External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G23190 717 / 0
CYP86B1
"cytochrome P450, family 86, subfamily B, polypeptide 1", cytochrome P450, family 86, subfamily B, polypeptide 1 (.1)
AT5G08250 697 / 0
Cytochrome P450 superfamily protein (.1)
AT1G24540 476 / 4e-164
CYP86C1
"cytochrome P450, family 86, subfamily C, polypeptide 1", cytochrome P450, family 86, subfamily C, polypeptide 1 (.1)
AT3G26125 466 / 2e-159
CYP86C2
"cytochrome P450, family 86, subfamily C, polypeptide 2", cytochrome P450, family 86, subfamily C, polypeptide 2 (.1)
AT1G13140 459 / 4e-157
CYP86C3
"cytochrome P450, family 86, subfamily C, polypeptide 3", cytochrome P450, family 86, subfamily C, polypeptide 3 (.1.2)
AT1G01600 453 / 2e-154
CYP86A4
"cytochrome P450, family 86, subfamily A, polypeptide 4", cytochrome P450, family 86, subfamily A, polypeptide 4 (.1)
AT1G63710 449 / 2e-153
CYP86A7
"cytochrome P450, family 86, subfamily A, polypeptide 7", cytochrome P450, family 86, subfamily A, polypeptide 7 (.1)
AT2G45970 447 / 2e-152
CYP86A8, LCR
LACERATA, "cytochrome P450, family 86, subfamily A, polypeptide 8", cytochrome P450, family 86, subfamily A, polypeptide 8 (.1)
AT4G00360 446 / 7e-152
ATT1, CYP86A2
"cytochrome P450, family 86, subfamily A, polypeptide 2", ABERRANT INDUCTION OF TYPE THREE 1, cytochrome P450, family 86, subfamily A, polypeptide 2 (.1)
AT1G13150 444 / 2e-151
CYP86C4
"cytochrome P450, family 86, subfamily C, polypeptide 4", cytochrome P450, family 86, subfamily C, polypeptide 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017394
1023 / 0
AT5G23190 739 / 0.0
"cytochrome P450, family 86, subfamily B, polypeptide 1", cytochrome P450, family 86, subfamily B, polypeptide 1 (.1)
Lus10040986
771 / 0
AT5G23190 751 / 0.0
"cytochrome P450, family 86, subfamily B, polypeptide 1", cytochrome P450, family 86, subfamily B, polypeptide 1 (.1)
Lus10032278
429 / 1e-144
AT1G63710 792 / 0.0
"cytochrome P450, family 86, subfamily A, polypeptide 7", cytochrome P450, family 86, subfamily A, polypeptide 7 (.1)
Lus10006232
413 / 5e-139
AT1G24540 524 / 0.0
"cytochrome P450, family 86, subfamily C, polypeptide 1", cytochrome P450, family 86, subfamily C, polypeptide 1 (.1)
Lus10018217
409 / 7e-138
AT5G58860 802 / 0.0
"cytochrome P450, family 86, subfamily A, polypeptide 1", cytochrome P450, family 86, subfamily A, polypeptide 1 (.1)
Lus10040687
409 / 1e-137
AT5G58860 803 / 0.0
