Lus10010493 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10010493 pacid=23154072 polypeptide=Lus10010493 locus=Lus10010493.g ID=Lus10010493.BGIv1.0 annot-version=v1.0
ATGTCTCAATCACGATCATCGGCCAGCGATAGTCCCTCTCATCGCACTGCTGCCAGCCCCGTCGTCAGTGGCGCCGCCGCCGCCTCCAGCGAGTACCGCG
CCTCGGCGAAGCGCAAGTTGGACGACTACGTGTCCTCCATCGAGGACGACGATGACCTCGACATTTGGGACGACCTGGTGAGCGTGAGGATGCGGAAGGA
CGATTCCCTCCTCGCCGTTAACTCTTCATCCGACTCTCAGAATAAGAATCGGCCGCCGTTTACCGACGATGATTTGGAAGGGGTCGTGTCCCTCTCGACT
ATGGCTTCGTGTTCATATGCGACTTCGGGTTTTCAAGCGTGTGGAGAATCCTCGGTTTCGAAGATCCAGTTCTTCGTCCGGATGATCGCTGAAGGTAAAC
ATGTTGTTGTCTACGCTAGCCCGGAGGATACCGTGGAGTCTATTATGGAGAGAATTCAGGTGATCACTGGGATTTCTTCTTACCAGATGAGGTTAATTTA
CAACGGGAAGCAGCTGCAGCTAGATCAATCAGTGGCAAAGTGTGGAATTCAGAAGGATGCCACACTTCACTTAGTTGCACGGATGAGGAGCACTAAGCAT
CCGCAGACTTGTCGGGTAGTCGAGGACATGGTAATGTTTATCTGGAGGCTGTGTAAAAGTGGAATGCCAGCTTATTCTTATGCGCCTAAGCATATCAAGG
ATTTGATTAGCGAGTTCTTCAAGCTGGCTGCAAATAGTGAAACGGAAGATGCAACTGGGCATTTTGAGATTTTCATGTCCTGCTCTGCTCCAGAGACATT
GACGATGCTGTATGTCTCGACCAACAAAGGTAACAGTGATTGTGCTGATAGTTCGATTAGGCACTTCTTGAATTCATGTAGGATCTCTTTACCGAAGCCA
TTGCATGCGAAGTGTACACCTATAGTATTAGAATTTTGCAAGTTGCTTAGGAAGGTTGCTAATGACGATCCTCTATATCTTTGCTGTCGGAATACTCTTG
GTTCATTGTTGGATAATGATTCTGGGTGGCGTTGGTCATCTAAGAATGGGGTGGGAGCGGATGATGCAAAGGGGTCGATTACGATAGCAGAGATTTTTCC
GTTTGTTCAAGAGTTAGCTACTAGGTTGTCCAATGATCTGGGTTCAAGTCTGAAAACTCCAGTGAGTTTAGGACCATTAGGATGTGATGTCCGTGACTTC
TCTGCATTCTTGACACCTTTACGAATGGCTATATCGGAGCAAGTGGGACGGCCTGTTACAATGCCTTTGAACAAGAAAGGTGTTGACCATCTGTTATATG
CCGGAGAGATGGAGCAGCTTAGTGTTATATTTAGGGACTTATTGACAAAGATGGATCAATGCCTTATGAAGGTGGAGGGCTCCTCGCTTTTGAAGTCAAG
TGGAGAGTGTGAGTTGAGCAGTACCTCATGGTCTCAGTATCTTGCCATTTTGAAGGAATTGAATGAAATAGCTAAGCTCTACAAGGATTCCGAAGAACAA
TTCTGGACAATTTTGAGGCTTAGAAAGGCTTCTATTTGTGTTTTGATTGTCAAATATGCAAAGAGAGCTGAGGATCATCAGTGGCTCCTCAAACACAAGG
ATGTGACGGATTTTGAGTCAAGAAGACATTTGGCTATGATGATGTTCCCTGAGGTGAAAGAAGACTATGAAGAGCTGCATGAAATGCTTATTGACAGGGC
TAACCTGCTGGAGGAGTCATTTGCATACATTGGTCGTGCCGAACCAGATTCTTTGCATGGAGGTTTGTTTATGGAGTTCAAGAATGAGGAAGCTACTGGT
CCTGGTGTATTACGTGAGTGGTTTCTCTTGGTTACACAGGCTATTTTCAACCCTCATAATGCTCTTTTTCTAGCATGCCCAAGTGATCGCAGAAGGTTCC
