External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G08720 127 / 4e-35
ATPK2, ATPK19, ATS6K2, S6K2
ARABIDOPSIS THALIANA SERINE/THREONINE PROTEIN KINASE 2, ARABIDOPSIS THALIANA PROTEIN KINASE 2, Arabidopsis thaliana protein kinase 19, serine/threonine protein kinase 2 (.1.2)
AT3G08730 122 / 2e-33
ATS6K1, ATPK6, ATPK1
P70 RIBOSOMAL S6 KINASE, ARABIDOPSIS THALIANA PROTEIN-SERINE KINASE 6, ARABIDOPSIS THALIANA PROTEIN-SERINE KINASE 1, protein-serine kinase 1 (.1)
AT5G04510 75 / 2e-16
ATPDK1, PDK1
3'-phosphoinositide-dependent protein kinase 1 (.1.2)
AT3G10540 74 / 8e-16
3-phosphoinositide-dependent protein kinase (.1)
AT1G45160 71 / 7e-15
Protein kinase superfamily protein (.1.2)
AT1G30640 70 / 2e-14
Protein kinase family protein (.1)
AT1G03920 69 / 4e-14
Protein kinase family protein (.1)
AT4G24400 66 / 4e-13
ATCIPK8, PKS11, CIPK8, SnRK3.13
SNF1-RELATED PROTEIN KINASE 3.13, PROTEIN KINASE 11, CBL-interacting protein kinase 8 (.1.2)
AT4G33080 63 / 3e-12
AGC (cAMP-dependent, cGMP-dependent and protein kinase C) kinase family protein (.1), AGC (cAMP-dependent, cGMP-dependent and protein kinase C) kinase family protein (.2)
AT5G09890 63 / 5e-12
Protein kinase family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033125
181 / 6e-55
AT3G08720 515 / 1e-179
ARABIDOPSIS THALIANA SERINE/THREONINE PROTEIN KINASE 2, ARABIDOPSIS THALIANA PROTEIN KINASE 2, Arabidopsis thaliana protein kinase 19, serine/threonine protein kinase 2 (.1.2)
Lus10036653
182 / 7e-55
AT3G08720 515 / 1e-178
ARABIDOPSIS THALIANA SERINE/THREONINE PROTEIN KINASE 2, ARABIDOPSIS THALIANA PROTEIN KINASE 2, Arabidopsis thaliana protein kinase 19, serine/threonine protein kinase 2 (.1.2)
Lus10025213
127 / 2e-34
AT3G08720 570 / 0.0
ARABIDOPSIS THALIANA SERINE/THREONINE PROTEIN KINASE 2, ARABIDOPSIS THALIANA PROTEIN KINASE 2, Arabidopsis thaliana protein kinase 19, serine/threonine protein kinase 2 (.1.2)
Lus10021386
115 / 4e-32
AT3G08720 445 / 4e-157
ARABIDOPSIS THALIANA SERINE/THREONINE PROTEIN KINASE 2, ARABIDOPSIS THALIANA PROTEIN KINASE 2, Arabidopsis thaliana protein kinase 19, serine/threonine protein kinase 2 (.1.2)
Lus10042337
73 / 9e-16
AT5G41310 96 / 1e-21
P-loop nucleoside triphosphate hydrolases superfamily protein with CH (Calponin Homology) domain (.1)
Lus10027086
72 / 2e-15
AT2G43940 163 / 5e-46
HARMLESS TO OZONE LAYER 3, S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10028834
70 / 1e-14
AT3G10540 817 / 0.0
3-phosphoinositide-dependent protein kinase (.1)
Lus10017448
69 / 3e-14
AT3G10540 821 / 0.0
3-phosphoinositide-dependent protein kinase (.1)
Lus10005470
68 / 1e-13
AT1G45160 977 / 0.0
Protein kinase superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G073700
