Lus10009940 [FLAX]

| External link |
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||
|
Arabidopsis homologues
|
| ||||||||||
|
Paralogs
|
|
||||||||||
|
Poplar homologues |
|
||||||||||
| PFAM info |
| ||||||||||
|
Representative CDS sequence |
>Lus10009940 pacid=23150088 polypeptide=Lus10009940 locus=Lus10009940.g ID=Lus10009940.BGIv1.0 annot-version=v1.0
ATGCACCGTACGTCGCTGCTAGGGCTGACGAGTACATCCGATGCTTGCAGACGGAGTTTGATACCGTGCTTGCAGACGGAGTTTGGTACCGTCAAGGCTG
CCGCAGATGCCAACGGCGTCACCGCTGAGCAGACTTGTTCGCTTCTCGAACACAAGTTTTTGTCTCTGTCCGCCGAGTTCTCTACGCTGCAGTCCCAGAA
CGCCCAGCTTCAGTCCTCGTTCGATGAGCGACTCTCGGAACTCGCCGACGTCCAGACTCAGAAGCACCAGCTTCACTTGCAATCGATGGAAAAGGATGGT
GAAATAGAGAGGCTGACAACGGAGCTTTCAGAGCTGCGTAAAACTAACAGGCAGTTGATGGAGTTGGTGGAACAAAAAGATGCGGAAATCAGCGATAAGA
AAGTTACTATTAACAGCTATCTCGATAAGATTGTTAATTTGACGGAGAGTGCATCAAAAAAGGAAGCCCGTTTAAGTGAAGTTGAAGCAGAGCTGTTGCG
ATCTCAGGCCCTCTGTTCTCGCCTTTCACAGGCTGAGAAACAACTGGGTGAATGCTCAAGCTCCGCGAAGAGAAAAATGGAGAGGGTAGAGGAACTCGAA
GCGAAGTTAACCTCCTTGCAAGAGGAATTATGCTCTACCAAGGATGGTGCTTTAGCAAATGAAGAGCAGTTTTTGGCTGAACTGACTACTGCTAATAAAC
TAATTGAGCTCTATAAGCAAAGTTCTGAGGAATGGTCACAAAAAGCTGGAGAGCTTGAAGGTGTGATTAGAGCTTTAGAGAAACACTTGAGTGAGCTTGA
TACTGATTACAAAGCAAGACTTGAAAAAGAAATACTTTCGAGAAGAGAATTAGAGAAGGAAGCTGCTGATCTCAGAAGTAGACTTGAACAATGTGAAGCT
GAGATGGAAACCAACAGAAAGAATAATGAGCTTAATCTACTTCCACTCAGTAGTTTTTCAACAGAAAGGTGGAAGTCACAGTCTGATGGTAATGATACTA
TGGAAGTTACTGACATGCTCATCCCCAAGATACCTCCTGGTGTTTCTGGGACTGCATTGGCAGCTTCTTTACTTCGCGATGGATGGAGTTTGGTCAAGAT
GTATGCGAAGTATCAAGAAGCAAATGATGCTTTTCTTCATGAACAACTAGGAAGGAAGGAATCTGAAGCTGTGTTGCAGCGGGTCCTGTTTGAACTGGAG
GAGAAAGCGGGAGTCATTTTGGAAGAAAGAGCGGATTATGAGAGAATGGTCGAATCATACTCTATTATGGAGAAAAGGCTGCAACAATCATTGTCTGAAC
