External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G42850 497 / 2e-173
CYP718
"cytochrome P450, family 718", cytochrome P450, family 718 (.1)
AT5G36110 230 / 1e-69
CYP716A1
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
AT5G45340 207 / 6e-61
CYP707A3
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
AT2G29090 207 / 8e-61
CYP707A2
"cytochrome P450, family 707, subfamily A, polypeptide 2", cytochrome P450, family 707, subfamily A, polypeptide 2 (.1.2)
AT4G19230 195 / 2e-56
CYP707A1
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
AT3G19270 171 / 2e-47
CYP707A4
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
AT3G50660 171 / 5e-47
PSC1, CYP90B1, CLM, SNP2, DWF4, SAV1
SUPPRESSOR OF NPH4 2, SHADE AVOIDANCE 1, PARTIALLY SUPPRESSING COI1 INSENSITIVITY TO JA 1, DWARF 4, CYTOCHROME P450 90B1, CLOMAZONE-RESISTANT, Cytochrome P450 superfamily protein (.1)
AT4G36380 164 / 1e-44
ROT3
ROTUNDIFOLIA 3, Cytochrome P450 superfamily protein (.1)
AT5G38970 163 / 1e-44
ATBR6OX, CYP85A1, BR6OX1
brassinosteroid-6-oxidase 1 (.1.2.3)
AT1G12740 156 / 5e-42
CYP87A2
"cytochrome P450, family 87, subfamily A, polypeptide 2", cytochrome P450, family 87, subfamily A, polypeptide 2 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10040055
238 / 2e-72
AT5G36110 318 / 7e-104
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
Lus10033308
192 / 4e-55
AT5G45340 732 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Lus10035685
191 / 9e-55
AT4G19230 714 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
Lus10034768
188 / 8e-54
AT5G45340 731 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Lus10019858
172 / 2e-47
AT3G19270 643 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Lus10024495
171 / 3e-47
AT1G12740 751 / 0.0
"cytochrome P450, family 87, subfamily A, polypeptide 2", cytochrome P450, family 87, subfamily A, polypeptide 2 (.1.2)
Lus10012675
169 / 9e-47
AT3G19270 586 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Lus10031391
159 / 9e-47
AT2G42850 141 / 8e-41
"cytochrome P450, family 718", cytochrome P450, family 718 (.1)
Lus10016515
164 / 9e-45
AT2G29090 554 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 2", cytochrome P450, family 707, subfamily A, polypeptide 2 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G060700
575 / 0
AT2G42850 582 / 0.0
"cytochrome P450, family 718", cytochrome P450, family 718 (.1)
Potri.011G001500
253 / 4e-78
AT5G36110 331 / 8e-109
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
Potri.012G115000
248 / 2e-76
AT5G36110 295 / 7e-95
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
Potri.004G017800
238 / 1e-72
AT5G36110 317 / 1e-103
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
Potri.011G001300
236 / 1e-71
AT5G36110 332 / 2e-109
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
Potri.019G057901
236 / 1e-71
AT5G36110 279 / 2e-88
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
