External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G50760 108 / 3e-29
GATL2
galacturonosyltransferase-like 2 (.1)
AT1G19300 102 / 1e-26
ATGATL1, GATL1, PARVUS, GLZ1
PARVUS, GAOLAOZHUANGREN 1, GALACTURONOSYLTRANSFERASE-LIKE 1, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
AT1G02720 75 / 1e-16
GATL5
galacturonosyltransferase 5 (.1.2)
AT3G28340 74 / 2e-16
GolS8, GATL10
galactinol synthase 8, galacturonosyltransferase-like 10 (.1)
AT3G06260 74 / 2e-16
GolS9, GATL4
galactinol synthase 9, galacturonosyltransferase-like 4 (.1)
AT1G13250 74 / 3e-16
GATL3
galacturonosyltransferase-like 3 (.1)
AT1G24170 71 / 4e-15
GATL8, LGT9
GALACTURONOSYLTRANSFERASE-LIKE 8, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
AT1G70090 71 / 5e-15
GATL9, LGT8
GALACTURONOSYLTRANSFERASE-LIKE 9, glucosyl transferase family 8 (.1.2)
AT4G02130 68 / 4e-14
GATL6, LGT10
galacturonosyltransferase 6 (.1.2.3)
AT3G62660 63 / 2e-12
GATL7
galacturonosyltransferase-like 7 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10016301
149 / 8e-45
AT1G19300 506 / 0.0
PARVUS, GAOLAOZHUANGREN 1, GALACTURONOSYLTRANSFERASE-LIKE 1, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
Lus10016302
134 / 9e-42
AT3G50760 94 / 1e-24
galacturonosyltransferase-like 2 (.1)
Lus10018801
116 / 7e-32
AT1G19300 518 / 0.0
PARVUS, GAOLAOZHUANGREN 1, GALACTURONOSYLTRANSFERASE-LIKE 1, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
Lus10032728
115 / 9e-32
AT1G19300 515 / 0.0
PARVUS, GAOLAOZHUANGREN 1, GALACTURONOSYLTRANSFERASE-LIKE 1, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
Lus10010727
76 / 8e-17
AT1G70090 546 / 0.0
GALACTURONOSYLTRANSFERASE-LIKE 9, glucosyl transferase family 8 (.1.2)
Lus10041489
74 / 2e-16
AT1G13250 491 / 2e-175
galacturonosyltransferase-like 3 (.1)
Lus10029215
74 / 3e-16
AT1G70090 543 / 0.0
GALACTURONOSYLTRANSFERASE-LIKE 9, glucosyl transferase family 8 (.1.2)
Lus10000262
74 / 3e-16
AT3G62660 405 / 2e-141
galacturonosyltransferase-like 7 (.1)
Lus10010025
74 / 4e-16
AT3G62660 568 / 0.0
galacturonosyltransferase-like 7 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G132900
109 / 2e-29