AGGGCAATCTGGAGAGAATGATCCAACAGTCAAAGGCTGAACTAAGAAGGCATGAGCGACAGTATGATTATGCTCAGAAGGATATTGTTGACCTTCAGAA
GCAGATTACAATTCTTCTAAAGGAATGTCGTGATGTGCAACTACGATGTGGATTGGGGGATAATGCAGATGATGATTTTCCGGTACATGGTGCTGTTGGT
ATGGAGACAGACACTGTTGAGGATAATGTTCTTTCTGAAAGGCTTTTGACTTTCAAGGACATAAATGGATTAGTTGAGCAGAATGTGCAACTGAGGAGTC
TCGTGCGAAGTCTTTCAGATCAGATTGAGAATAATGAGGAGGAGTTCAAGGTGAAATTGGACATGGAACTCAAAAAAAAAACTGATGAAGCGGCTTCTAG
AGTTGCTATAGTCTTACAAAGGGCAGAAGAACAAGGGCAGATGATTGAATCTCTTCATACAGCTGTTGCTATGTATAGAAGGTTATATGAGGAAGAACAG
AAACGTCATTCGCCTAATTCTCGTCCGCATTCATTGGAGGAAGCTTCAGAGGATGCGAGGGGGGATCTTTTGGTTCTACTTGAGGGTACTCAGGAAGCTA
GCAAGAAGGCCCAGGAAAAGGTAGCTGAGCAGCTGGGATCGCTTGAGGAAAAGCTATCAAAATATAGGAGTGACATTGTGTCGCTGCAATTGGAGCGTGA
CAAGTTGGCTTCAGAGGCAAAACTTGCGGTAGAAAAACTTGAGAGATTTATGAAAGAATTCGAACATCAGAACACTCAGATCCACGGTCTTCAAGATGGA
AATTTGGAGTACTCGCGCCTTATTCTCGACTTTCAGAAGAGGTTAAATGAAAGTATGCATGCTCAAAATGCTTCCGAGGATCATTGTCGGAGGTTGAACA
TGGAGGTGTCGACCCTAAAGCACGAGAAGGAAATGTTATCAAATGCTGAAAAGAGAGCATGTGATGAAGTTGGTAGATTGTCAGAGAGGGTGCACAGGCT
GCAGTCTAGCTTAGGCACTTTGCAGAGTGCTGAGGAGGTTCGGGAGGAAGCAAGGGCTGCCGAGAGGAGAAAACAGGAGGAACATGTTCAACAAATTGAG
AGGGAATGGGTGTTGGCGAAAAAAGAGCTCGAACTGGAGAAGGACAATGTCAGGGCCCTCACATGTGGTCGTGAGCAGATTTTGAAGGATGCAATGAAAC
AAGTGGGTGAGATTGGAAAAGAGTTAGCTAATGCGTTAGGTGCAGTAACAGTCGCTGAGACCAGGGCTGCTGTTGCTGAGACTAAAGTTGAGGATTTGGA
GAAGAAGCTCAGGATTTCAAATCAAAAGGCCGCCACTGATTTGATTATGCTGAAGGAGGAGTTAGAAAAGTTGAGACAAGAGGCACAAACTAATAAGGAC
TACATGGTGCAGTATAAGAATATAGCTGAGGTAAATGAAGATGCACTGAAGCAAATGGAAGCTGCTCATGAGAACTTCAAGAGAAAGTCAGAAAAGCTCA
AAGAGTCTTTGGAAGCTGAGTTGATTGCACTAAGAGGGAAGATATCTGAACTTGAAAATGAATTGCGTTTGAAGTCTGAGGAAGTAGTTGCCCTTGCTGC
TGGTAATGGAGAAACTCTTACCTCTGCTCTGGCTGAAGTTACTGCCTTGAAGGAGGAAGCTTCGAATAAAATGTCTCAAACTGCTGCATTGGAAATGCAA
