External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G69600 122 / 7e-33
ZF_HD
ATHB29, ZFHD1, ZHD11
ZINC FINGER HOMEODOMAIN 11, ARABIDOPSIS THALIANA HOMEOBOX PROTEIN 29, zinc finger homeodomain 1 (.1)
AT5G60480 111 / 2e-29
ZF_HD
ATHB26, ZHD12
ZINC FINGER HOMEODOMAIN 12, homeobox protein 26 (.1)
AT2G02540 113 / 4e-29
ZF_HD
ATHB21, ZFHD4, ZHD3
ZINC FINGER HOMEODOMAIN 3, ZINC FINGER HOMEODOMAIN 4, homeobox protein 21 (.1)
AT1G14440 111 / 3e-28
ZF_HD
ATHB31, ZHD4
ZINC FINGER HOMEODOMAIN 4, homeobox protein 31 (.1.2)
AT1G75240 103 / 2e-25
ZF_HD
ATHB33, ZHD5
zinc-finger homeodomain 5, homeobox protein 33 (.1)
AT4G24660 97 / 1e-23
ZF_HD
ATHB22, MEE68, ZHD2
ZINC FINGER HOMEODOMAIN 2, MATERNAL EFFECT EMBRYO ARREST 68, homeobox protein 22 (.1)
AT5G65410 92 / 2e-21
ZF_HD
ATHB25, ZFHD2, ZHD1
ZINC FINGER HOMEODOMAIN 1, ZINC FINGER HOMEODOMAIN 2, ARABIDOPSIS THALIANA HOMEOBOX PROTEIN 25, homeobox protein 25 (.1)
AT2G18350 92 / 2e-21
ZF_HD
ATHB24, ZHD6
ZINC FINGER HOMEODOMAIN 6, homeobox protein 24 (.1)
AT1G74660 85 / 2e-20
ZF_HD
MIF1
mini zinc finger 1 (.1)
AT3G50890 87 / 1e-19
ZF_HD
ATHB28, ZHD7
ZINC FINGER HOMEODOMAIN 7, homeobox protein 28 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038135
274 / 1e-90
AT2G02540 223 / 7e-71
ZINC FINGER HOMEODOMAIN 3, ZINC FINGER HOMEODOMAIN 4, homeobox protein 21 (.1)
Lus10010693
198 / 9e-64
AT2G02540 104 / 7e-28
ZINC FINGER HOMEODOMAIN 3, ZINC FINGER HOMEODOMAIN 4, homeobox protein 21 (.1)
Lus10026010
123 / 4e-33
AT1G75240 172 / 2e-52
zinc-finger homeodomain 5, homeobox protein 33 (.1)
Lus10007147
102 / 1e-24
AT2G18350 222 / 1e-70
ZINC FINGER HOMEODOMAIN 6, homeobox protein 24 (.1)
Lus10036753
79 / 4e-17
AT5G15210 112 / 3e-30
ZINC FINGER HOMEODOMAIN 8, ZINC FINGER HOMEODOMAIN 3, homeobox protein 30 (.1)
Lus10033016
75 / 9e-17
AT3G28917 103 / 4e-30
mini zinc finger 2 (.1)
Lus10014302
78 / 3e-16
AT1G75240 176 / 9e-54
zinc-finger homeodomain 5, homeobox protein 33 (.1)
Lus10015900
76 / 3e-16
AT4G24660 145 / 6e-44
ZINC FINGER HOMEODOMAIN 2, MATERNAL EFFECT EMBRYO ARREST 68, homeobox protein 22 (.1)
Lus10030679
76 / 1e-15
AT5G15210 145 / 2e-42
ZINC FINGER HOMEODOMAIN 8, ZINC FINGER HOMEODOMAIN 3, homeobox protein 30 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G000400
180 / 8e-55
AT1G14440 238 / 9e-78
ZINC FINGER HOMEODOMAIN 4, homeobox protein 31 (.1.2)
Potri.005G122500
136 / 2e-37
AT2G18350 179 / 2e-54
ZINC FINGER HOMEODOMAIN 6, homeobox protein 24 (.1)
Potri.004G229600
115 / 8e-30
AT1G14440 244 / 1e-79
ZINC FINGER HOMEODOMAIN 4, homeobox protein 31 (.1.2)
Potri.004G135100
105 / 1e-26
AT2G02540 213 / 4e-68
