Lus10010642 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10010642 pacid=23143639 polypeptide=Lus10010642 locus=Lus10010642.g ID=Lus10010642.BGIv1.0 annot-version=v1.0
ATGGAGTCTCAGCCGGCTTCTTCTGCCGCTGATGCCCATGCCAACGGCCGGAAGCACGAGTACGGCATGTTTAGCGGTCCAATGAAGCTCGGGTCTAGAG
TGGAGGATTACAGGGAGATATTTGAATGCTCGGGAACGTCGTCCATTCCGGTGTTGGATGTTCCCGAGTTGAATGAGAGGAAGGCTTCCGTCGAGCCTCG
GAACTCTCGGCTGGACTACTCCAAGATTTTCGGCGGACTCGGAGAAGCGGACATTGCTATGCCTTACGACAACTTGAATTCGAGAGCGGAGACGGATATT
CGGGTTCCAGCAGAATCAAGATCACAGTTTTTAGGGCATGAGCAATCTACGTTTGCAAGGAAGAGAATGCCAGCGGAAGCATCTCTTCAGCCATTGGATG
GTGGGAAAAATGCCAACGTGTTGCATCTTAAGACAAACTCGGGAGGCAGAATGGGGAGAAGTGGCATGACCCACGTTGCTCAGTTCCATGCTGTTCCAGG
CTTCACTCATCTAGTTGACGAAAATGTGCCCTCGGGATCTACTGGAGGGACCAAGTCTGGACGCTTTGTGCGTACCAATGTTCATCTTAATCTCAGTGAG
GAGATGAAGGAAACCAGATCCAAGAGGAGAGCTTACTCCGGACCACTCCTCTTCAGCAATGCTTCGGGACAGTACACAGAACATCACAGGACGTCTAGTT
CCATATTGAATGATGTATCGTCTGACCTTCACAGAGGCTCACATCCATCTGCAGTTTGTTCATCGTCTCCAAGCTCACCACTTAGTTTTGGTAATGAGAA
GGCTGCTTCCAACAGGTTGCTGAACTCCAAATTTGGGTTTTCTAAAAGTGCTACTGGCTATTATTCACCACCTTTCTTGGATGATGAAGTAGATGCAAAT
TCAACTGCTGTCGCTTCTGCAGCTGCGGTGAGAAAGGCAATAGATGATGCTGAAGCAAAAATTAAATATGCAAAGGAACTCATGGAAAGGAGAAAAGAAG
GCCGACATAATCGTTCGAAACCAAAGGTCAATCATGGCTTGAAGTCTGAGACGAAACTGGAAGCTATATCAGCTGAAATGTACAGGAGAGATGATGCTCC
AAAACGGCCAGTCGTAAACTGTTCACCAGCTCAAATTTCAGGAGAAGTCCGCCAAGTGAATTTGGATTCCAGGGACCAGAAGAAAGATTCTCTTGCTAGA
CCTTCTCTTACAGTAAACCACCAGCTAGAAACCACCTTTACTAAAGTTGATTCCAGACTGGAAGAAATAGAGATCTGCAAAACTCCGCAAGAGTTTTCTG
AAGCTTCCATTAGCAAAGGGCTCAGGCCATCAATATCAGAGTTAAACGAACAAGACCTGGGGCAAAAAGTAATACCAAGTAGTTATGAGAACAAATGGAA
GGAGAAGATGAAAACATCATTTCTGCAGGACAAATTTGAAAACATAATGAAAATGCTGCGAGCACCAGAAGACGTTCCAGATGTTGAATACCAACTGATG
GTCGAAAGAAAATCAGGTGGGATAAAAGGAAATGTAGCTCATGTGAGAAGTGAAGAGAAACAAGAGGCTGCATGCACCAAAGAGGAAATAGAAGGTAGAT
CAAACGAAGGCACCGCAAGGAAGGAGGATGAGAGCACGGTTAAAGTTAATTCCTGGCTGAATAATATTGCCAAGGGGCATAAGGAACAAAAAGATTATCA