GTTTCGGTATTGAAAGTGGATCTTGAGAAGGAGCATGACCGATGGCGATCTGCTCAAAATAATTACGAGAGACAGGTCATTTTACAATCTGAGACAATTC
AGGAGTTAACTAGGACATCGCAGACTTTGGCATCATTACAAGAAGAAGCAACTACCTTGCGTAAATTGGCCGATGAGCGAGAACTCGGAAATAATGAGTT
GAAAGCCAAGTGGGAGGCTGCACAATCAATGCTAGAAGAATCAAGGAGGGAATCTCAGAAGAAATATGACGAGCTCAATGAACAGAATAAAGTACTGCAC
GACAGACTGGAGGCTCTGCACATCAAGTTTGCTGAGAGTCACCGCAGCTCTGCTGGAGTGTCTTCTAGAAACGGTGTTGATTCAAGCGATGATGGAGGAT
TGCAGAAAGTTGTTGGCTATCTTCGACGATCCAAGGAGATGGCAGAGACAGAGAGGGCTTTATTAAAGCAGGAAAATTCTCGATTGAAATCACAACTTGA
TAATGCTCTCAAGGCAGCTGAAAATGCTAAAGTTTCACTCGGTGCTGAGCGTGCAAACACGAGGGCATTAATTTTTTCAGAGGAAGAAATGCAATCTCTT
CAGAATCAGGTTATGGAGATGAACTTGTTACGTGGGAACAATATGCAACTTATAGAAGAGAACAAGCAGAATGTTGAGGAATGTCAGAAATTGCGTGATT
TAGCACAAGAGAGTGATGCCAAGGCTGAAGAACTGAAACTTCTACTGAGGGAGAAAGAAAAGGAAATCGATGCTTGCAAGAAAGAATTAGAAATGTTAAG
GGCGGATAAAGTTTTTTTGGATAATAAGATATCTGAAAAGAATAATGAGTTGGAGGAGGCCAAGAACTGCATGTTGAAGCAGCAAGAGACCCTGTCGAAG
TTGGAGCATGATCTTGGAAGAAGTGAAGTAGAGCTGAAACAAAGAAGTAACGATCTTCTGGAAGCAGAGGCTAACATAAAGAGTGAATTAGAAAGGCAGA
AGAGGCAATTCACTCTTAACAGGAGAAAATTGGACAATTTAACGAAGCAAAGGGATGAGTTAATCAAAGAAAAGAATGCAGCTAACAAACAGATTGAAGA
ACTAAAGCAAGCGAAGAAGGCCATTGGCAATGTTGGTGGTGAGCACTCCCATAAGGAAAAGGAAGACAAAACAAGATTACAGATACTGGAAAAAACAGTG
GAGAAGCGAAGAGAGGAGTTGAAGAAAGCAAAAGATGAGTTGAGGAAGGAAAAGGAGGATCATCAAATGGAGAAATCGAAGTTTGCGGACATATATGATA
AAAGTAGGGAGGCCCTGAGGCGCGTTTTGGATGCACTAGAAAAGCTTAAGCACGTTGAGGATGCTCTCCCTGAGTTTCTTTCTGGAAGCGTTTTGGATGA
TCTCATTGCTGTTTGCACATCAGCCATTGAGATTTTTGGGAAAGAAGTGAATTCGTTGCTGGGTGACCTTGGGCAAAGCACACCCTCAGTAGAGACTCCA
TCTCTGGATGCTCCTGCACCAGTTTCAGCTAGTGAAACAACTTCTCAAATTCCTCTCATCCCAGCTGCTTCTGCCTCTCAAGCTGCTGGAAAAGCTGCTG
AAGAGAGGGAGAAAAGATTCTCAGCTCCCCAGATAAATGCCTCCGAAGGTCGTAAAACAGCAAGAAGGTTGGTGAGGCCCAGTTTTTCCAAGACAGAGGA
