Lus10010010 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10010010 pacid=23176683 polypeptide=Lus10010010 locus=Lus10010010.g ID=Lus10010010.BGIv1.0 annot-version=v1.0
ATGTCGTCGGCAAAGAAACGATCCGCTGGCCGATCGCCGCTGGTGAATCAGCAATCGCAGATCACTTCCTTCTTCACTAAAAGGACCAACTCGTCTTCTC
CTTCGCCTTCTCCCTCGCCTTCTCCGTCTGTATTGAAACAGACAAAACTAAAACCCCAATCCAACATCAGCAACCCTAACCCTAGCTCTCCGAGTCCGTC
TCATTCACCGGTAACTCCTTCTCCGATCCAGTCCAGCTCCAAGAAGCCTCTCCTCGTCATCGGCCAAAGCCCTTCTCCGTCGACGCCTGAGGTCGGTGTG
AGAACCTACGGGAAGGAGGTGGTGGATAAGAGGATTAGGGTTTATTGGCCGCTGGACAAGGCCTGGTACGAAGGCTGTGTGAAGTCTTATGAAAAGGGTT
CGGGTAAGCATTTGATTCAGTACGATGATTCGGAAGAGGAGATGTTGGATTTGGGGAAGGAGAAGGTTGAGTGGATCGGGGAAGCTGTCAAGAAATTGAA
GCGGTTGAGGAAGGGTTCTCTGGCGGCTGTTATCAAGCAGGCGGTGGTTGAGGGAGAAGATTTGGAGGACGCTGAGGTGGAGAATGGCGGTGGTGGTGAT
AGTCATGGTGATGATTCTAGTGATGAAGATTGGGGAAAGAATGTGGTGGAAGATGTCAGCGATGGCGAGGAGGACCAGATGGATCTGGAAGAGGAGGATG
AGGATGATGCTGGAGATAGGAAGAGAGTAGGGAAGCAAGGAGTGAAGAGAGCAAAGAAGAACGACTCGCGAAAACGGAAGGGAAGCTCTGAAGTGAAGGT
AGATTCTGGTAAGAAGGGCAGGAACAATGGAGAATTGAGTACAGAAGCCTCGAAATGGCCAGCCTTGGAGCCAGTAAAGAATGTGCACACTGGAGAAACC
GTTGCATTGGGAAGATTTGAGATGCGTGAATCCGAGAAGCTGTGGTTTCTTGGAGTAAGACGTCGAGATGCAAAAAGAAGACTGCCTACAGACGGAGACT
ATGATCCTAGGACCCTATACTTGCCTCCTGATTTTGTAAGGAGTTTATCTGATGGCCAGAAGCAATGGTGGGAGTTTAAGTCCAAGCACATGGACAAGGT
TTTATTCTTCAAGATGGGAAAGTTTTATGAGCTTTTTGAAATGGATGCTCATATAGGAGCTAAAGAACTTGATTTAAACTTTTCAATGAATGTAGAGAAA
TTGGCTAGAAAGGGCTATCGAGTTCTTGTGGTGGAGCAGACTGAAACACCTGAACAGCTAGAGCGTCGTCGCAAAGAGAAAGGCTCTAGGGACAAGGTTG
TTAAACGTGAGATCTGTGCGGTTGTCACGAGGGGAACACTAACAGAGGGAGAACTGCTTTCAGCAAATCCTGATGCGTCTTATTTGATGGCACTGACTGA
AGTTTGTCAGTCAACCCAAAATTTTGAACGGGTGTTTGGCGTTTGTGTAGTTGATGTTGCAACGAGCAGAATAATTCTTGGACAGTTTGGTGACGATGCA
GACTGCAGCTCCTTGTGTTGCCTGTTGTCTGAGTTGAGGCCTGTGGAGATTATAAAACCTGCAAAATCTCTGAGTTCTGAGACTGAGAGAGTTATTTCAA
GGCATACGAGAAATGTTTTATTTAACGAGTTGGTTCCACTTTCAGAATTCTGGAGTGCCGAGAAAACTATTCATGAAGTCAAAGCTGCTTTAAAGCTCAT
