Lus10011064 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10011064 pacid=23148623 polypeptide=Lus10011064 locus=Lus10011064.g ID=Lus10011064.BGIv1.0 annot-version=v1.0
ATGGAGAAGCTGGTCAAGAACCTAACGCAGCTCAAAGTTCTCCATCTAGATTGGGTTGATCTCTCCCAAACCATCAACAACAGGTCTTGGTCAACCTTAC
CTGATCTTCAGGAGCTCAGCCTGATCTATTGTAGAATCACCGGAGAGGCACTGCAGTATTCATCTCTTCTACACTTGAAATCCCTCACTCATTTGATACT
CTCATATAATTACCTTGCGTGGAATGTTTCTGGGTATTCCTTCATCGAGTTTCCTCATCTCAAACATCTAGAGCTTAGCAACTGTAGGTTGTATGGAAGA
TTTCTTAATAGCGTCTTCTTCATTCCCAATCTACAACATCTTGATGTTTCTTCCAATGATCAACTCTCTGGTACTCTGCCCCCCTTCCCTGCCGGCAACA
ATTCACTACAATATATTAAACTTTCGGAAACATTATTTTATGGTGAAGTACCTGATTCCATAAGCTATTTAGTGTCATTGCAGCAACTCTATCTCTCCAA
CTGCCACTTCCATGGAACATTCCCGAGTAGTATTTTCTTAAATCCAAATTTACAATCCATTAGTGTTTCCAACAACTCCCTTCTCACCGGTACTCTACCC
GAATTCCCTACTGCCGGTAGCCCTCTTCAATTCATCCAACTTTCAGATACAAAATTTCAGGGGAAACTGCCTGATTCTATCAACAATTTAGTTTCGTTGA
AGGAAGTGTATCTCTTAAACTGCATGTTTCATGGAGTTATCCCACCTTCGATTGACAGTCTAAGAGAACTACAACGCCTTCACCTTGCTGGAAACAACTT
CAATGGTCCACTCCCATTATTATCTTCATCTCGCAATATGTTGGTTCTTAACCTGGCTTTCAATAATTTCAGTGGAAACATTCCTCTATCTTACGGCAGC
GACCAGTTTCATGATCTTTGGAGGTTGTACTTGAACAACAACAACCTACAAGGACAAATTCCACCCTCTTTATTCAGTTTGCCATCCCTCCTCTCATTGG
AACTCGGCTGGAACCAACTAGAAGGGCAGCTGGAAGAAATTCACAGTTACTCATTTCAACTAAGGAAAATAGACTTGCGTCATAACAAGCTTCGAGGCCA
TCTTCCATTGTCATTCTTCAATATGACCGGCCTTAATTATCTATCGGTTGCTTTCAATGATTTCCAAGGAAGAATCAACCTTACGGAGACAGCCAACGGG
TTGCTCTTCTATTTAGACCTCTCACACAATCACTTCCAAGTATATCTTACTCCTAGACGGTCAGGTTTGCAGGTCCTTGGTTTGAGTTCATGCAATATAG
ACTCGTTGCCTGCAGATTTCTCAAACGCAAACCTTCAATATCTCGATCTTTCGCGAAATAGGATCAAGGGAAACATCTCTCCATCCATATGCAATATTCA
AGGTTTAGAAATCCTAAATCTCTCCTCTAATAACTTCAGTGGTCCAATTCCACCATGCTTGGCTTCACTTCCCTATCTCCGTGTGTTAAGCTTAAGAAAT
AACCAACTCAGTGGGATGATCCCTGAAGGTTTTACAGCAAACTGCACTCTGACTACTCTCGATGTGAATCAGAACCGCCTCCGAAACAAACTTCCTGGAT
CTCTAAGCCTGTGTAAAGGTCTGCAAGTTATAGATGTTGGGAAAAATAACTTGAAGGGGAAGTTCCCAGTTTGGTTGGAGAATTTGCCTTTGTTGCGAGT
TCTCATCTTGCACTCCAACCAATTCAGTGGTACTATTGGTTCCCGAAACAAGACTGGATTTCCTCTGCTCCAAATATTTGATGTCTCTTCTAACAATTTT
GTCGGTGAGATGCCTTCAGGGTGGTTCGTAAGTTGGAAAGAAATGATGAAGCATGCTACTCCAACGGGTTTCGATATCGAAGTTATCGGTTCCAGTTACT
CTTCAGGAACAGGTCTTGTCGCACCTGTCCTTGAACAAGAGTACCAAGATTCCATTACGGTCATTAATAAAGGAATTGTAACAGAGTACAGCAAGATCCT
CACCATCCTCATCTTAATTGATGTATCCGACAACGGATTCACTGGCGAAATTCCTGAAGCAATATCTCGTCTCGAATCGCTACATCTCCTCAACCTCTCC