ZINC FINGER HOMEODOMAIN 3, ZINC FINGER HOMEODOMAIN 4, homeobox protein 21 (.1)
Potri.007G024100
100 / 3e-24
AT1G75240 186 / 2e-56
zinc-finger homeodomain 5, homeobox protein 33 (.1)
Potri.005G158800
97 / 1e-23
AT4G24660 204 / 1e-65
ZINC FINGER HOMEODOMAIN 2, MATERNAL EFFECT EMBRYO ARREST 68, homeobox protein 22 (.1)
Potri.002G035200
97 / 5e-23
AT1G14440 169 / 2e-50
ZINC FINGER HOMEODOMAIN 4, homeobox protein 31 (.1.2)
Potri.019G081300
96 / 7e-23
AT4G24660 166 / 8e-51
ZINC FINGER HOMEODOMAIN 2, MATERNAL EFFECT EMBRYO ARREST 68, homeobox protein 22 (.1)
Potri.002G102900
95 / 1e-22
AT4G24660 209 / 9e-68
ZINC FINGER HOMEODOMAIN 2, MATERNAL EFFECT EMBRYO ARREST 68, homeobox protein 22 (.1)
Potri.004G126600
91 / 1e-22
AT3G28917 129 / 5e-40
mini zinc finger 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04770
ZF-HD_dimer
ZF-HD protein dimerisation region
Representative CDS sequence
>Lus10010694 pacid=23141674 polypeptide=Lus10010694 locus=Lus10010694.g ID=Lus10010694.BGIv1.0 annot-version=v1.0
ATGGAATTAACCAGTCATCATCATCATCATGTTCATGAGGAAGAGGAGGATCACATGGCAATTCCAGTAAACTCTTCCTATGCAGGAGGAGGCCTAAACC
ACTTGATCCACAATCCTCATCCTCCTCTGTCTCCTTCTACTGTTACACCCGCAACTTCTTCTGCACCATTATCTCTCCCCAACAATAATAACAATAATGG
ATCTGGAAGTCAACACCATCACCATATCAACAACAACAGCAAGAGAAGGTCTTCTTCAGTGATCAGATACAGGGAATGCCTGAAGAATCACGCGGCATCA
ATGGGAGGAAACGCTACAGACGGATGTGGGGAGTTCATGCCCAGCGGAGAAGAAGGCACCATCGAAGCACTCACTTGCTCCGCCTGCAACTGCCACCGCA
ACTTCCACCGCAAACAAACCATCCTCCACCACCACCCGCATCCTCATAATCACCACCACAATCCCCTCCTGCTACAACAAGTACCACTTCATCAACATCA
TCAAGCTCCACCGCCATTGGTGGTCCCCCACCACCACCACCACCACATGCTCATGTCATACAACATGGCTGCTGCTGCTGCTGCTGGATCACTGCTGCCC
TCTGAATCCGACAACCACAACAACGATGCTGAAGATCATCACCAGCATGAGTACCATAACAACAACAAGCAGCAGCAAGGGATGAATACGAAGAGGCACA
GGACAAAGTTCAGCCAGGAGCAGAAGGAGAAGATGCTTGCTTTTGCTGAGAGTGTTGGTTGGAAGCTACAGAAACAAGAGGAGAGCGTGGTCCAACAATT
CTGCCACGAGATTGGTGTTAACCGTCGTGTGCTTAAGGTCTGGATGCACAATAACAAGCACACTTACCAACATCATCATCATCAAAACGCTGTCGTATCT
CGTCCTCCAGTGCTTCATCCATCCACCACCACCACTGCTCCACTCCCGGCTTCTTCTTAA
AA sequence
>Lus10010694 pacid=23141674 polypeptide=Lus10010694 locus=Lus10010694.g ID=Lus10010694.BGIv1.0 annot-version=v1.0
MELTSHHHHHVHEEEEDHMAIPVNSSYAGGGLNHLIHNPHPPLSPSTVTPATSSAPLSLPNNNNNNGSGSQHHHHINNNSKRRSSSVIRYRECLKNHAAS
MGGNATDGCGEFMPSGEEGTIEALTCSACNCHRNFHRKQTILHHHPHPHNHHHNPLLLQQVPLHQHHQAPPPLVVPHHHHHHMLMSYNMAAAAAAGSLLP
SESDNHNNDAEDHHQHEYHNNNKQQQGMNTKRHRTKFSQEQKEKMLAFAESVGWKLQKQEESVVQQFCHEIGVNRRVLKVWMHNNKHTYQHHHHQNAVVS
RPPVLHPSTTTTAPLPASS
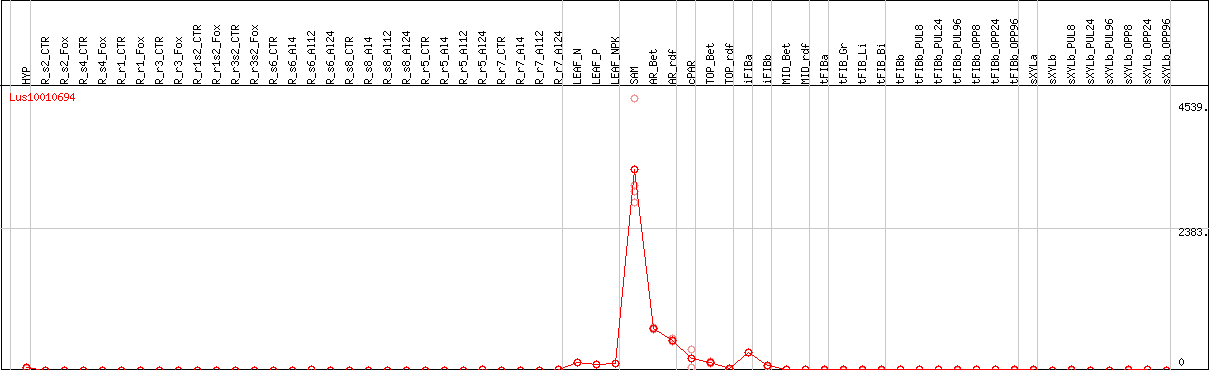
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010694 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.