CAATTACCAGTCAACTTCTCAATCAGTGAATGAGGCGACCTTGCTTACTAATGTGGAGCAAGAGGATGTGTCCTGTCTACCAGCCGTTTTGCTGCAGCTA
GTGAATAAAGGTGAAGATGGAAGCTTAGCACTCTCTGCTTTTGGAGCCACTCTCTATTATCTGAAACAGTCATTTCTGGACGAGGCATTGCTTAGATCAG
CGAAATTTGAACGACTCCCATGCTCCGAATTTTGCGATGTTGCTCAAAAACCGTATATGATTCTTGATGCTTCTGCCCTTGAAAACCTTGAGATTTTCGA
GAACAGCATAACTGGAGACTCATTTGGGACGCTATATGCTCAAATAAATCATTGTGTGACTGCGTTTGGAAAGAGGTTACTTAAAACGTGGCTTGCAAGA
CCTTTATATCATGTTAAATCTATCAAGGATCGCCAGGATGCTGTCTCAGATTTACAGGGGACTAATCAAACAATGGCACTTGAATTCCGGAAAGCATTGT
GTAGGCTGCCAGACATGGAGAGACTGCTTACCCGTATGTTTGCCTGCAGTGAAGCTAACGGGAGGAATGCAAGCAAGGTGGTACTGTATGAAGATGCGTC
AAAGAAACAGCTGCAAGAATTCACATCTGCTCTACGTGGATGTGAATTAATGGCTCAAGCATGTTCTTCACTCAGTGTCATTCTGCAAACATCAGAGTCT
AAAAAACTACGATTTTTGCTGACAGCTGGTGAAGGTCTTCCTGATATCAATTCAGATCTTAATTATTTCAAGGATGGGTTTGATTGGGTGGAAGCGAACA
ACACTGGTCGTATAATACCCCACGAGGGAGTTGATATGGTGTATGACTCTGCTTGTAAGAACGTCCATGATATTGAATCTGGTCTGAAAAAAGAATTGAA
GGAACAGCGAAAGATCCTTGGAGATGCATCTATAACTTACGTCACGGTTGGGAAAGATGCATATTTATTGGAAGTTCCAGAACATATGAGTGAAAGTATC
CCTCAAGAATATGAGCTGCGTTCATCAAAAAAGGGCGCTTGCAGGTATTGGACTCCAAATATTAAGAAACTTTTACCCGAACTGGCCCAAGCTGAATCCG
AAAAGGAGTCGGCGTTGAAGAGCATTTTGCAAAGGTTCATTGGTCGTTTTTGTCTAAATCATACCAAGTGGAGACAACTAGTTTCTGTAGTTGCAGAACT
TGATGTTTTGATCAGTCTGGCAATTGCTTGTGATTTTTATGAGGGACCAACGTGCCGCCCGGTCATCTTGGATTCTTCATCATCGGCTAGTGAAACCCCA
TATCTTTCTGCAAAAGGCTTAGGACATCCTATACTGAGAAGTGATTCCTTAGGCAAGGGATCTTTTGTTTCCAACAATATTAGCATCGGTGGTTTGAATG
ATGGAAATTTTATCCTTCTCACGGGTCCAAATATGGGTGGAAAGTCTACTCTTCTTCGACAAATTTGTTTGGCTACAGTTCTAGCGCAGATAGGAGCTGA
TGTCCCTGCTGAGAGCTTTGAGTTGTCACCTGTTGACCGGATCTTTGTCCGTATGGGTGCCAAAGATCACATTATGGCTGGCCAAAGTACATTTTTAACA
GAGCTTTCAGAAACTGCATTAATGTTGTCATCGGCAACCCGTAATTCATTTGTGGCACTTGATGAACTTGGACGCGGCACTTCAACTTCAGATGGACAAG
CAATAGCTGAAGCTGTTCTTGAACATTTTGTAAACAAGGTGCAGTGCCGTGGAATGTTCTCTACTCACTATCACAGATTAGCTGTAGACTATGAGAAGGA
TCCCAAGGTCTCCTTATGTCATATGGCATGCAAAATTGGAATTGGAGCTGGTGGAGTGGAGGAAGTAACTTTTCTGTATAGGTTAACACCCGGTGCTTGC
