberellin 3-oxidase 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
Representative CDS sequence
>Lus10009663 pacid=23140019 polypeptide=Lus10009663 locus=Lus10009663.g ID=Lus10009663.BGIv1.0 annot-version=v1.0
ATGTCTCAATCTGCGACTGTAAAGCTCCCTGTGTTCGACATCTCGGTCCCGTTCTCGAAATCGGCCATTGATGACCTCTCCGCAGCTTGCAAGGAGTGGG
GGTTCTTCCACATAACCAACCATGGAGTTTCCAAGGAGCTCTACAGCAAGGTCAGAGCTTTCTCCTTGGAGGATCTCTTCTCTCAGCCTTTGGAGATGAA
GATCAGAGCTGGACCGAGCTCCGATGTCAAGTCCTACACTCCGCATTTCATCGCTTCGCCTTTCTTCGAGAGCTTTCGCGTCTCCGGTCCCGGTTTTTTG
GATTCTGTTCATGAATCGGTTCGAGTGGTCCGTGGTCCGGTTTCAAGCTATCCAGAATTAAGTCAAGTAGTGGAAGAGTACGGGAAGAAAATGACGAATC
TATCTGAAACCATAGTAAAAACCATCCTCGCAACCATGAGTAAGGAATTGGTGGAGAAATTGTACATTTCCCACTTCAAAGACTGCCATGGATACCTAAG
AATCATCAATTACTCCCCACCAAACCCACATCAAGAAGAACAAGAAGTTGAAGGGCTCGGGATGCACACAGACATGAGCTGCGTGACCATAGTATGCCAG
GACGAGATCGGGGGGCTACAGGTTCGAACAAAAGAAGGGAAGTGGATGGACATCGAGCCACGACCCGATTCACTCGTGGTGAACATCGGAGACATGTTGG
AAGCTTGGAGCAATGGGGAGCTGAGATCATCCGAACACAGGGTTGTGCTGAAGAAACCTGTCGAAAGGTTCTCTTTGGCATTTTTCTGGTGCTTTGAGGA
CCAAAAGGTAATTTTGGCGCCCGAGGAGGTTGTAATTGGGAAGAGGAGAAAGTATAGGGAGTTTGTTTGTAAAGAGTATTTGAGTTATAGAGAGACTAGT
GTTAAGGGGAGGTTTGATAAGGTTGGGTTCACTGTTAGGGATTTCGCTGCACTCGATGGATCGATCGTAAATTCGTGA
AA sequence
>Lus10009663 pacid=23140019 polypeptide=Lus10009663 locus=Lus10009663.g ID=Lus10009663.BGIv1.0 annot-version=v1.0
MSQSATVKLPVFDISVPFSKSAIDDLSAACKEWGFFHITNHGVSKELYSKVRAFSLEDLFSQPLEMKIRAGPSSDVKSYTPHFIASPFFESFRVSGPGFL
DSVHESVRVVRGPVSSYPELSQVVEEYGKKMTNLSETIVKTILATMSKELVEKLYISHFKDCHGYLRIINYSPPNPHQEEQEVEGLGMHTDMSCVTIVCQ
DEIGGLQVRTKEGKWMDIEPRPDSLVVNIGDMLEAWSNGELRSSEHRVVLKKPVERFSLAFFWCFEDQKVILAPEEVVIGKRRKYREFVCKEYLSYRETS
VKGRFDKVGFTVRDFAALDGSIVNS
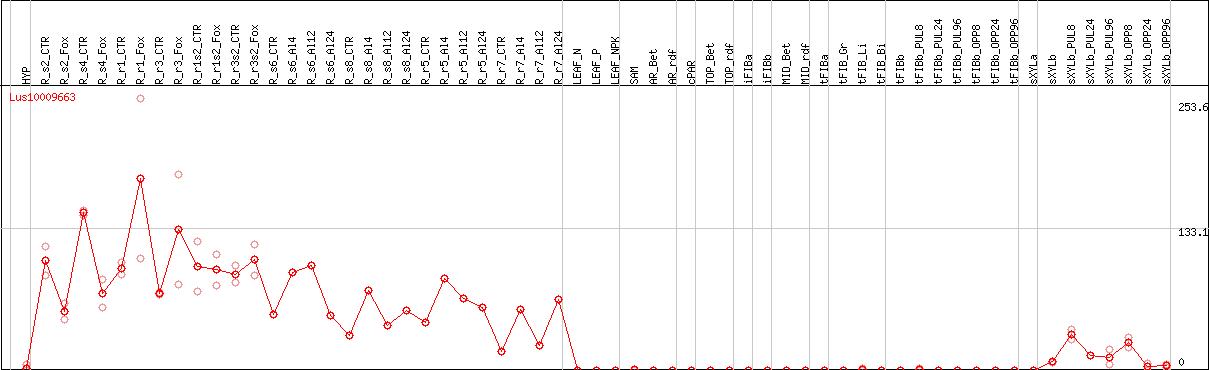
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009663 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.