External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G44800 197 / 1e-60
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G07480 194 / 3e-59
KUOX1
KAR-UP oxidoreductase 1 (.1)
AT3G60290 193 / 7e-59
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G36690 183 / 5e-55
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G10490 150 / 2e-42
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G24530 136 / 2e-37
DMR6
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G10500 134 / 2e-36
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G20400 102 / 7e-25
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G17010 100 / 4e-24
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G22880 100 / 1e-23
TT18, TDS4, ANS, LDOX
TANNIN DEFICIENT SEED 4, ANTHOCYANIDIN SYNTHASE, leucoanthocyanidin dioxygenase (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032502
214 / 8e-67
AT5G07480 396 / 3e-138
KAR-UP oxidoreductase 1 (.1)
Lus10023890
172 / 2e-50
AT2G36690 486 / 5e-173
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10014398
169 / 2e-49
AT2G36690 482 / 4e-171
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10032930
144 / 4e-40
AT5G24530 504 / 0.0
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10035782
140 / 1e-38
AT5G24530 258 / 4e-84
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10030185
139 / 3e-38
AT4G10490 442 / 3e-156
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10015573
129 / 8e-35
AT5G24530 510 / 0.0
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10024882
128 / 4e-34
AT5G24530 271 / 2e-89
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10042999
114 / 8e-31
AT5G07480 183 / 6e-58
KAR-UP oxidoreductase 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G080600
242 / 7e-78
AT5G07480 453 / 9e-161
KAR-UP oxidoreductase 1 (.1)
Potri.008G029700
175 / 7e-52
AT2G36690 478 / 1e-169
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.010G231500
165 / 7e-48
AT2G36690 474 / 3e-168
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451700
153 / 9e-44
AT4G10500 451 / 6e-160
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451600
148 / 8e-42
AT4G10500 406 / 5e-142
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451900
148 / 1e-41
AT4G10490 498 / 3e-178
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.014G106700
145 / 8e-41
AT4G10490 468 / 2e-166
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.017G048900
143 / 1e-39
AT2G36690 250 / 5e-80
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.011G150100
142 / 1e-39
AT4G10490 501 / 2e-179
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.017G048700
140 / 1e-38
AT2G36690 255 / 3e-82
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
Representative CDS sequence
>Lus10011002 pacid=23148697 polypeptide=Lus10011002 locus=Lus10011002.g ID=Lus10011002.BGIv1.0 annot-version=v1.0
ATGGAGGTTTCACCAGCTCCAACCATCGACTTCACCCATCTCCTCAGCTCAGCCAACCCCGAAACCAAATCAGAAGCCATTTCCCAAATCGCCGTCGCCT
GCAAATCCTTCGGCTTCTTCAATGTGATTAACCACGGAATCAGCGACGAAACGGTGCTCGGAGCGGTCGAATCCAACGCCAAGTTCTTCGATTCGCCATC
GGAGGAGAAAGAGGAGTTCCAATCCGGCGACGTCAGAAGGCCGGTGAGATTCGGTACGCTCGCTGCACAGAGCAAGCTAAGAAATGGTGATGGTGGGACT
TTTGCTAGGGATTTCCTCAAGCTCTATGGCTTCCCCTTGGAGGATTGGATCGGGTTCTGGCCAAAAAACCCGCCCGATTACAGGTTGAAAATGGGGAGAT
ACACTAAGGAAGCAAGAAACCTAAGCCTCTGTCTATTCGAATCGATACTCGAAAGCCTCGACATAGACCCAACCCCATTCATACAAAAGCTAACCTCTGA
AGGCATGCAAATGATGGCTGTAAACCATTACCATACTGGGTTGATTGGTTCCGGTTCGACCCAAACCGGCATTCCTCCCCATACAGACTTCTCCTTGATC
ACTGTTCTGCTCCAGACCAAACCAGGCCTCCATGTCTTCGACCCGACCGAAGGGAAATGGAAAGATGCCCCGGGATCAAGTTCGTCGATGCAGGTACTCG
TTGGGGATCATCTCGAGGTTCTGAGCAATGGAAGGTGCAAGGCAGCTGTGCATCGTGCAGCCTGCGGCACGAACTCCGAGGAGTTCACGAGTAGGATTTC
TGTGGCGAGCCTGCTTACCTTCGGGTTCGATGAAGTGGTTGAACCGGTTTTGGTGACGAAAAATCAAGAAGAAGTTGGGAAGAGTTTGTACAAGGGTAGT
AGCTTGAGGGAGTTTCTTGAGTACCTTTCATCTGGAGAAAAACTACCCTTCATTGACACTCTTAAGGTTTGTTGA
AA sequence
>Lus10011002 pacid=23148697 polypeptide=Lus10011002 locus=Lus10011002.g ID=Lus10011002.BGIv1.0 annot-version=v1.0
MEVSPAPTIDFTHLLSSANPETKSEAISQIAVACKSFGFFNVINHGISDETVLGAVESNAKFFDSPSEEKEEFQSGDVRRPVRFGTLAAQSKLRNGDGGT
FARDFLKLYGFPLEDWIGFWPKNPPDYRLKMGRYTKEARNLSLCLFESILESLDIDPTPFIQKLTSEGMQMMAVNHYHTGLIGSGSTQTGIPPHTDFSLI
TVLLQTKPGLHVFDPTEGKWKDAPGSSSSMQVLVGDHLEVLSNGRCKAAVHRAACGTNSEEFTSRISVASLLTFGFDEVVEPVLVTKNQEEVGKSLYKGS
SLREFLEYLSSGEKLPFIDTLKVC
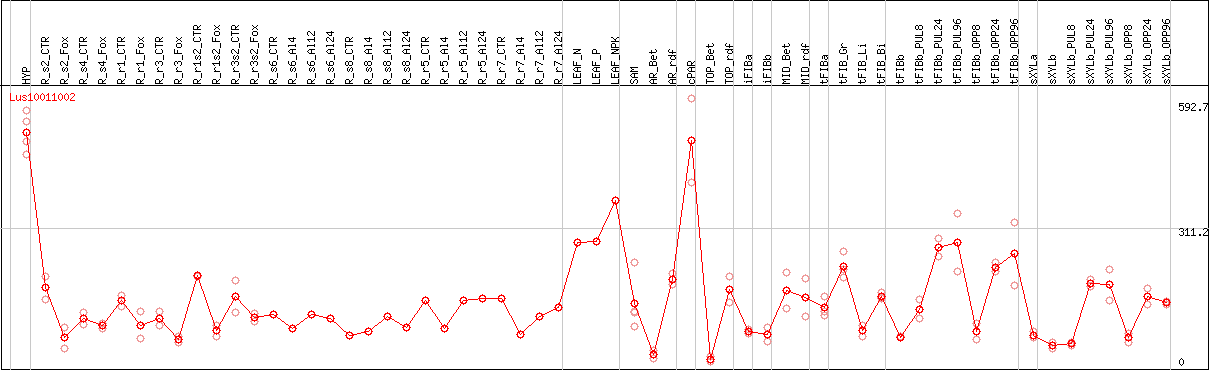
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011002 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.