External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G54340 261 / 1e-88
MADS
AP3, ATAP3
APETALA 3, K-box region and MADS-box transcription factor family protein (.1)
AT5G20240 124 / 2e-35
MADS
PI, PISTILLATA
PISTILLATA, K-box region and MADS-box transcription factor family protein (.1)
AT5G60910 120 / 3e-33
MADS
FUL, AGL8
FRUITFULL, AGAMOUS-like 8 (.1.2)
AT4G18960 117 / 5e-32
MADS
AG
AGAMOUS, K-box region and MADS-box transcription factor family protein (.1.2)
AT2G45660 115 / 7e-32
MADS
ATSOC1, SOC1, AGL20
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
AT2G45650 116 / 9e-32
MADS
AGL6
AGAMOUS-like 6 (.1)
AT2G42830 114 / 6e-31
MADS
AGL5, SHP2
SHATTERPROOF 2, AGAMOUS-like 5, K-box region and MADS-box transcription factor family protein (.1.2)
AT5G23260 113 / 1e-30
MADS
ABS, TT16, AGL32
TRANSPARENT TESTA16, AGAMOUS-like 32, ARABIDOPSIS BSISTER, K-box region and MADS-box transcription factor family protein (.1.2.3)
AT5G51870 112 / 1e-30
MADS
AGL71
AGAMOUS-like 71 (.1.2.3)
AT3G58780 112 / 2e-30
MADS
AGL1, SHP1
SHATTERPROOF 1, AGAMOUS-like 1, K-box region and MADS-box transcription factor family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031960
426 / 1e-153
AT3G54340 275 / 5e-94
APETALA 3, K-box region and MADS-box transcription factor family protein (.1)
Lus10033187
415 / 3e-149
AT3G54340 275 / 4e-94
APETALA 3, K-box region and MADS-box transcription factor family protein (.1)
Lus10029719
128 / 2e-36
AT5G20240 221 / 3e-73
PISTILLATA, K-box region and MADS-box transcription factor family protein (.1)
Lus10026678
123 / 1e-34
AT5G15800 225 / 3e-74
SEPALLATA1, AGAMOUS-like 2, K-box region and MADS-box transcription factor family protein (.1.2)
Lus10042749
121 / 8e-34
AT5G20240 219 / 2e-72
PISTILLATA, K-box region and MADS-box transcription factor family protein (.1)
Lus10010184
120 / 3e-33
AT5G23260 218 / 3e-71
TRANSPARENT TESTA16, AGAMOUS-like 32, ARABIDOPSIS BSISTER, K-box region and MADS-box transcription factor family protein (.1.2.3)
Lus10004638
119 / 4e-33
AT3G02310 246 / 2e-82
SEPALLATA 2, AGAMOUS-like 4, K-box region and MADS-box transcription factor family protein (.1)
Lus10017388
119 / 5e-33
AT5G23260 220 / 4e-72
TRANSPARENT TESTA16, AGAMOUS-like 32, ARABIDOPSIS BSISTER, K-box region and MADS-box transcription factor family protein (.1.2.3)
Lus10021140
117 / 1e-31
AT5G60910 291 / 1e-99
FRUITFULL, AGAMOUS-like 8 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G028400
249 / 1e-84
AT3G54340 229 / 7e-77
APETALA 3, K-box region and MADS-box transcription factor family protein (.1)
Potri.005G118000
244 / 4e-82