Potri.006G085500
228 / 1e-68
AT5G36110 573 / 0.0
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
Potri.004G017633
227 / 1e-68
AT5G36110 301 / 3e-97
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
Potri.007G002400
227 / 3e-68
AT5G36110 585 / 0.0
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
Potri.001G003000
227 / 3e-68
AT5G36110 389 / 3e-131
"cytochrome P450, family 716, subfamily A, polypeptide 1", cytochrome P450, family 716, subfamily A, polypeptide 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00067
p450
Cytochrome P450
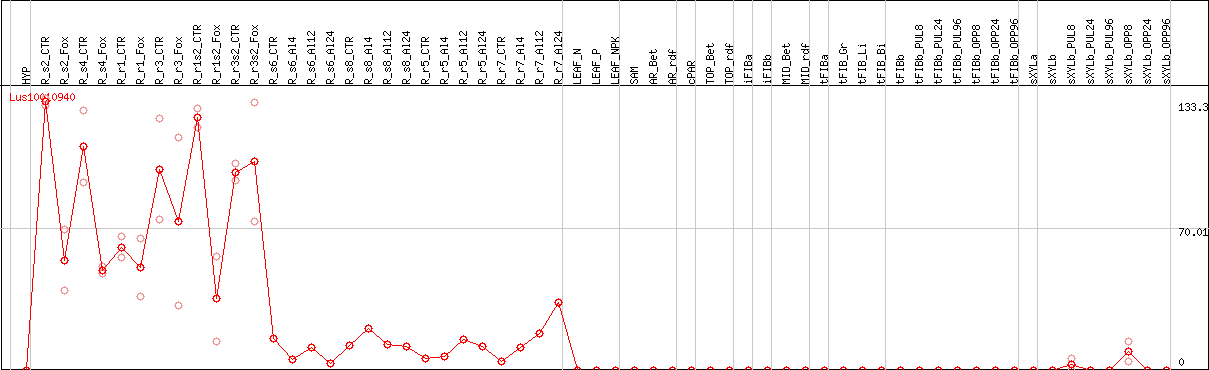
Representative CDS sequence
>Lus10010940 pacid=23172492 polypeptide=Lus10010940 locus=Lus10010940.g ID=Lus10010940.BGIv1.0 annot-version=v1.0
ATGTCCTATGAGATCATCATTGCCATTGTCACAACAACAACCTTAATCATACTTTCAACACGCTTGATAAGGATGATTGTCATCACCATATCCTCCAAAC
CAAATGGGAAGAAGAAAAACAAGAAGCTCAACCTCCCGCCGGGAGAAATGGGGCTCCCGTGGATCGGAGAGACGATGGACTTCTACAAATCCCAACGCAA
CAATCGCTTATTTGACCACTTCGTCCGCCCTCGAATGGCCAAGCACGGTAAGATCTTCAAGACAAGGCTAATGGGTTCCCCGACAATCGTGGTCAACGGT
GCGGAAGCCAACAAGTTCTTCCTCTCCAACGAGTTCAAGCTCGTGGTCAGCTCGTGGCCGACCTCGTCGGTCCAACTGATGGGGAGGAACTCCATCATGG
AGGCGAAGGGAGACAAGCACCGGACGGTCCGAGGTCTGATCGCCACGAGCGTGGGATACTCGTCGTTGGAGGTGCTGGTACCGAAGATGTGCAGCCTGGT
GCAAGCCAGCCTGGCCGAACACTGGGGTGTGATTGATCACATCCCAAAGGAGGTTAGTTTGTACTACTGGACGAAAAAGTTGACTTTTAAGATCGTGTTT
GAGTGCCTTTTGGGGATAAAGACGGAGTCAGGGACGTTGGAGATGTTTGAAAAAGTGCTAGAAGGGGTGTTCGCCCCTCCGGTAGGGTTTCCGGGGTCAA
AGTTTTCGCGGGCAAAGAAGGCCAGAGAGGAGATTGAGAGAATGTTGTTGGAGGTTGTGAGAGAAAAGAGGAAGGAAGTGGAAGAAAAAACCACCTCCGC
CCCCAGAGCGGACGGAGGAGGAAAAGAAGAGGAAGGACAGCTCATGTGGAGGCTCGTAAAGGCGATGATACAAGGAGAGATCACCGAGGAGGAAGTGGTT
GACAATGTGGTGTTGCTCGTGTTTGCCGCTCATGATACAACGTCGTTTGTTGTTGCCATGATGTTCAAGATGTTGGCTCAACACCCCGATTGCTACAATC
TCCTCTTGCAAGAGCATATCGAGATAGTGAAGAATAATGGAAACATTTTAAGTCTGGAAGAAGTAAAGAAGATGAAGTATACATGGGAAGTAGCACGTGA
GAGCATGAGGTTATTCCCTCCCATCTTTGGCTCCTTCAGGAAAGCTGTTCAAGACATTGAATTCGATGGCTTCACCATTCCCAAAGGATGGAAGGTGTTG
TGGACAGCGTACGGGACGCATTACAACGAGGAGTACTTCGAGGATCCGATGAGCTTCAAGCCGGTCCGGTTCCAACAACACAACAACATCCCACCCTACG
TCTACGTGCCGTTTGGTGGAGGTCCCAGGGTCTGCGCCGGTAGCCAGCTGGCCAAGCTCAGCATCCTAGTACTGGTTCACTACGTTGTAACTGGTTACGA
CTGGTCGCTTGTCCTACCAAACGAGGGAATCACAATGGATCCTCTTCCTTTTCCTTCTCATGGAATGCCCATCCGGATAACTCCCAAAGCTTGTAGCACA
AGTAACCGAGTTTAA
AA sequence
>Lus10010940 pacid=23172492 polypeptide=Lus10010940 locus=Lus10010940.g ID=Lus10010940.BGIv1.0 annot-version=v1.0
MSYEIIIAIVTTTTLIILSTRLIRMIVITISSKPNGKKKNKKLNLPPGEMGLPWIGETMDFYKSQRNNRLFDHFVRPRMAKHGKIFKTRLMGSPTIVVNG
AEANKFFLSNEFKLVVSSWPTSSVQLMGRNSIMEAKGDKHRTVRGLIATSVGYSSLEVLVPKMCSLVQASLAEHWGVIDHIPKEVSLYYWTKKLTFKIVF
ECLLGIKTESGTLEMFEKVLEGVFAPPVGFPGSKFSRAKKAREEIERMLLEVVREKRKEVEEKTTSAPRADGGGKEEEGQLMWRLVKAMIQGEITEEEVV
DNVVLLVFAAHDTTSFVVAMMFKMLAQHPDCYNLLLQEHIEIVKNNGNILSLEEVKKMKYTWEVARESMRLFPPIFGSFRKAVQDIEFDGFTIPKGWKVL
WTAYGTHYNEEYFEDPMSFKPVRFQQHNNIPPYVYVPFGGGPRVCAGSQLAKLSILVLVHYVVTGYDWSLVLPNEGITMDPLPFPSHGMPIRITPKACST
SNRV