Lus10010069 [FLAX]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10010069 pacid=23158855 polypeptide=Lus10010069 locus=Lus10010069.g ID=Lus10010069.BGIv1.0 annot-version=v1.0
ATGGATGATAGTTGTGCTGTTTGCGCTGATTCCTTGGAATGGGTCAGTTATGGTGCTTGTGGCCATCGCGAGGTCTGCTCTACCTGCGTCGCTCGCCTCC
GTTTCATCATCGAGGATCGACGATGCTGCATCTGCAAGGCGGAATCCGATGTTGTCTTTGTCACCAAGGCTCTTGGAGACTATACGAGGATGATTAATGA
CTTTTCAGTATTGCCATCTGAACCAAGGGAGGGTCGGGTTGGGTCATATTGGTATCATGAGGATACCCAGGCATTTTTTGACGACGTGGAACACTACAAG
ATGGTTAAGGCAATGTGCAAACTCTCATGCAGTGTATGCGACAAGGCGGAGGATGAGACTAACGATGGTGCCAAAAGGCGCTCCAAGTTTAGGAACATTG
AGCATCTGAAGGGTCATTTGTTTCACAAACATAAATTGGTTATGTGTAGCCTATGTTTGGAAGGGAGGAAGGTATTTATTTGTGAGCAAAAGCTATATAC
TAGATCACAGCTTAGTCAGCACATCAGCACAGGGGACTCTGAGGTTGATGGTAGTGAAAGTGAAAGAGGCGGGTTTAGGGGTCATCCCATGTGTGAATTT
TGCAAGACTCCATTCTATGGCGAAAATGAGTTGTACTCCCATATGTCCAGAGATCACTACACCTGCCATATATGCCAAAGGCAGCACCCTGGACAATATG
TATATTATAAAAACTACCAGGACCTAGAAATGCACTTTCGTCATGATCATCTTCTCTGTGAGGATGAGTCGTGCCTTGCCAAAAAGTTTGTTGTTTTTCA
AAGTGAAGCAGAATTAAAGAGGCATAATACCATTGAACATGGTGGCCGGATGTCCCGTTCGAAGCGTAATGCTGCTCTACAGGCATCTCATGACATAAGC
CGTTTATTGCATTTTCAGATACCAACAAGCTTTAGGTTTCACCGTGAACAAGATCCTCGTCGTGGAAGGGGACGTACATCTCGGCGTGATGAACCTGATA
ATCAGCTTTCTATGGAAATACAAGCAAGTATGGATGCTTCCTTCCCTAGAGGAACGTCTCATGACCCCCCATTGCCATCATCTGGTACCCAAGTTACGGA
TCTTGGTTATGATATGGATCCAGACGTTCAGTCTTTAGAATCATTAGCTTTAACAGAATCTACCCCTTCAAGGTATCTTCAAGCCTTAGGGCACAGTTCT
AGGAATGGAGCTTTGCAAGAATCCGCCTTTCCTCCACTTGCTATGATTCCTAGTAGTAGCCAGCAGAAGCCCACACAAGAACCAGAAGGGTTACCAAATA
CGACTATGGCAACTCATCTCAGACGCAAGAACAATAAGGTTACTGTTATTAGCTCACCCCAGGGATGGGCGACAAGTAATGGCCCTGTTTCCAGTAGTTC
TACACAGTGGCCTGCCATAAATACTGCACCGTCAATCTCACAGAGTACTGACAGTGCTCCTTCAGTTTCCAGTCATGCAAGTCCTCCTGTTCGACCAGGG
CATGCTCCTGGTCCTCCTGGACATCAAGGGAGTTCTACACAGTGGCCTGCCATAAATAATGCACCATCAATCCCACACAGTACTTTCAGTTCTCCTTCAG
TTTCCATAGTGCATGCTCCTGGAAATCAAGGGAGAACCAGTAGAATGAGTCATTCTGCATCAGCTCCAAGTCTTGCTGAGAGTAGACTATCAAAGCCAAC
