External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G22900 112 / 4e-27
ATCHX3
cation/H+ exchanger 3, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 3, cation/H+ exchanger 3 (.1)
AT3G44910 111 / 6e-27
ATCHX12
cation/H+ exchanger 12, cation/H+ exchanger 12 (.1)
AT3G44900 107 / 2e-25
ATCHX4
cation/H+ exchanger 4, cation/H+ exchanger 4, cation/H+ exchanger 4 (.1)
AT1G08150 99 / 1e-22
ATCHX5
CATION/H+ EXCHANGER 5, Cation/hydrogen exchanger family protein (.1)
AT1G08140 96 / 1e-21
ATCHX6a
cation/H+ exchanger 6A, cation/H+ exchanger 6A, cation/H+ exchanger 6A (.1)
AT5G22910 92 / 2e-20
ATCHX9
cation/H+ exchanger 9, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 9, cation/H+ exchanger 9 (.1)
AT3G44930 91 / 6e-20
ATCHX10
cation/H+ exchanger 10, cation/H+ exchanger 10, cation/H+ exchanger 10 (.1)
AT1G08135 90 / 1e-19
ATCHX6B, CHX6B
cation/H+ exchanger 6B, cation/H+ exchanger 6B (.1)
AT2G13620 87 / 1e-18
ATCHX15
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
AT2G28170 87 / 1e-18
ATCHX7
CATION/H+ EXCHANGER 7, Cation/hydrogen exchanger family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000161
541 / 0
AT5G22900 320 / 6e-99
cation/H+ exchanger 3, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 3, cation/H+ exchanger 3 (.1)
Lus10040962
341 / 1e-117
AT3G44910 64 / 2e-15
cation/H+ exchanger 12, cation/H+ exchanger 12 (.1)
Lus10040963
317 / 3e-101
AT3G44900 376 / 2e-117
cation/H+ exchanger 4, cation/H+ exchanger 4, cation/H+ exchanger 4 (.1)
Lus10013423
314 / 1e-100
AT3G44900 375 / 2e-117
cation/H+ exchanger 4, cation/H+ exchanger 4, cation/H+ exchanger 4 (.1)
Lus10026408
110 / 3e-26
AT2G13620 441 / 2e-142
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
Lus10041344
109 / 4e-26
AT5G22900 511 / 4e-170
cation/H+ exchanger 3, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 3, cation/H+ exchanger 3 (.1)
Lus10017540
108 / 1e-25
AT5G41610 1013 / 0.0
cation/H+ exchanger 18, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 18, cation/H+ exchanger 18 (.1), cation/H+ exchanger 18 (.2)
Lus10037374
107 / 3e-25
AT3G44900 502 / 8e-167
cation/H+ exchanger 4, cation/H+ exchanger 4, cation/H+ exchanger 4 (.1)
Lus10043128
106 / 5e-25
AT3G44900 540 / 0.0
cation/H+ exchanger 4, cation/H+ exchanger 4, cation/H+ exchanger 4 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G182900
250 / 8e-77
AT3G44900 365 / 1e-114
cation/H+ exchanger 4, cation/H+ exchanger 4, cation/H+ exchanger 4 (.1)
Potri.010G171200
247 / 2e-75
AT2G13620 360 / 4e-112
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
Potri.007G099000
222 / 3e-66
AT5G22900 385 / 1e-121
cation/H+ exchanger 3, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 3, cation/H+ exchanger 3 (.1)
Potri.019G123700
131 / 1e-33
AT2G13620 536 / 4e-179
