External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G32150 338 / 7e-118
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT3G62040 258 / 2e-86
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT5G02230 240 / 4e-79
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
AT5G59480 224 / 1e-72
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
AT5G59490 223 / 2e-72
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004914
511 / 0
AT2G32150 338 / 6e-118
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10009256
237 / 2e-78
AT3G62040 364 / 1e-128
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10038016
239 / 5e-78
AT3G62040 363 / 6e-127
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10010848
236 / 6e-77
AT5G02230 394 / 7e-139
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Lus10024382
235 / 1e-76
AT5G02230 394 / 5e-139
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Lus10004971
229 / 8e-75
AT5G59480 392 / 1e-138
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Lus10016518
226 / 2e-73
AT5G59480 383 / 5e-135
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Lus10028107
226 / 3e-73
AT5G02230 369 / 4e-129
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Lus10040786
224 / 2e-72
AT5G59480 382 / 4e-134
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G151700
431 / 8e-155
AT2G32150 319 / 2e-110
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.010G089300
423 / 2e-151
AT2G32150 317 / 9e-110
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.014G110800
251 / 3e-83
AT3G62040 356 / 8e-125
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.002G185300
247 / 2e-81
AT3G62040 360 / 5e-126
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.006G086900
246 / 3e-81
AT5G02230 402 / 4e-142
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Potri.001G242300
238 / 4e-78
AT5G59480 409 / 2e-145
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Potri.009G033500
232 / 7e-75
AT5G59480 361 / 4e-125
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0137
HAD
PF00702
Hydrolase
haloacid dehalogenase-like hydrolase
Representative CDS sequence
>Lus10010537 pacid=23156642 polypeptide=Lus10010537 locus=Lus10010537.g ID=Lus10010537.BGIv1.0 annot-version=v1.0
ATGGATTTCCGCAGCAACACTACTACTTGCGACTCCTTTTCCCCTTTCCACTGCCTTCTCTTTGATTTAGACGATACGCTTTACTCTTCCAAGCTCGGAA
TAGCTGAAGCTCTCCGCAACAACATCGACGATTTTCTTGTTGAGAAATGTGGATTCCCTCCGAGCAAAGCTCCAAGTCTCCGATCAGAGCTCTTCAAAAC
CTACGGCAGCTCTCTCGCCGGCTTGCGCGCATTAGGTTACAACATCGATGCCGATGATTACCATAGTTTCGTGCACGGAAGATTGCCGTACCACCTGATC
AAACCAGACCCGCAGCTATCCAACCTCCTCACAACCATTCCGCAAAGAAAACTCATATTCACTAACTCGGACAGAAACCACGCGATCGCAGTGCTGGAGA
GGTTAGGATTGGAGGACTGTTTCGATCAGATCATCTGCTTTGAGACCATGAACCCTAACCTCCCCAATTCCACCCGACCCGACGAATTCCCCGTTCTCCT
CAAGCCCTCCATGGACGCCATGAGAATCGCACTCCGTGCCGCCGATGTCGATCCCCGTCGCACGCTGTTTCTGGACGATAACTCACGCAACGTGGCAGCA
GGGAAAGCGCTGGGCCTCCGCACCGTTCTGGTTGGGAAGACGGTAAAGATCCCAGAAGCTGACTACGTCCTGGAGAAGATACACAATCTTGCCCAGGCTG
TTCCGGAGATTTGGGTCACCGGCGATCAAAGGATTAGCCGCAGCAAGAGCGATATGGATTCCATTCTCGCCACCACTGCAGCCACCGTTGGGGCCTAG
AA sequence
>Lus10010537 pacid=23156642 polypeptide=Lus10010537 locus=Lus10010537.g ID=Lus10010537.BGIv1.0 annot-version=v1.0
MDFRSNTTTCDSFSPFHCLLFDLDDTLYSSKLGIAEALRNNIDDFLVEKCGFPPSKAPSLRSELFKTYGSSLAGLRALGYNIDADDYHSFVHGRLPYHLI
KPDPQLSNLLTTIPQRKLIFTNSDRNHAIAVLERLGLEDCFDQIICFETMNPNLPNSTRPDEFPVLLKPSMDAMRIALRAADVDPRRTLFLDDNSRNVAA
GKALGLRTVLVGKTVKIPEADYVLEKIHNLAQAVPEIWVTGDQRISRSKSDMDSILATTAATVGA
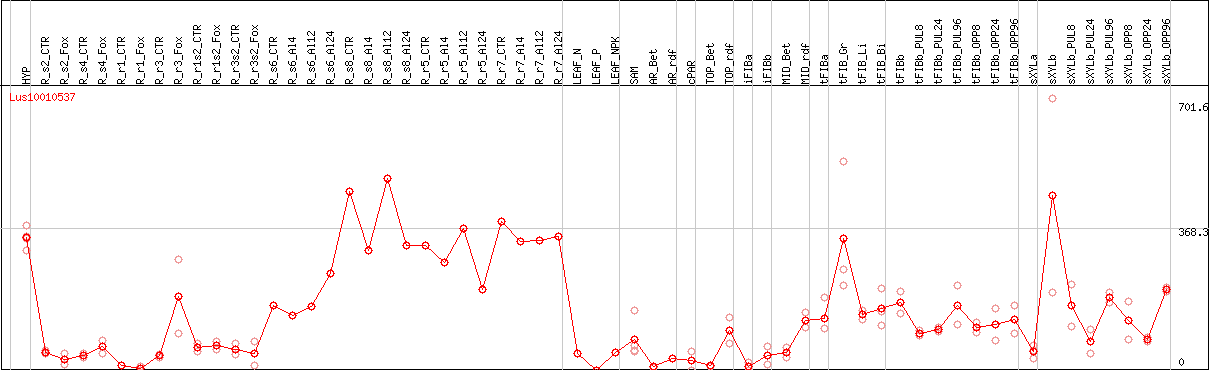
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010537 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.