AACAACCACTTTACAGGTCACATCCCAGGATCATTCGGACAACTAAGACAGCTAGAGTCACTAGACCTTTCGTCAAACAAGATTTCTGGCAACATCCCAA
CACAACTTACAGAGCTCACATTTATGTCAAAACTGAACCTGTCAAAGAACTCACTCACGGGTGAGATACCTCGAGGACGGCAATTCGCAACTTTCGAAGT
AGAATCTTATGAAGGCAATCCAGGATTGTGCGGCCTTCCATTGCCTAAACCATGCCACAATGTGAAGTCCCCGCCTATAAGTAGCAAACCAGATGATGCA
AAAGCCAAAGGAGGGAATAGTGTGATTGACTGGGAGTACTATCGAGTTGGGATCGCGTGTGGAGGCGGGACAGGACTCATTGCTGGGTTCGTAGCAGCCT
ATTTTGTGTTTGGAAATTTCTTCATCAAGACACGAAGAGTTCGTCGTAATGCTCAAAGTAAGAAGATTGCAGACGCTAAGCTTGCGAAAGATTTCCAAGC
AGTTCTGAAAGAGTTTCAGAAGGCTCAACGTCTCTCAGCTGAGAGAGAAACCGCTTATTCTCCATTCGCTGCCAAGGCAAACCCTCCATCCAGACAAGAA
GTCGTGCTGTTGGACAATGAAATTTCTTTCAACGAGGCAATCATCGAGGAAAGAGAGCAAGGGATACACGAAATTTCGCAGCAAATCGGTGAGGTGAACG
AGATTTTTAAGGATCTTGCAGTCCTTGTCCACGAACAAGGAACCATGATCGACGATATCGGTTCGCACATCGATAATGCACATGCTGCAACTGCACAAGG
GAAACAACATCTCGCCAAAGCCGCGAAAACCCAAAAATCGAATTCATCTCTGACCTGCTTACTCATGGTCATCTTCGCAATTGTGCTCCTCATCGTCATT
GTAGTGCTTGCAGCCTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10011064 pacid=23148623 polypeptide=Lus10011064 locus=Lus10011064.g ID=Lus10011064.BGIv1.0 annot-version=v1.0
MEKLVKNLTQLKVLHLDWVDLSQTINNRSWSTLPDLQELSLIYCRITGEALQYSSLLHLKSLTHLILSYNYLAWNVSGYSFIEFPHLKHLELSNCRLYGR
FLNSVFFIPNLQHLDVSSNDQLSGTLPPFPAGNNSLQYIKLSETLFYGEVPDSISYLVSLQQLYLSNCHFHGTFPSSIFLNPNLQSISVSNNSLLTGTLP
EFPTAGSPLQFIQLSDTKFQGKLPDSINNLVSLKEVYLLNCMFHGVIPPSIDSLRELQRLHLAGNNFNGPLPLLSSSRNMLVLNLAFNNFSGNIPLSYGS
DQFHDLWRLYLNNNNLQGQIPPSLFSLPSLLSLELGWNQLEGQLEEIHSYSFQLRKIDLRHNKLRGHLPLSFFNMTGLNYLSVAFNDFQGRINLTETANG
LLFYLDLSHNHFQVYLTPRRSGLQVLGLSSCNIDSLPADFSNANLQYLDLSRNRIKGNISPSICNIQGLEILNLSSNNFSGPIPPCLASLPYLRVLSLRN
NQLSGMIPEGFTANCTLTTLDVNQNRLRNKLPGSLSLCKGLQVIDVGKNNLKGKFPVWLENLPLLRVLILHSNQFSGTIGSRNKTGFPLLQIFDVSSNNF
VGEMPSGWFVSWKEMMKHATPTGFDIEVIGSSYSSGTGLVAPVLEQEYQDSITVINKGIVTEYSKILTILILIDVSDNGFTGEIPEAISRLESLHLLNLS
NNHFTGHIPGSFGQLRQLESLDLSSNKISGNIPTQLTELTFMSKLNLSKNSLTGEIPRGRQFATFEVESYEGNPGLCGLPLPKPCHNVKSPPISSKPDDA
KAKGGNSVIDWEYYRVGIACGGGTGLIAGFVAAYFVFGNFFIKTRRVRRNAQSKKIADAKLAKDFQAVLKEFQKAQRLSAERETAYSPFAAKANPPSRQE
VVLLDNEISFNEAIIEEREQGIHEISQQIGEVNEIFKDLAVLVHEQGTMIDDIGSHIDNAHAATAQGKQHLAKAAKTQKSNSSLTCLLMVIFAIVLLIVI
VVLAA
|
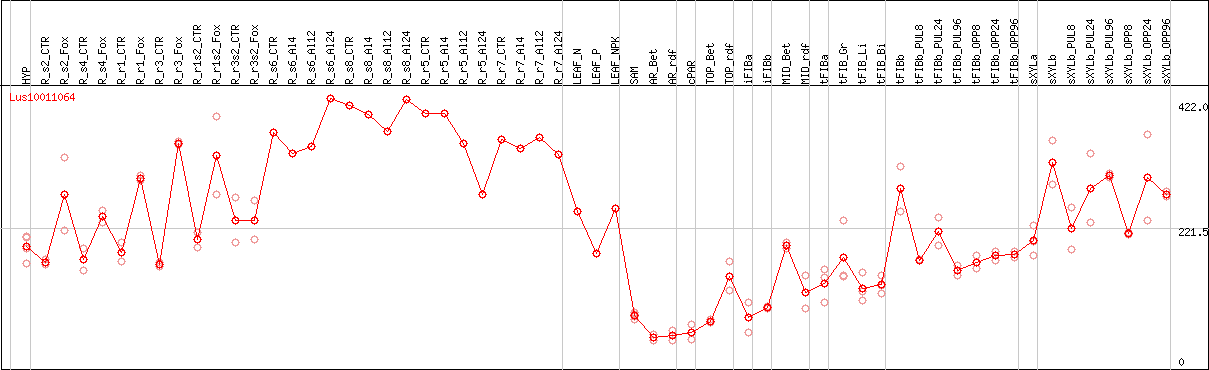
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011064 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
