External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G55340 254 / 9e-87
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
AT3G03880 176 / 6e-56
Protein of unknown function (DUF1639) (.1)
AT4G20300 99 / 1e-24
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
AT3G60410 65 / 2e-12
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2), Protein of unknown function (DUF1639) (.3)
AT1G68340 56 / 1e-09
Protein of unknown function (DUF1639) (.1)
AT1G25370 54 / 1e-08
Protein of unknown function (DUF1639) (.1)
AT4G17440 47 / 2e-06
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013412
289 / 2e-101
AT1G55340 165 / 1e-52
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10010650
137 / 1e-41
AT1G55340 148 / 3e-46
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10013616
102 / 1e-28
AT1G55340 111 / 1e-32
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10038379
96 / 3e-24
AT4G20300 188 / 1e-58
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10036240
95 / 5e-23
AT4G20300 217 / 3e-67
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10035144
60 / 5e-11
AT1G25370 87 / 2e-20
Protein of unknown function (DUF1639) (.1)
Lus10004378
57 / 1e-09
AT4G17440 96 / 4e-23
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10010984
52 / 3e-08
AT3G60410 89 / 3e-20
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2), Protein of unknown function (DUF1639) (.3)
Lus10040176
43 / 6e-05
AT3G60410 76 / 1e-16
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2), Protein of unknown function (DUF1639) (.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G004000
286 / 1e-99
AT1G55340 268 / 4e-92
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.003G221000
279 / 1e-96
AT1G55340 261 / 3e-89
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.019G035300
230 / 6e-77
AT1G55340 232 / 9e-78
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.013G057600
214 / 9e-71
AT1G55340 212 / 1e-69
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.018G059200
124 / 8e-35
AT1G55340 133 / 3e-38
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.006G233100
120 / 3e-33
AT1G55340 130 / 9e-37
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.003G157500
104 / 1e-26
AT4G20300 201 / 2e-61
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.001G073400
103 / 2e-26
AT4G20300 226 / 3e-71
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.010G122700
65 / 1e-12
AT1G25370 110 / 1e-28
Protein of unknown function (DUF1639) (.1)
Potri.008G122600
59 / 3e-10
AT1G25370 100 / 6e-25
Protein of unknown function (DUF1639) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF07797
DUF1639
Protein of unknown function (DUF1639)
Representative CDS sequence
>Lus10010312 pacid=23169514 polypeptide=Lus10010312 locus=Lus10010312.g ID=Lus10010312.BGIv1.0 annot-version=v1.0
ATGGACAAGGAGAAAAAGAATTCAGAGAATGATTTTGTGTTACAATGGGGAACCAAAAAGAGGCTCAGATGCGTGAAATTGAAGAAAAATCAAAACTTGG
CCAACAGTTCAAGACCAACTGATTCTTTCCCCAAGAAGAGGCTCACTTCTGCTGTCGTCTCTTCAAAGAAAGATTCTCCTTCTCGTTTTATTAAGAACTC
CGATATGTTAGCAAGCAGGAGGAAAGTATCGTCACCGGAGAAAGAAGACCGGTACTACACAACTAGGGGATCAATGGGACTGGATGAAAGCAGTAAGCTT
ATATTCGATAGTGTGAAGTTGGAAGACAAAGGTCATGTCTGGCCTAAATTGTTCATCTCGCTGTCAAATAAAGAAAAGGAGGAGGATTTCATGGCAATGA
AAGGATGTAAACCATCACAGAGGCCTAAGAAAAGAGCCAAGTTGATCCAACGAACGATACTCTTGGTGAGCCCAGGGACATGGCTGTCAGATTTGACTCA
AGAGAGATATGAAGTCAGGGAGAAGAAAGCTGCCAAAAGGAGACCTAGGGGGTTGAAGGCAATGGGCAGCAAGGAAAGTGATTCAGAATGA
AA sequence
>Lus10010312 pacid=23169514 polypeptide=Lus10010312 locus=Lus10010312.g ID=Lus10010312.BGIv1.0 annot-version=v1.0
MDKEKKNSENDFVLQWGTKKRLRCVKLKKNQNLANSSRPTDSFPKKRLTSAVVSSKKDSPSRFIKNSDMLASRRKVSSPEKEDRYYTTRGSMGLDESSKL
IFDSVKLEDKGHVWPKLFISLSNKEKEEDFMAMKGCKPSQRPKKRAKLIQRTILLVSPGTWLSDLTQERYEVREKKAAKRRPRGLKAMGSKESDSE
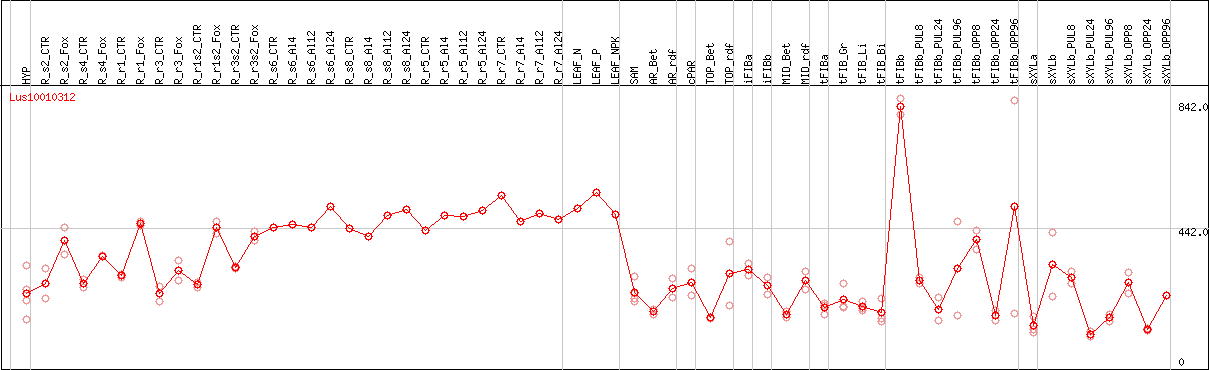
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010312 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.