147 / 6e-43
AT3G08720 483 / 1e-168
ARABIDOPSIS THALIANA SERINE/THREONINE PROTEIN KINASE 2, ARABIDOPSIS THALIANA PROTEIN KINASE 2, Arabidopsis thaliana protein kinase 19, serine/threonine protein kinase 2 (.1.2)
Potri.006G109600
125 / 2e-34
AT3G08720 637 / 0.0
ARABIDOPSIS THALIANA SERINE/THREONINE PROTEIN KINASE 2, ARABIDOPSIS THALIANA PROTEIN KINASE 2, Arabidopsis thaliana protein kinase 19, serine/threonine protein kinase 2 (.1.2)
Potri.016G138400
120 / 9e-33
AT3G08720 639 / 0.0
ARABIDOPSIS THALIANA SERINE/THREONINE PROTEIN KINASE 2, ARABIDOPSIS THALIANA PROTEIN KINASE 2, Arabidopsis thaliana protein kinase 19, serine/threonine protein kinase 2 (.1.2)
Potri.008G028800
77 / 5e-17
AT3G10540 809 / 0.0
3-phosphoinositide-dependent protein kinase (.1)
Potri.010G232800
76 / 2e-16
AT3G10540 834 / 0.0
3-phosphoinositide-dependent protein kinase (.1)
Potri.002G263000
73 / 2e-15
AT1G45160 1162 / 0.0
Protein kinase superfamily protein (.1.2)
Potri.004G209700
63 / 4e-12
AT5G58140 1332 / 0.0
NON PHOTOTROPIC HYPOCOTYL 1-LIKE, phototropin 2 (.1.2.3.4)
Potri.006G068400
63 / 5e-12
AT5G35410 686 / 0.0
SNF1-RELATED PROTEIN KINASE 3.11, CBL-INTERACTING PROTEIN KINASE 24, SALT OVERLY SENSITIVE 2, Protein kinase superfamily protein (.1)
Potri.006G224800
62 / 5e-12
AT4G33080 832 / 0.0
AGC (cAMP-dependent, cGMP-dependent and protein kinase C) kinase family protein (.1), AGC (cAMP-dependent, cGMP-dependent and protein kinase C) kinase family protein (.2)
Potri.002G125800
62 / 9e-12
AT5G09890 828 / 0.0
Protein kinase family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF00069
Pkinase
Protein kinase domain
Representative CDS sequence
>Lus10010054 pacid=23155625 polypeptide=Lus10010054 locus=Lus10010054.g ID=Lus10010054.BGIv1.0 annot-version=v1.0
ATGTTGAGAACGATACCTCAGGGGAAGTTTGAGAAGGTGTTCATGGTGCAAGGGAGCAAGCCTCAACTTGGCAATGGCCATGTCATTTTTGCAATGAAAG
AGATGAGCAGGGATAATATAATCAAGAAGAAGCACGTGGATTATATAAGAGTTGACAAATATGTTCTCACTGAACTCGTCTATCCTTTCTCCATCGAACT
TCCTTACTCTTTTTGTACCAACACAAAGCTAAATTTGATCATCTATGTCATGTACAGAGGCCATCTCTTCTTCCACATCTATCGCCAAGGGATCTTCAGT
GAGGATCGGTTGAAGCTTAGTGCTGCTGAATTTGTGTCTTTTGTTTCACATTTCCATGATTACTGGATCGTGCATCAAGATTTCGGGCCTGGAAATATTC
TCTCGAGTGTCGAAGGATATGTTGTGCTACTTGATTTTGCAAAGGAGATGAATGAATCCAGTAGGTAG
AA sequence
>Lus10010054 pacid=23155625 polypeptide=Lus10010054 locus=Lus10010054.g ID=Lus10010054.BGIv1.0 annot-version=v1.0
MLRTIPQGKFEKVFMVQGSKPQLGNGHVIFAMKEMSRDNIIKKKHVDYIRVDKYVLTELVYPFSIELPYSFCTNTKLNLIIYVMYRGHLFFHIYRQGIFS
EDRLKLSAAEFVSFVSHFHDYWIVHQDFGPGNILSSVEGYVVLLDFAKEMNESSR
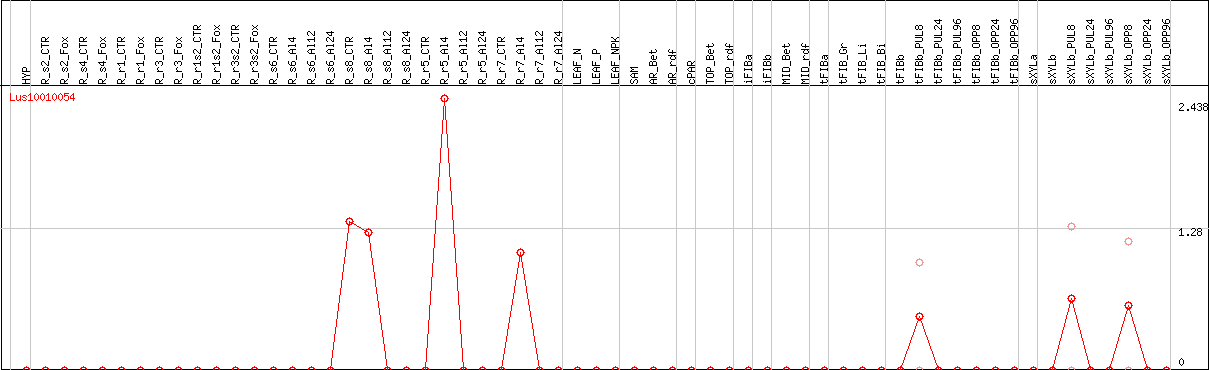
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010054 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.