External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G46760 154 / 1e-44
D-arabinono-1,4-lactone oxidase family protein (.1)
AT2G46740 153 / 2e-44
D-arabinono-1,4-lactone oxidase family protein (.1)
AT2G46750 147 / 2e-42
D-arabinono-1,4-lactone oxidase family protein (.1)
AT5G56490 139 / 4e-39
D-arabinono-1,4-lactone oxidase family protein (.1)
AT1G32300 137 / 2e-38
D-arabinono-1,4-lactone oxidase family protein (.1)
AT5G11540 102 / 5e-26
D-arabinono-1,4-lactone oxidase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002688
127 / 7e-35
AT2G46760 674 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
Lus10030206
126 / 2e-34
AT2G46760 657 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
Lus10004735
120 / 2e-32
AT2G46760 653 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
Lus10012610
115 / 1e-30
AT2G46760 761 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
Lus10007795
115 / 2e-30
AT5G47380 562 / 0.0
Protein of unknown function, DUF547 (.1)
Lus10010105
94 / 4e-23
AT2G46760 709 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
Lus10038620
90 / 1e-21
AT5G11540 732 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
Lus10012609
52 / 3e-08
AT2G46760 642 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G178300
160 / 7e-47
AT2G46760 785 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
Potri.002G178400
157 / 1e-45
AT2G46760 809 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
Potri.018G039600
83 / 3e-19
AT5G11540 771 / 0.0
D-arabinono-1,4-lactone oxidase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0077
FAD_PCMH
PF01565
FAD_binding_4
FAD binding domain
Representative CDS sequence
>Lus10010106 pacid=23158866 polypeptide=Lus10010106 locus=Lus10010106.g ID=Lus10010106.BGIv1.0 annot-version=v1.0
ATGATCATCATTATCATCATCATCATCTCTCTCCGACCAGCAAGCTGCACACCTCCGGAAGAGCCAATCAAGTGCACCTCCACAAGCAACACCAACTGCA
CCATCACAAACTCCTACGGCGCATTCCCTGACCGTTCCACGTGTCGCGCATCCAACGCCGCCTACCCAACCACCGAGCAAGAACTCATCGACATCGTCCA
AAACGCCACGAGGGCTCAACGGAAGATGAAAGTGGCCACAAGATACTCCCACAGCATCCCCAAGCTGGTCTGTCCCGACGGCGACGACGGCCTCCTCATA
AGCACCAAGTTCCTCAACCGGGTGTTGGCGGTGGACAGTGCGGCGATGACGATGAGCGTGGAGAGTGGGGTGACGCTACGCCAGCTGGTGGAGGAGGCGG
CTAAGGCAGGGATGGCTGACGCTGCGGCAGCTGGTGGGAGAGGCGGCTAA
AA sequence
>Lus10010106 pacid=23158866 polypeptide=Lus10010106 locus=Lus10010106.g ID=Lus10010106.BGIv1.0 annot-version=v1.0
MIIIIIIIISLRPASCTPPEEPIKCTSTSNTNCTITNSYGAFPDRSTCRASNAAYPTTEQELIDIVQNATRAQRKMKVATRYSHSIPKLVCPDGDDGLLI
STKFLNRVLAVDSAAMTMSVESGVTLRQLVEEAAKAGMADAAAAGGRGG
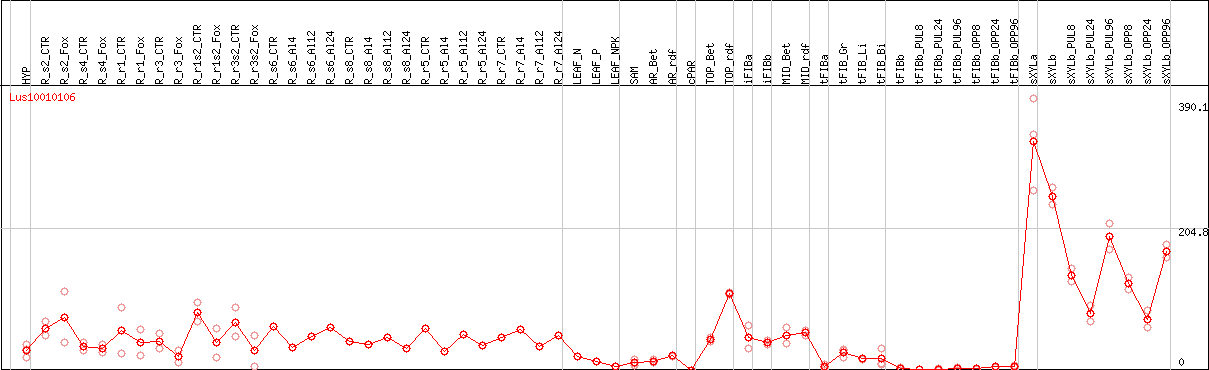
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010106 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.