ACCTCAGGGTGATGCAGAGATATCTGAGGCAGATGGATCCAGCGTTGAGGCGAAACCTGCTGTGTTGCAAGATTCTGAAATGCAACCTAACCCAACTGTG
TCGTTTCAGCCACTCAGTCGTAAACGATCTTTACAGTCTGGTATAGAGAGCAGTGAGCAACAAATCAATCAGGGGGACGCAGTTCCCCAGGGAGCTGTCC
AGATGCCGAAGAAGGCAAAATTACAGGAATCCGCAGATGGAGAGGTGGAAGTAGAGCCTAATACATCAGCAGTTGACCTTACGCTGGATTCCACTAGTGA
TGTAATTCCTGATGGACTCGTTTCTGAAAAAGAAGATGCGGACGTAATTACTGGGAAAAAGATGGAGGCTTCAAAGGATGCAGCAGAGGATGTAGGTGAA
GAAGAGATTCAGAAAACTGATATTTCAGAGAAGCCGAGTGGAACCGGGATGGAGATTGACGAGAGCTTGAAGGATGAGATGGTAGAGGAGGACCAACAAC
CGATATTGGAGTCTGATGGTGACCGAGAAGAGGGAGAGATGCTACCAGATGTCTCAGAAGTGGAAGTGTCCAATACGGCAGACAGCCCCGAAAGTGGCGA
GTTTGTACCCGAGACAACACCCGAGGGGTCGCCATTGAAGATGGAAGATGAAACCAACTCCCCCCTAAATGCTCCCAACGATGAAGCTGACATCACGGTT
GAGGATACCGGTGAAGGTTCGGAGAAGGCAAATGACATTGGCGAGCCGAATGTGGTGGAAACTGACCCGATTCTTGAAAGTCCGCCAGTTGTCGGGGACA
GTACAACTAAGCCGAGTGCAGAAGTAGTTACCGTTGTGAAACCAAGAAGTCCTTGCGCAATCACGGCGGAGGAGGTTAAGCACCCAACACCAACTCTCGT
TAATTTGACGGATAGAGCAAGGCAGAATGCTGAGAAAAGACTACAAAGACCTGCACCAACACAACCCAATGCATCGCCGCCGCCCTCAATCCCAGGCAAT
AGAGGTCGAGCCAGAGGAGGATCAGTACGTGGTCGCGGTGTTCAAAGAACTTTTCGTGGAGGCCGGACTGGGCGAGGTGGCCCGTCTCCTGGCCAACCAG
GTCAAGGCTAA
|
||||||||||
|
AA sequence
|
>Lus10009940 pacid=23150088 polypeptide=Lus10009940 locus=Lus10009940.g ID=Lus10009940.BGIv1.0 annot-version=v1.0
MHRTSLLGLTSTSDACRRSLIPCLQTEFGTVKAAADANGVTAEQTCSLLEHKFLSLSAEFSTLQSQNAQLQSSFDERLSELADVQTQKHQLHLQSMEKDG
EIERLTTELSELRKTNRQLMELVEQKDAEISDKKVTINSYLDKIVNLTESASKKEARLSEVEAELLRSQALCSRLSQAEKQLGECSSSAKRKMERVEELE
AKLTSLQEELCSTKDGALANEEQFLAELTTANKLIELYKQSSEEWSQKAGELEGVIRALEKHLSELDTDYKARLEKEILSRRELEKEAADLRSRLEQCEA
EMETNRKNNELNLLPLSSFSTERWKSQSDGNDTMEVTDMLIPKIPPGVSGTALAASLLRDGWSLVKMYAKYQEANDAFLHEQLGRKESEAVLQRVLFELE