ACCCTAACCCTGCTTCGAAGGTTGAACCTAGGCATCTCGAGTATTTCGCCTTTTCCGGTCGAGTGATTGCATTAGCTTTGATGCACAAAGTGCAAGTGGG
CATAGTCCTAGATCGGGTCCTTTTCTTACAATTAGCTGGAGGGCCTATATATTTGGAAGACATAAAGGATGCTGATCCGATCCTGTACAGTAGCTGCAAG
CAAATCCTGGATATGGATGCCGAGTTTATTGATTCCGATGCTTTAGGGCTGACATTTGTTAGAGAAGTCGAGGAGTTGGGATCCAGGAACGTTGTAGAAC
TCCGTCCTGGAGGGAAAAACATCGTAGTGAATAGCAAGAACAGAGGGGAGTACGTAACCCTTCTGGTTCGGCATCGATTTGTCACATCCATCTCTGAGCA
GGTCTCACATTTTGCAAAAGGATTCACTGATATTCTTTCCAATGGAAGTTTCCGACACTCTTTTTTCCAAAGCTTGGAGCTAGAAGACCTTGATTGGATG
CTTTACGGAAGTGAAAGCGCAATCTGTGTTGAAGACTGGAAGGCACATACTGAGTACAATGGCTTCAAAGAAACAGATCCCCAAATATCCTGGTTCTGGA
AGATTGTTGAGGAAATGCCAGCCGAGCGGAGGAAAGTATTGCTCTTCTTCTGGACATCAATTAAATACCTACCGGTTGAAGGTTTCCGAGGACTTGCTTC
TCGTCTATACATATACAAGTCCACGGAGCCACAAGATCGTCTCCCTTCGTCCCATACATGCTTCTACAGGCTGTGCTTCCCACCCTATCCATCCATGGAC
ATCATGCAAGATCGACTCCGCGTCATAACCCAAGAGCATATTGGTTGCAGCTTCGGCACCTGGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10010493 pacid=23154072 polypeptide=Lus10010493 locus=Lus10010493.g ID=Lus10010493.BGIv1.0 annot-version=v1.0
MSQSRSSASDSPSHRTAASPVVSGAAAASSEYRASAKRKLDDYVSSIEDDDDLDIWDDLVSVRMRKDDSLLAVNSSSDSQNKNRPPFTDDDLEGVVSLST
MASCSYATSGFQACGESSVSKIQFFVRMIAEGKHVVVYASPEDTVESIMERIQVITGISSYQMRLIYNGKQLQLDQSVAKCGIQKDATLHLVARMRSTKH
PQTCRVVEDMVMFIWRLCKSGMPAYSYAPKHIKDLISEFFKLAANSETEDATGHFEIFMSCSAPETLTMLYVSTNKGNSDCADSSIRHFLNSCRISLPKP
LHAKCTPIVLEFCKLLRKVANDDPLYLCCRNTLGSLLDNDSGWRWSSKNGVGADDAKGSITIAEIFPFVQELATRLSNDLGSSLKTPVSLGPLGCDVRDF
SAFLTPLRMAISEQVGRPVTMPLNKKGVDHLLYAGEMEQLSVIFRDLLTKMDQCLMKVEGSSLLKSSGECELSSTSWSQYLAILKELNEIAKLYKDSEEQ
FWTILRLRKASICVLIVKYAKRAEDHQWLLKHKDVTDFESRRHLAMMMFPEVKEDYEELHEMLIDRANLLEESFAYIGRAEPDSLHGGLFMEFKNEEATG
PGVLREWFLLVTQAIFNPHNALFLACPSDRRRFHPNPASKVEPRHLEYFAFSGRVIALALMHKVQVGIVLDRVLFLQLAGGPIYLEDIKDADPILYSSCK
QILDMDAEFIDSDALGLTFVREVEELGSRNVVELRPGGKNIVVNSKNRGEYVTLLVRHRFVTSISEQVSHFAKGFTDILSNGSFRHSFFQSLELEDLDWM
LYGSESAICVEDWKAHTEYNGFKETDPQISWFWKIVEEMPAERRKVLLFFWTSIKYLPVEGFRGLASRLYIYKSTEPQDRLPSSHTCFYRLCFPPYPSMD
IMQDRLRVITQEHIGCSFGTW
|
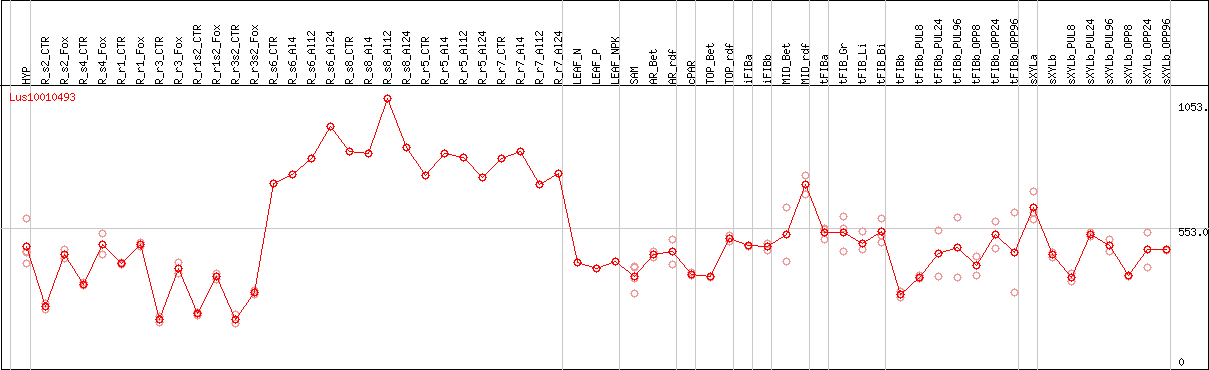
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010493 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