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
Potri.018G098300
125 / 1e-31
AT5G22900 470 / 3e-154
cation/H+ exchanger 3, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 3, cation/H+ exchanger 3 (.1)
Potri.009G078000
108 / 6e-26
AT5G37060 677 / 0.0
cation/H+ exchanger 24, ARABIDOPSIS THALIANA CATION/H+ EXCHANGER 24, cation/H+ exchanger 24 (.1)
Potri.006G176400
107 / 2e-25
AT3G44900 442 / 8e-144
cation/H+ exchanger 4, cation/H+ exchanger 4, cation/H+ exchanger 4 (.1)
Potri.016G127800
103 / 4e-24
AT5G01680 436 / 6e-142
cation/H+ exchanger 26, ARABIDOPSIS THALIANA CATION/HYDROGEN EXCHANGER 26, cation/H+ exchanger 26 (.1)
Potri.019G123800
102 / 1e-23
AT2G13620 443 / 5e-144
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
Potri.007G105200
100 / 6e-23
AT2G13620 1112 / 0.0
CATION/H+ EXCHANGER 15, cation/hydrogen exchanger 15 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0064
CPA_AT
PF00999
Na_H_Exchanger
Sodium/hydrogen exchanger family
Representative CDS sequence
>Lus10009846 pacid=23180193 polypeptide=Lus10009846 locus=Lus10009846.g ID=Lus10009846.BGIv1.0 annot-version=v1.0
ATGATATTCCCACCATTCCCAAACCAAGTGTTGGGTTCCTTATCCAAGATAGGATACGTATTATTCGCATTCTTGGCAGGAATAAGGATGGATCCAAGCC
TGATCCCGCAATCCGGCAGGAAAGCAGTATTCATGGGGTTCCTTCTATTTGCTTACCCTTTCACTATGATACACACAATTGTAGTTGAGTTCCAAAAAGA
CAACATTTTCAACTTCGAGCTCAAACTTGCAATTTACAAAGGTGTTGGAATTGTCTTTGGTTCCATCACAACTTCCCAATTCGTTGGAGTGAGTCACCTC
TTGACACAGCTTAGGATAACCAACTCTCAGCTTGGACACCTTGCCTTAGCATCCACATTGGTCAGTGAGTTGCTCCGATTCGCCTACTCCTCCACTCTTG
GTGTCCTGGAAGCGTTCATGTTCGTAGCTAGGAGATCGGGGATCCAATCCTTACTCTTCAGCATGCTTTTGGTAGGCTTCATTATGGTTGTGTTGAGAGG
GTTTGTGCTTTGGGCAATCAGGAACACCCCACAGGGGAAACCCATGAAGGAAATCTACATAACTATGGTTGTTGGTGTGCTTCTCAGTTTAGTCACACTT
GGGGATTCTGTTGGAATGCATTACCTTTTTGGCCCTTTCCTTTTGGGTCTTATCATACCAGCTGGATCTCCATTGGCAACCAGTTTGATTGCCAAGCTTG
AAACTGCTGTCAATGGCTTGTTTATCCCCATCATTCTCATGTTCTGCTCCACAAGCTTTGATTTGGTAGCCTTTTCCCAGGATTACTCCAAAGTTCTTCC
CTTTAGGATTACCTTGCTTGGTTATTTCTTGAAGTTGGCCTTCACTTTCTTAATGGGACTCGGTTTTAAGCTATCTGCTAAAGATTGTGCAGCTTACACA
CTCATTGTCAATGCTAAAGGTGTTCTTGAAGTTGGCGTCTTCCTCAGCTTTGGTAACCTTGCGGTACATCTGATTCATTCATTATCTCTACATACATATA
TAGAGTAG
AA sequence
>Lus10009846 pacid=23180193 polypeptide=Lus10009846 locus=Lus10009846.g ID=Lus10009846.BGIv1.0 annot-version=v1.0
MIFPPFPNQVLGSLSKIGYVLFAFLAGIRMDPSLIPQSGRKAVFMGFLLFAYPFTMIHTIVVEFQKDNIFNFELKLAIYKGVGIVFGSITTSQFVGVSHL
LTQLRITNSQLGHLALASTLVSELLRFAYSSTLGVLEAFMFVARRSGIQSLLFSMLLVGFIMVVLRGFVLWAIRNTPQGKPMKEIYITMVVGVLLSLVTL
GDSVGMHYLFGPFLLGLIIPAGSPLATSLIAKLETAVNGLFIPIILMFCSTSFDLVAFSQDYSKVLPFRITLLGYFLKLAFTFLMGLGFKLSAKDCAAYT
LIVNAKGVLEVGVFLSFGNLAVHLIHSLSLHTYIE
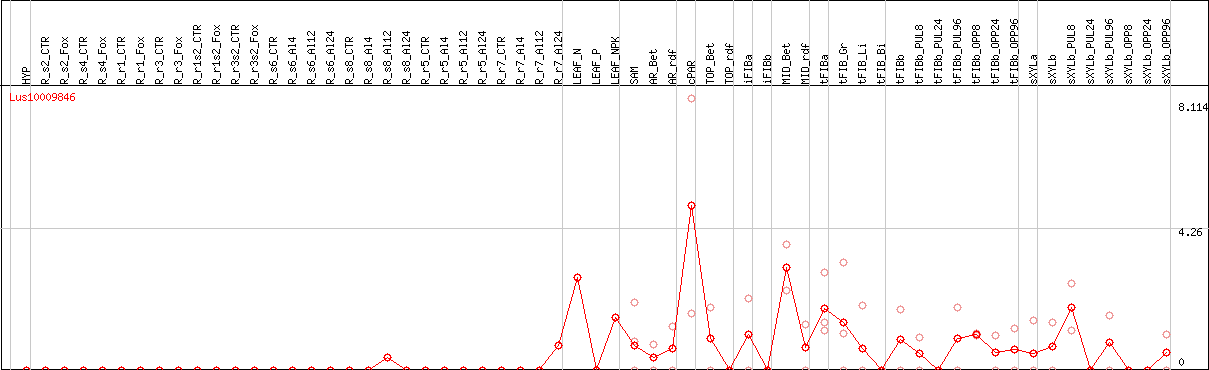
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009846 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.