AT3G54340 226 / 9e-75
APETALA 3, K-box region and MADS-box transcription factor family protein (.1)
Potri.007G017000
231 / 5e-77
AT3G54340 208 / 1e-67
APETALA 3, K-box region and MADS-box transcription factor family protein (.1)
Potri.002G079000
133 / 1e-38
AT5G20240 238 / 2e-80
PISTILLATA, K-box region and MADS-box transcription factor family protein (.1)
Potri.005G182200
129 / 3e-37
AT5G20240 244 / 2e-82
PISTILLATA, K-box region and MADS-box transcription factor family protein (.1)
Potri.002G151700
121 / 5e-34
AT2G45660 212 / 8e-70
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
Potri.014G074200
121 / 9e-34
AT2G45660 242 / 3e-81
SUPPRESSOR OF OVEREXPRESSION OF CO 1, AGAMOUS-like 20 (.1)
Potri.014G074100
121 / 1e-33
AT2G45650 311 / 5e-108
AGAMOUS-like 6 (.1)
Potri.007G073000
118 / 1e-32
AT5G23260 256 / 3e-86
TRANSPARENT TESTA16, AGAMOUS-like 32, ARABIDOPSIS BSISTER, K-box region and MADS-box transcription factor family protein (.1.2.3)
Potri.010G154100
118 / 2e-32
AT1G69120 324 / 7e-113
APETALA1, AGAMOUS-like 7, K-box region and MADS-box transcription factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01486
K-box
K-box region
PF00319
SRF-TF
SRF-type transcription factor (DNA-binding and dimerisation domain)
Representative CDS sequence
>Lus10010630 pacid=23143661 polypeptide=Lus10010630 locus=Lus10010630.g ID=Lus10010630.BGIv1.0 annot-version=v1.0
ATGGCGAGGGGGAAGATCCAGATCAAGAGGATCGAGAACTCCACCAATCGCCAGGTCACCTACTCTAAGCGCCGCAACGGTCTCTTCAAGAAGGCCAACG
AGCTCACCGTCTTATGTGACGCTAAGGTCTCCATCATCATGTTCTCCAGCACCGACAAGCTTCACGAGTTCATCAGCCCTGCCGCCACAACTAAGGGGAT
GTTGGATCAGTACCAGAGAACACTTGGCGTCGACATCTGGACCACGCACTATGAGAAAATGCAAGATCACTTGAAGAAACTGAAAGACGTGAACAGAGAT
CTGCGCAAGGAGATACGGAGAAGGATGGGGGAGTGCTTGAACGATCTGGAGTTGGATCAGATGCGAGCTCTTGACATGGAAATGGATAATGCTGTTAAGA
TCATCCGTGAGCGCAAGAATCAGGTGATCTCCAATCAGATTGAGAGGCAAAAGAAAAAGGTGAGGAATGTGGAAGCAATACACAGAAACCTTCAGCATGC
ACTTCGAGAGAGAGAGGATGATCCACACTATGGTTTAGTGGATAATGGAGGGGATTATGAACCAGCTGTGGTTGGTTTCCCAACTGGGGGTTCTCGAGTT
TTCGCTTTAAGGCTAAATCACCACAATCTCCACACTGCTACTACTGCTACTGGATCCGATCTCACCACTTTCCACTTGCTTGAGTAA
AA sequence
>Lus10010630 pacid=23143661 polypeptide=Lus10010630 locus=Lus10010630.g ID=Lus10010630.BGIv1.0 annot-version=v1.0
MARGKIQIKRIENSTNRQVTYSKRRNGLFKKANELTVLCDAKVSIIMFSSTDKLHEFISPAATTKGMLDQYQRTLGVDIWTTHYEKMQDHLKKLKDVNRD
LRKEIRRRMGECLNDLELDQMRALDMEMDNAVKIIRERKNQVISNQIERQKKKVRNVEAIHRNLQHALREREDDPHYGLVDNGGDYEPAVVGFPTGGSRV
FALRLNHHNLHTATTATGSDLTTFHLLE
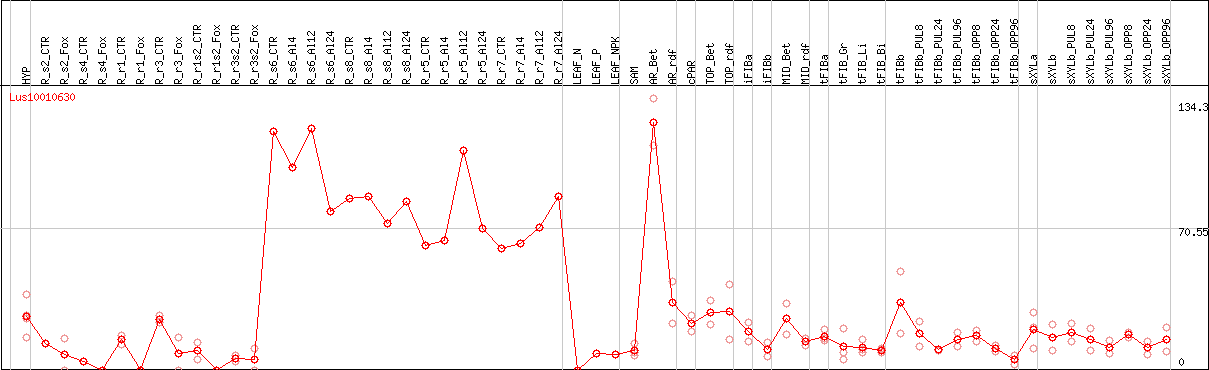
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010630 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.