GAATGGCGACGAAATGGAGAAGAAACAAAGTTTTGATAGGAGGGAAGATGACTCAGAGAACACTTCTGCTAAAATTGATGGCGGGGAACCATCAAAAGTA
AGGTTTGACCATTTTGGGATAGAGAAGGAAGGTCGAGACAAGTCTACAGCAAGTGATCAGGAAGTAGAGATAGAGAAGGTTGAAGAAGTTGAGAAAGAAA
GACGGATTGGTGATGATGATCCTATTAATTCTACTAGCAGAGGGCGAGAAAACTTTTCCAATTCCATGGACAGTAAGAGGTTAGACTCGGTGGTTCATCA
AAGGTTGGAAGGTGATGCAAAGGATAACGTTGCTCAAGAGCCTCTTGGATTGCTGAATGGCAACGTTGAAGTGCTTCATGCACTGCACGAGCGAGAGGAA
AGTGAGAATGCCGAACAGATTAATGAATCCCATAGATACAGAGAGCTTGTAGTTACAGCGAATGACAGGGCCACCAACCCTTTCTCTGGGCCCGGAGACA
TTGAAAAACAATCAAATGTTTTTGAAGAGACATATATCTTTGAGGGAGGGGAGGATTGTACAAGGGTTGACCCTTTTAATTCTACTGGATTCGATGGGAT
CGGAAAACAGAGCGATGAGCCCCTTATTGTTGATGAAAATGGGGTCTGTTCAAGTGGACTTGTTTTCAACTTTAAGGACCAGCATACTGAAGACAGTGCA
GCAAAAAGCAAAGGAACCTATTGTCCTGAAAATCACGTAGAAGAAGCGACTAGACTGTCAGGAGAAAGCAAAGATAACAATAAGGATGCTGAAGTTGGCA
GCACAGATGAAGATACTGCTAGTAACCTTGCATCTCTTAATGAGGTAAAGGTGTTACCAAATGGGGAAGAATCCAGGCCGCTCGCAGAACTGGAAAAGCA
TGTAGAAGAAGCCATTGGTGATTTGGAAGATCATGATAAGGATAACGTTAACAATGGTAAATCGGTTAGTGAAAGGAAATTATCGGATGATGGAATAAGT
AAAAGAACAAGTCAAGCATTTCGGGAGTCTGAGAAGGAAGAACCTGTCGAGGCAGCAGTATCTTCACATGAGAAAACAACTAAAAGTATCATTGATGACT
CTGAGGATGCTTATTATGAGGTGCTCCAAATGCAAGAGACAGAAGAAAACCAAAGCTCGCGAATGGACGTGGAGGTGGAAAGGCAGCAACCCAGGGAAAA
GGAAGTTGCAAAAGAGAGAGACTTGGAGAAAGAGAAAGAGCGAAGAGTTGTCGAAAGGGCAATAAGGGAAGCACGCGAGAGGGCATTTACTGAAGCTCGA
GAGAAGGCAGAAAAGTTAGCTGCAGCAGAAGCTCAGCGAAGAGCAAAAGCTGTGGCTGCTGCTCGAGAGAGGTCTGAGAGGGCTATCGCATCTGAGAAAG
CTTCCACAGAAACCAAACTGAAAGCTGAGCGTGCGGCAGTTGAAAGAGCAACTGCAGAAGCTAGAGAGCGGGCTCTGGAGAAAGCATTATCTGAAAAGGC
TTCCCTCAAAGGGAAAAATTGTGCAGAGAAGTCTCCTGCCGTTTCAACTGATGATGGAAGACCCATTAATCAACGACACAAAAACTCAGGTTACAACACC
AGCTCAAGACATCCAAGTTCTACAACTAGCGGTGCTGAAATCTCTGGTAATGGAGCAAACGGTGAATCTCTGAAATCTAAAGCCTCGTTGGAGAAGCAGC
AGCAAATGGCTGAGCGTGCGGCGAAAGCTCTGGCAGAGAAGAATATGCGCGATCTTCTTGTTCAGAAGGAGCAAGCTGAAAAAAATAGACTAGCAGAAGT
TGTGGATGCGGAGATAAAAAGGTGGTCCAGCGGGAAAGAGCGGAACTTGCGAGTTCTGCTTTCTACACTTCACTATATCCTTGGACCTGACAGTGGTTGG