"cytochrome P450, family 86, subfamily A, polypeptide 1", cytochrome P450, family 86, subfamily A, polypeptide 1 (.1)
Lus10014768
424 / 2e-136
AT4G00360 756 / 0.0
"cytochrome P450, family 86, subfamily A, polypeptide 2", ABERRANT INDUCTION OF TYPE THREE 1, cytochrome P450, family 86, subfamily A, polypeptide 2 (.1)
Lus10003669
402 / 8e-135
AT2G23180 440 / 4e-150
"cytochrome P450, family 96, subfamily A, polypeptide 1", cytochrome P450, family 96, subfamily A, polypeptide 1 (.1)
Lus10002527
395 / 1e-132
AT2G23180 454 / 7e-156
"cytochrome P450, family 96, subfamily A, polypeptide 1", cytochrome P450, family 96, subfamily A, polypeptide 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G072100
736 / 0
AT5G23190 799 / 0.0
"cytochrome P450, family 86, subfamily B, polypeptide 1", cytochrome P450, family 86, subfamily B, polypeptide 1 (.1)
Potri.005G092200
728 / 0
AT5G23190 788 / 0.0
"cytochrome P450, family 86, subfamily B, polypeptide 1", cytochrome P450, family 86, subfamily B, polypeptide 1 (.1)
Potri.010G050100
501 / 2e-173
AT1G24540 684 / 0.0
"cytochrome P450, family 86, subfamily C, polypeptide 1", cytochrome P450, family 86, subfamily C, polypeptide 1 (.1)
Potri.008G183300
491 / 7e-170
AT1G24540 682 / 0.0
"cytochrome P450, family 86, subfamily C, polypeptide 1", cytochrome P450, family 86, subfamily C, polypeptide 1 (.1)
Potri.014G085800
458 / 3e-156
AT2G45970 866 / 0.0
LACERATA, "cytochrome P450, family 86, subfamily A, polypeptide 8", cytochrome P450, family 86, subfamily A, polypeptide 8 (.1)
Potri.003G129100
427 / 2e-144
AT1G63710 823 / 0.0
"cytochrome P450, family 86, subfamily A, polypeptide 7", cytochrome P450, family 86, subfamily A, polypeptide 7 (.1)
Potri.009G043700
422 / 7e-143
AT5G58860 827 / 0.0
"cytochrome P450, family 86, subfamily A, polypeptide 1", cytochrome P450, family 86, subfamily A, polypeptide 1 (.1)
Potri.001G249700
414 / 6e-140
AT5G58860 790 / 0.0
"cytochrome P450, family 86, subfamily A, polypeptide 1", cytochrome P450, family 86, subfamily A, polypeptide 1 (.1)
Potri.015G132800
398 / 1e-133
AT4G39490 589 / 0.0
"cytochrome P450, family 96, subfamily A, polypeptide 10", cytochrome P450, family 96, subfamily A, polypeptide 10 (.1)
Potri.007G080300
398 / 2e-133
AT4G39490 574 / 0.0
"cytochrome P450, family 96, subfamily A, polypeptide 10", cytochrome P450, family 96, subfamily A, polypeptide 10 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00067
p450
Cytochrome P450
Representative CDS sequence
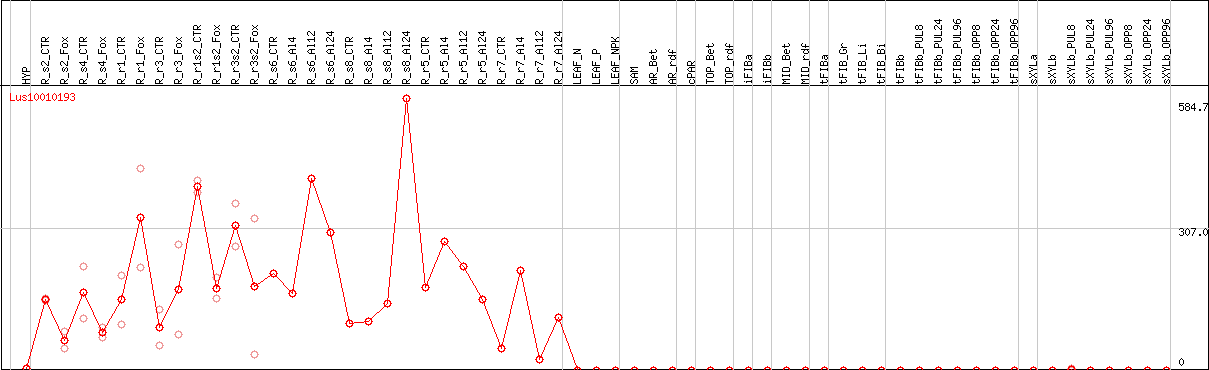
