External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G49710 101 / 5e-26
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G59200 100 / 8e-26
OTP80
ORGANELLE TRANSCRIPT PROCESSING 80, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G05750 97 / 1e-24
PDE247, CLB19
pigment defective 247, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G09410 97 / 2e-24
pentatricopeptide (PPR) repeat-containing protein (.1)
AT2G29760 97 / 2e-24
OTP81
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G06540 97 / 2e-24
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G39350 96 / 7e-24
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G25060 95 / 8e-24
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G37380 94 / 3e-23
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G13770 93 / 5e-23
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024376
238 / 5e-77
AT3G15130 370 / 1e-119
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10040680
109 / 9e-29
AT2G29760 926 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10018223
106 / 1e-27
AT2G29760 931 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10029344
100 / 2e-25
AT5G06540 803 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10031423
99 / 6e-25
AT5G66520 538 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10004253
98 / 1e-24
AT3G49170 832 / 0.0
embryo defective 2261, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10009913
97 / 2e-24
AT3G09040 1114 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10014594
97 / 2e-24
AT3G02330 901 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10019098
97 / 3e-24
AT3G47840 697 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G076100
172 / 7e-52
AT4G33170 383 / 1e-121
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.007G050200
106 / 1e-27
AT4G37380 817 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G006400
105 / 1e-27
AT1G56690 993 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.004G237000
105 / 2e-27
AT5G66520 544 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G071800
104 / 6e-27
AT1G08070 606 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.013G005400
102 / 2e-26
AT1G56690 1018 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.016G065500
101 / 4e-26
AT5G06540 828 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.004G111300
100 / 2e-25
AT3G02330 1077 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.005G012600
100 / 2e-25
AT2G29760 540 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G187800
100 / 2e-25
AT3G09040 1201 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10010853 pacid=23163414 polypeptide=Lus10010853 locus=Lus10010853.g ID=Lus10010853.BGIv1.0 annot-version=v1.0
ATGGTTGACACTTACGCAATTGTTCCTGAGACAGAGCATCCATCAAGTCTCATCGATATACTAGGACGCGCAGGGAGACTACACGACGCGGAAGAGTATG
TTGACAGATTCCGGTTCGGAAACGATCCTATTGTGTTGGGGAGTCTCCCTGGATGTGTTTCACATGGAGATTTCGCCATGGGAGAGCGTATAGGGAAGAG
GCTTCTGAAGCTTCAGCCGGAGACTACTTCACCTTATGTGTTGCTGTCGAATCTTTACGCTAGTGGTGAGATATGGAGAGGAGTTGCAGAGGCGAAGAGG
ATGCTTAAGGGTAGCGGGCTGAAGAAAGAGCCTGGGCATAGCCTGATAGAATTGGATGGAGCTTTTGAAAAGTTCAGCAAGTGA
AA sequence
>Lus10010853 pacid=23163414 polypeptide=Lus10010853 locus=Lus10010853.g ID=Lus10010853.BGIv1.0 annot-version=v1.0
MVDTYAIVPETEHPSSLIDILGRAGRLHDAEEYVDRFRFGNDPIVLGSLPGCVSHGDFAMGERIGKRLLKLQPETTSPYVLLSNLYASGEIWRGVAEAKR
MLKGSGLKKEPGHSLIELDGAFEKFSK
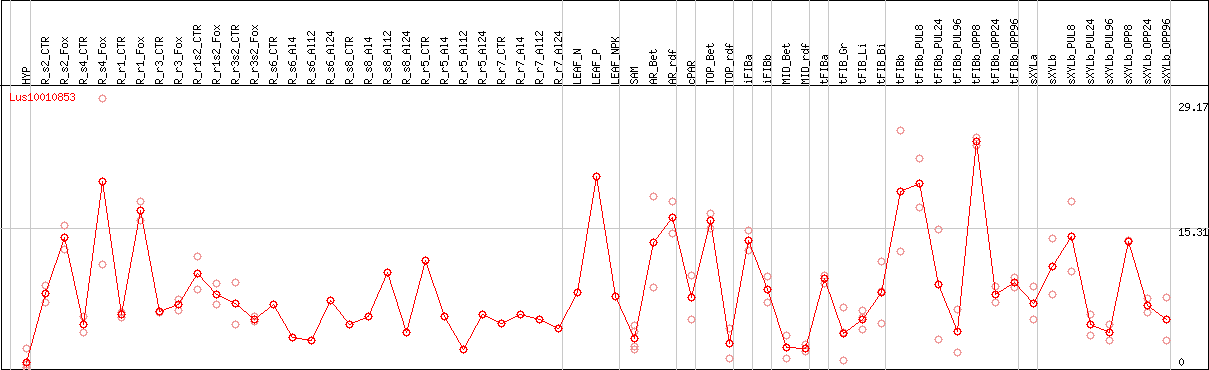
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10010853 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.