AT1G19300 483 / 2e-172
PARVUS, GAOLAOZHUANGREN 1, GALACTURONOSYLTRANSFERASE-LIKE 1, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
Potri.014G040300
108 / 3e-29
AT1G19300 489 / 1e-174
PARVUS, GAOLAOZHUANGREN 1, GALACTURONOSYLTRANSFERASE-LIKE 1, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
Potri.007G031700
102 / 5e-27
AT3G50760 459 / 5e-163
galacturonosyltransferase-like 2 (.1)
Potri.005G128000
101 / 1e-26
AT3G50760 463 / 9e-165
galacturonosyltransferase-like 2 (.1)
Potri.010G129400
76 / 5e-17
AT1G13250 521 / 0.0
galacturonosyltransferase-like 3 (.1)
Potri.008G192600
76 / 7e-17
AT1G70090 556 / 0.0
GALACTURONOSYLTRANSFERASE-LIKE 9, glucosyl transferase family 8 (.1.2)
Potri.010G038300
74 / 2e-16
AT1G70090 580 / 0.0
GALACTURONOSYLTRANSFERASE-LIKE 9, glucosyl transferase family 8 (.1.2)
Potri.008G116900
72 / 2e-15
AT1G13250 509 / 0.0
galacturonosyltransferase-like 3 (.1)
Potri.008G018100
70 / 7e-15
AT3G06260 533 / 0.0
galactinol synthase 9, galacturonosyltransferase-like 4 (.1)
Potri.010G242300
67 / 9e-14
AT3G06260 524 / 0.0
galactinol synthase 9, galacturonosyltransferase-like 4 (.1)
PFAM info
Representative CDS sequence
>Lus10010790 pacid=23170758 polypeptide=Lus10010790 locus=Lus10010790.g ID=Lus10010790.BGIv1.0 annot-version=v1.0
ATGCCTGCATTTCTCTTTTCTTTCATTGTCATTCTCCTTCTTCTACCTTTGACTTCCAATGCATCCCTCACTTTCAAAGAAGCTCCCCAGTTTTACAATT
CCCCTGACTGCCCCTCCGCCGCTGACCATGACGCCATCTTCTGTTCAGATCGCGCAGTCCACGTGGCCATGACCTTAGACACCGCCTACCTCCGCGGCTC
CATGGCCGCTATCCTCTCAGTTCTCCAGCACTCCTCCTGCCCACAGAACATCGCTTTCCATTTCGTAGCATCAGAATCAGCCAACGCCTCCTCCGCCGTA
CTGCAATGCCACTTTCACTTTCTACTTCACTCCGACCTTCTGGTCCAATCCGTCTCTTTCGTTGACCTTCGCCGACAGGAAGGCCTGCTACTTCAATACC
GGGGTTATGGTGATTGA
AA sequence
>Lus10010790 pacid=23170758 polypeptide=Lus10010790 locus=Lus10010790.g ID=Lus10010790.BGIv1.0 annot-version=v1.0
MPAFLFSFIVILLLLPLTSNASLTFKEAPQFYNSPDCPSAADHDAIFCSDRAVHVAMTLDTAYLRGSMAAILSVLQHSSCPQNIAFHFVASESANASSAV
LQCHFHFLLHSDLLVQSVSFVDLRRQEGLLLQYRGYGD
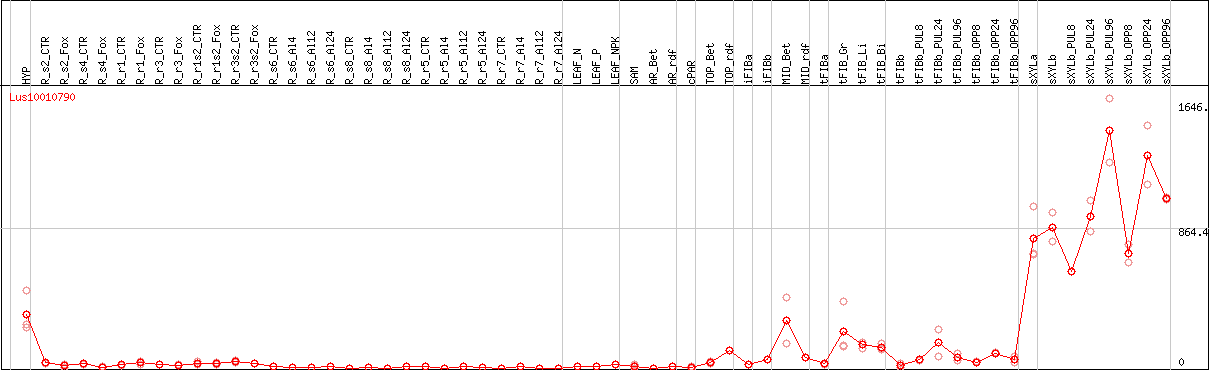
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010790 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.