CCTAAGAGCTACGGTGTCAATGTTGCAAGACTGGCAGGCCTTCCTGAAAATGTCCTTCAGACAGCTGCTGCTAAGTCTAGCGAAATCGAAGCCCTCTACG
GTAAGCACGTCAAGAAAGGCGCGGTGATCAAGGAGAATCAAAGGTCAAAATATGCTGCGTTATGTGGCATCCTTAATTTGGTCAATGGCACAATAAAATC
CGATAGCTACGAGCCTTCTCTGGGTACTGGTATCAGTCACCTGATTGAACACCAGCACCAAGCAAGAACACTTCTACTCCAAAACTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10010010 pacid=23176683 polypeptide=Lus10010010 locus=Lus10010010.g ID=Lus10010010.BGIv1.0 annot-version=v1.0
MSSAKKRSAGRSPLVNQQSQITSFFTKRTNSSSPSPSPSPSPSVLKQTKLKPQSNISNPNPSSPSPSHSPVTPSPIQSSSKKPLLVIGQSPSPSTPEVGV
RTYGKEVVDKRIRVYWPLDKAWYEGCVKSYEKGSGKHLIQYDDSEEEMLDLGKEKVEWIGEAVKKLKRLRKGSLAAVIKQAVVEGEDLEDAEVENGGGGD
SHGDDSSDEDWGKNVVEDVSDGEEDQMDLEEEDEDDAGDRKRVGKQGVKRAKKNDSRKRKGSSEVKVDSGKKGRNNGELSTEASKWPALEPVKNVHTGET
VALGRFEMRESEKLWFLGVRRRDAKRRLPTDGDYDPRTLYLPPDFVRSLSDGQKQWWEFKSKHMDKVLFFKMGKFYELFEMDAHIGAKELDLNFSMNVEK
LARKGYRVLVVEQTETPEQLERRRKEKGSRDKVVKREICAVVTRGTLTEGELLSANPDASYLMALTEVCQSTQNFERVFGVCVVDVATSRIILGQFGDDA
DCSSLCCLLSELRPVEIIKPAKSLSSETERVISRHTRNVLFNELVPLSEFWSAEKTIHEVKAALKLINYQSTSQSVNEATLLTNVEQEDVSCLPAVLLQL
VNKGEDGSLALSAFGATLYYLKQSFLDEALLRSAKFERLPCSEFCDVAQKPYMILDASALENLEIFENSITGDSFGTLYAQINHCVTAFGKRLLKTWLAR
PLYHVKSIKDRQDAVSDLQGTNQTMALEFRKALCRLPDMERLLTRMFACSEANGRNASKVVLYEDASKKQLQEFTSALRGCELMAQACSSLSVILQTSES
KKLRFLLTAGEGLPDINSDLNYFKDGFDWVEANNTGRIIPHEGVDMVYDSACKNVHDIESGLKKELKEQRKILGDASITYVTVGKDAYLLEVPEHMSESI
PQEYELRSSKKGACRYWTPNIKKLLPELAQAESEKESALKSILQRFIGRFCLNHTKWRQLVSVVAELDVLISLAIACDFYEGPTCRPVILDSSSSASETP
YLSAKGLGHPILRSDSLGKGSFVSNNISIGGLNDGNFILLTGPNMGGKSTLLRQICLATVLAQIGADVPAESFELSPVDRIFVRMGAKDHIMAGQSTFLT
ELSETALMLSSATRNSFVALDELGRGTSTSDGQAIAEAVLEHFVNKVQCRGMFSTHYHRLAVDYEKDPKVSLCHMACKIGIGAGGVEEVTFLYRLTPGAC
PKSYGVNVARLAGLPENVLQTAAAKSSEIEALYGKHVKKGAVIKENQRSKYAALCGILNLVNGTIKSDSYEPSLGTGISHLIEHQHQARTLLLQN
|
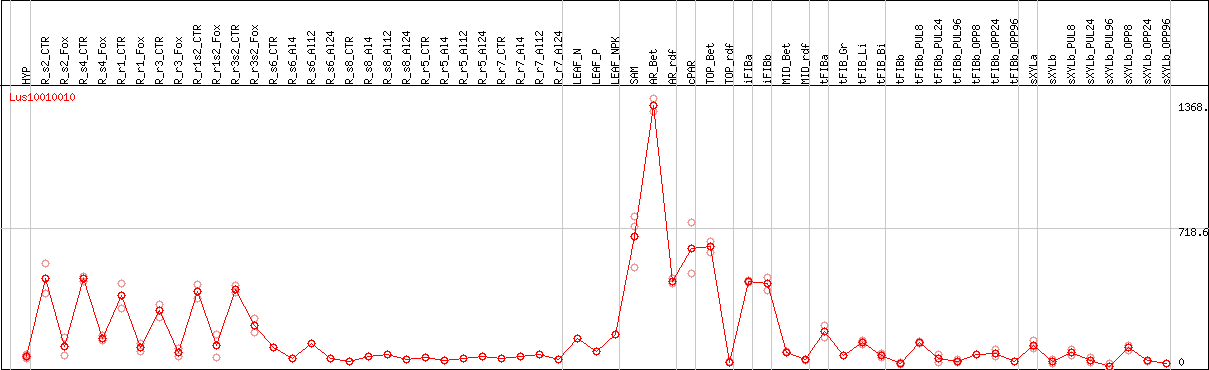
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010010 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