CGTTTCAGATTTCCCACCAGTTTCAGCAACACAAGTACGGAAAACAACCCCTGCTGTGCAAGTTGTGCCAAATGTTGAAGACGTTCAAACTGCAAACAGG
TCTTTGGTTGAAAAAATTCGTGATGCCCTTCAGCATGACGAGGCTAAGTATGCTTCTTTCAAAGAAATCTCTGGTCAGTATCGTCAGGGTTCCATTAGCA
CTCCAAGGTACTTGGATAACGTGAGGCAGTTTGGATTGTCGCATCTTGTTATCGATCTGGCTAGTCTATGTCCTGATCCTCAGAAGCGCCAGGAGTTAGT
TGGCACCTATTTCGATAGTTTGGAGAGTAACGGCCAGCAGGAAAAGGTTGGTGGCCGCCGGGAAACCCAATTGCAGGAGAGTAAAGGTAAAAAGGGCAAG
GGGATAGCTACTGAAGGCAGTAGCTCAAGGGACAGATTGGCCGACAATATTATCAGCACTGTTAGGACACTACAGTCAAACCACAAGCCTCAGGAGGAAG
AGGTTGAGGTGTTGTCCAAGGATGGTTACCGGAATTCGAAAGGCAATTTAGTGGCTGATCAACGAAGTTCCAAGCAGGGGGGTCAACGTGACTCTACATC
TGGTGATGTATCCAAGTATAAGAAGAAAACTCCAAAGTTCCTCAGAGCGAGACTGGGTGATGGGTCAATGGCGGCGATGTTAGACACGAGAGATTCTGAA
CCAGACCTGGATCCAGGTTCCCTTTCTGAAACATCAGATAGTAATGGTTTGCCTGTTCGCGGGGTGTGGCGAAAGGGAGGAAGTAAGAAACTGTTTGCTT
AA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Lus10010069 pacid=23158855 polypeptide=Lus10010069 locus=Lus10010069.g ID=Lus10010069.BGIv1.0 annot-version=v1.0
MDDSCAVCADSLEWVSYGACGHREVCSTCVARLRFIIEDRRCCICKAESDVVFVTKALGDYTRMINDFSVLPSEPREGRVGSYWYHEDTQAFFDDVEHYK
MVKAMCKLSCSVCDKAEDETNDGAKRRSKFRNIEHLKGHLFHKHKLVMCSLCLEGRKVFICEQKLYTRSQLSQHISTGDSEVDGSESERGGFRGHPMCEF
CKTPFYGENELYSHMSRDHYTCHICQRQHPGQYVYYKNYQDLEMHFRHDHLLCEDESCLAKKFVVFQSEAELKRHNTIEHGGRMSRSKRNAALQASHDIS
RLLHFQIPTSFRFHREQDPRRGRGRTSRRDEPDNQLSMEIQASMDASFPRGTSHDPPLPSSGTQVTDLGYDMDPDVQSLESLALTESTPSRYLQALGHSS
RNGALQESAFPPLAMIPSSSQQKPTQEPEGLPNTTMATHLRRKNNKVTVISSPQGWATSNGPVSSSSTQWPAINTAPSISQSTDSAPSVSSHASPPVRPG
HAPGPPGHQGSSTQWPAINNAPSIPHSTFSSPSVSIVHAPGNQGRTSRMSHSASAPSLAESRLSKPTVSDFPPVSATQVRKTTPAVQVVPNVEDVQTANR
SLVEKIRDALQHDEAKYASFKEISGQYRQGSISTPRYLDNVRQFGLSHLVIDLASLCPDPQKRQELVGTYFDSLESNGQQEKVGGRRETQLQESKGKKGK
GIATEGSSSRDRLADNIISTVRTLQSNHKPQEEEVEVLSKDGYRNSKGNLVADQRSSKQGGQRDSTSGDVSKYKKKTPKFLRARLGDGSMAAMLDTRDSE
PDLDPGSLSETSDSNGLPVRGVWRKGGSKKLFA
|
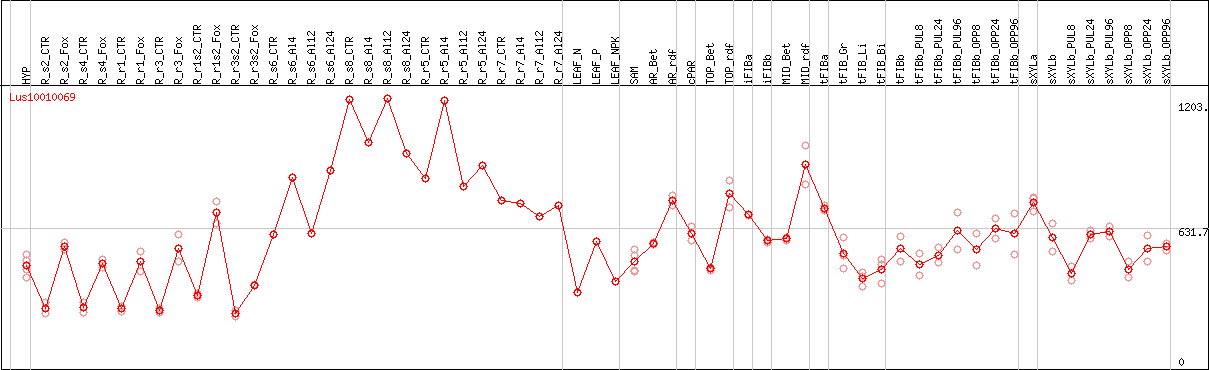
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010069 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