>Lus10010193 pacid=23154218 polypeptide=Lus10010193 locus=Lus10010193.g ID=Lus10010193.BGIv1.0 annot-version=v1.0
ATGGCACTTCTCATTTCCACCACCGCCGGAACACCGCCCACCACCGCCACCGTCCATACCATCCTCCGCGACATCCACCTCCTCGAGCTCCTGGTCGCCC
TCCTCGTATTCGTCACCATACACTCCCTCCGCAAAGCCCACCGCCACGGCCTCCCCATATGGCCCTTCCTAGGCATGCTCCCTTCTCTATTCTCCGCCTT
ACAAGGCGACAAGAACGTCTACGAATGGCTCACCGACGTCCTCCGACGCCTCAACGGCACCTTCCTATTCAAAGGCCCGTCTTTAAGCTCCCTCAACTGC
GTCGTCACTTCCGACCCTCGCAACCTCGAGTACTTGCTCAAGACAAAATTCTCCAACTTCCCTAAGGGGCCCTACTTCCGCGGCAACCTCCGTGACCTTC
TGGGCGAGGGCATCTTCAACGCGGATGGAGCCACTTGGAACAAGCAACGCAAGGCTGCGAGCCTCGAGTTTCATTCTTCGAGGTTCCGGAAGCTTACTTA
CTCGTCGTTGTTCGAGCTCGTGCATGCTAGGTTGCTCCCTGTGCTTCTTGATGCCTCGTCCTCGAAACGAAACGTTTCGATCGATCTCCAGGATATATTG
TTGAGGCTCACATTCGACAACGTGTGCATGCTCGCATTCGGAACAGACCCCGGCTGCCTAAGCCAGGGTCTCCCCGAGGTACAATTCGCCCGTGCATTTG
AGGACGCCACAGAAGCCACCGCACTACGATTCGTGATGCCAACGTGTCTATGGAAGGCAATGAGGTGGATGAACATCGGAAGCGAAAGAGTGCTCCGAAA
GTCGATCGAAGGAGTGGACGATTTCGCGGAGGAAGTGATCCGAGCTCGGAAGAAGGAGCTGGCATTACTGGACGACGACGTGGACAGCCACCTGGCCATC
AAGAGGTCCGACATACTGACGGTGTTCATGGGGATGAAGGACGAGGAAGGCAAAGCGTATACGGAGAAGTTTTTAAAGGACATATGTGTCAACTTTATAT
TGGCTGGGAGGGACACGTCATCGGTGGCGCTGAGCTGGTTTTTCTGGCTGGTTGATCGGCATCCGGAGGTGGAAGATAAGATTCTGAAGGAGATTCGGAG
CGTAGTGAAGGAGAGGGAACCGGAGATGATCGACGGTGGAGATGAAGTGATGCTGAAGCCTGAGGAGATCAAGAGGATGGATTATCTACAGGGTGCTCTG
TCTGAAGCTCTCAGGCTGTATCCTTCTGTTCCGGTCGATCACAAGGAGGTGCTGGAGGATGACGTGTTCCCAGACGGTACCGTTTTGAAGAAAGGGACCA
AGGTGCTGTACGCAATCTACTCAATGGGGAGGATGGAGTCAATCTGGGGGGAAGACTGCAGGGAGTTCAAGCCAGAGAGGTGGCTGAGGGACGGCCGGTT
CACGAGCGAATCGGCCTACAAGTTCACGGCTTTCAACGGAGGGTCCAGGCTGTGCCTGGGGAAGGACTTCGCTTACTACCAGATGAAGTTCGTGGCGGCT
TCGATCATGTACCGGTTCAAGGTTAGAGTTGTCGAGGGTCACCCTGTCACTCCGAAGCTCGCCTTGACCATGTACATGAAACACGGGCTCAAGGTGAATC
TGTACAGGCGTGGGGGTGGTGTTGGTGATGATGAAACATCATGA
AA sequence
>Lus10010193 pacid=23154218 polypeptide=Lus10010193 locus=Lus10010193.g ID=Lus10010193.BGIv1.0 annot-version=v1.0
MALLISTTAGTPPTTATVHTILRDIHLLELLVALLVFVTIHSLRKAHRHGLPIWPFLGMLPSLFSALQGDKNVYEWLTDVLRRLNGTFLFKGPSLSSLNC
VVTSDPRNLEYLLKTKFSNFPKGPYFRGNLRDLLGEGIFNADGATWNKQRKAASLEFHSSRFRKLTYSSLFELVHARLLPVLLDASSSKRNVSIDLQDIL
LRLTFDNVCMLAFGTDPGCLSQGLPEVQFARAFEDATEATALRFVMPTCLWKAMRWMNIGSERVLRKSIEGVDDFAEEVIRARKKELALLDDDVDSHLAI
KRSDILTVFMGMKDEEGKAYTEKFLKDICVNFILAGRDTSSVALSWFFWLVDRHPEVEDKILKEIRSVVKEREPEMIDGGDEVMLKPEEIKRMDYLQGAL
SEALRLYPSVPVDHKEVLEDDVFPDGTVLKKGTKVLYAIYSMGRMESIWGEDCREFKPERWLRDGRFTSESAYKFTAFNGGSRLCLGKDFAYYQMKFVAA
SIMYRFKVRVVEGHPVTPKLALTMYMKHGLKVNLYRRGGGVGDDETS