CAAGCAATTCCTAGGACGGATCTTAACTCAGCTACTGCTGTTAGGAAAGCTTACAAGAAAGCAACTCTGTTCGTCCATCCTGATAAGCTGCAGCAGCGCG
GTGCCAGCATTCGCCACAAGTACGTCTGCGAGAAGGTCTTCGAACTTCTGAAGATTCTCGCGGAAGCCTTTGGTGGGGATGCAACCAGCAGCGGAGGACC
AACAGAAAATAGTGAAGAAGTCATGGAAATGTTCACCAATTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10010642 pacid=23143639 polypeptide=Lus10010642 locus=Lus10010642.g ID=Lus10010642.BGIv1.0 annot-version=v1.0
MESQPASSAADAHANGRKHEYGMFSGPMKLGSRVEDYREIFECSGTSSIPVLDVPELNERKASVEPRNSRLDYSKIFGGLGEADIAMPYDNLNSRAETDI
RVPAESRSQFLGHEQSTFARKRMPAEASLQPLDGGKNANVLHLKTNSGGRMGRSGMTHVAQFHAVPGFTHLVDENVPSGSTGGTKSGRFVRTNVHLNLSE
EMKETRSKRRAYSGPLLFSNASGQYTEHHRTSSSILNDVSSDLHRGSHPSAVCSSSPSSPLSFGNEKAASNRLLNSKFGFSKSATGYYSPPFLDDEVDAN
STAVASAAAVRKAIDDAEAKIKYAKELMERRKEGRHNRSKPKVNHGLKSETKLEAISAEMYRRDDAPKRPVVNCSPAQISGEVRQVNLDSRDQKKDSLAR
PSLTVNHQLETTFTKVDSRLEEIEICKTPQEFSEASISKGLRPSISELNEQDLGQKVIPSSYENKWKEKMKTSFLQDKFENIMKMLRAPEDVPDVEYQLM
VERKSGGIKGNVAHVRSEEKQEAACTKEEIEGRSNEGTARKEDESTVKVNSWLNNIAKGHKEQKDYQNGDEMEKKQSFDRREDDSENTSAKIDGGEPSKV
RFDHFGIEKEGRDKSTASDQEVEIEKVEEVEKERRIGDDDPINSTSRGRENFSNSMDSKRLDSVVHQRLEGDAKDNVAQEPLGLLNGNVEVLHALHEREE
SENAEQINESHRYRELVVTANDRATNPFSGPGDIEKQSNVFEETYIFEGGEDCTRVDPFNSTGFDGIGKQSDEPLIVDENGVCSSGLVFNFKDQHTEDSA
AKSKGTYCPENHVEEATRLSGESKDNNKDAEVGSTDEDTASNLASLNEVKVLPNGEESRPLAELEKHVEEAIGDLEDHDKDNVNNGKSVSERKLSDDGIS
KRTSQAFRESEKEEPVEAAVSSHEKTTKSIIDDSEDAYYEVLQMQETEENQSSRMDVEVERQQPREKEVAKERDLEKEKERRVVERAIREARERAFTEAR
EKAEKLAAAEAQRRAKAVAAARERSERAIASEKASTETKLKAERAAVERATAEARERALEKALSEKASLKGKNCAEKSPAVSTDDGRPINQRHKNSGYNT
SSRHPSSTTSGAEISGNGANGESLKSKASLEKQQQMAERAAKALAEKNMRDLLVQKEQAEKNRLAEVVDAEIKRWSSGKERNLRVLLSTLHYILGPDSGW
QAIPRTDLNSATAVRKAYKKATLFVHPDKLQQRGASIRHKYVCEKVFELLKILAEAFGGDATSSGGPTENSEEVMEMFTN
|
DESeq2's median of ratios [FLAX]
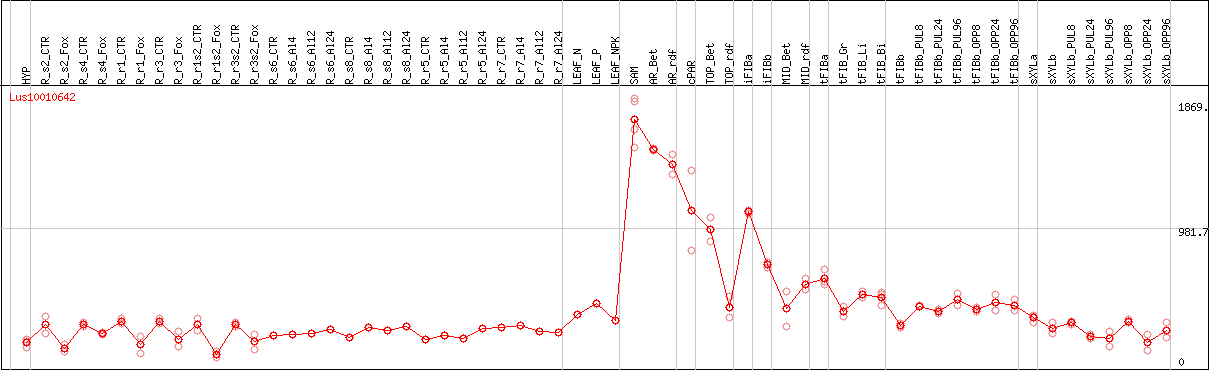
Coexpressed genes
Lus10010642 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