EKAGVILEERADYERMVESYSIMEKRLQQSLSEQGNLERMIQQSKAELRRHERQYDYAQKDIVDLQKQITILLKECRDVQLRCGLGDNADDDFPVHGAVG
METDTVEDNVLSERLLTFKDINGLVEQNVQLRSLVRSLSDQIENNEEEFKVKLDMELKKKTDEAASRVAIVLQRAEEQGQMIESLHTAVAMYRRLYEEEQ
KRHSPNSRPHSLEEASEDARGDLLVLLEGTQEASKKAQEKVAEQLGSLEEKLSKYRSDIVSLQLERDKLASEAKLAVEKLERFMKEFEHQNTQIHGLQDG
NLEYSRLILDFQKRLNESMHAQNASEDHCRRLNMEVSTLKHEKEMLSNAEKRACDEVGRLSERVHRLQSSLGTLQSAEEVREEARAAERRKQEEHVQQIE
REWVLAKKELELEKDNVRALTCGREQILKDAMKQVGEIGKELANALGAVTVAETRAAVAETKVEDLEKKLRISNQKAATDLIMLKEELEKLRQEAQTNKD
YMVQYKNIAEVNEDALKQMEAAHENFKRKSEKLKESLEAELIALRGKISELENELRLKSEEVVALAAGNGETLTSALAEVTALKEEASNKMSQTAALEMQ
VSVLKVDLEKEHDRWRSAQNNYERQVILQSETIQELTRTSQTLASLQEEATTLRKLADERELGNNELKAKWEAAQSMLEESRRESQKKYDELNEQNKVLH
DRLEALHIKFAESHRSSAGVSSRNGVDSSDDGGLQKVVGYLRRSKEMAETERALLKQENSRLKSQLDNALKAAENAKVSLGAERANTRALIFSEEEMQSL
QNQVMEMNLLRGNNMQLIEENKQNVEECQKLRDLAQESDAKAEELKLLLREKEKEIDACKKELEMLRADKVFLDNKISEKNNELEEAKNCMLKQQETLSK
LEHDLGRSEVELKQRSNDLLEAEANIKSELERQKRQFTLNRRKLDNLTKQRDELIKEKNAANKQIEELKQAKKAIGNVGGEHSHKEKEDKTRLQILEKTV
EKRREELKKAKDELRKEKEDHQMEKSKFADIYDKSREALRRVLDALEKLKHVEDALPEFLSGSVLDDLIAVCTSAIEIFGKEVNSLLGDLGQSTPSVETP
SLDAPAPVSASETTSQIPLIPAASASQAAGKAAEEREKRFSAPQINASEGRKTARRLVRPSFSKTEEPQGDAEISEADGSSVEAKPAVLQDSEMQPNPTV
SFQPLSRKRSLQSGIESSEQQINQGDAVPQGAVQMPKKAKLQESADGEVEVEPNTSAVDLTLDSTSDVIPDGLVSEKEDADVITGKKMEASKDAAEDVGE
EEIQKTDISEKPSGTGMEIDESLKDEMVEEDQQPILESDGDREEGEMLPDVSEVEVSNTADSPESGEFVPETTPEGSPLKMEDETNSPLNAPNDEADITV
EDTGEGSEKANDIGEPNVVETDPILESPPVVGDSTTKPSAEVVTVVKPRSPCAITAEEVKHPTPTLVNLTDRARQNAEKRLQRPAPTQPNASPPPSIPGN
RGRARGGSVRGRGVQRTFRGGRTGRGGPSPGQPGQG
|
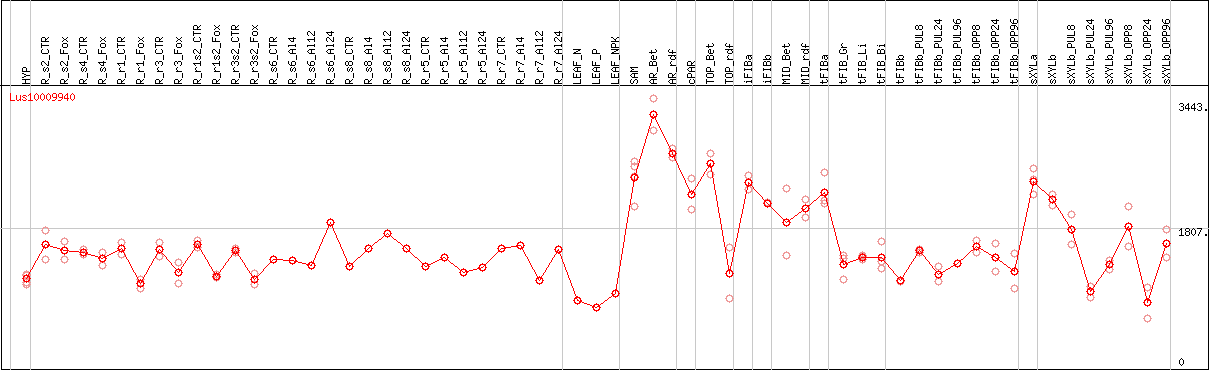
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10009940 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
