External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G33910 228 / 4e-74
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G23096 164 / 4e-49
P4H13
prolyl 4-hydroxylase 13, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT1G20270 105 / 2e-26
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G66060 101 / 4e-25
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
AT2G17720 100 / 1e-24
P4H5
prolyl 4-hydroxylase 5, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G35810 94 / 3e-22
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G28490 93 / 5e-22
Oxoglutarate/iron-dependent oxygenase (.1)
AT3G28480 88 / 7e-20
Oxoglutarate/iron-dependent oxygenase (.1.2)
AT2G43080 85 / 6e-19
AT-P4H-1
P4H isoform 1 (.1)
AT3G06300 84 / 9e-19
P4H2, AT-P4H-2
prolyl 4-hydroxylase 2, P4H isoform 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028404
110 / 3e-28
AT5G66060 486 / 8e-176
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10041857
108 / 1e-27
AT5G66060 488 / 2e-176
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10012014
93 / 1e-21
AT5G18900 448 / 2e-160
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10016271
93 / 1e-21
AT5G18900 444 / 2e-158
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10032183
90 / 1e-20
AT3G28480 434 / 3e-154
Oxoglutarate/iron-dependent oxygenase (.1.2)
Lus10014502
89 / 2e-20
AT3G28480 398 / 4e-140
Oxoglutarate/iron-dependent oxygenase (.1.2)
Lus10031048
87 / 4e-20
AT2G43080 406 / 7e-145
P4H isoform 1 (.1)
Lus10005620
89 / 5e-20
AT1G20270 465 / 7e-167
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10017249
88 / 9e-20
AT1G20270 455 / 3e-162
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G052600
245 / 1e-80
AT4G33910 353 / 4e-123
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G296800
234 / 2e-76
AT4G33910 441 / 6e-158
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.009G091000
218 / 4e-70
AT4G33910 431 / 9e-154
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.005G245300
104 / 4e-26
AT1G20270 483 / 3e-174
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.005G108000
97 / 1e-23
AT5G66060 405 / 1e-143
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.007G060800
95 / 2e-22
AT5G66060 349 / 6e-121
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Potri.008G197700
92 / 1e-21
AT5G18900 448 / 3e-160
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.002G232100
91 / 3e-21
AT2G43080 456 / 4e-164
P4H isoform 1 (.1)
Potri.017G075300
90 / 8e-21
AT3G28480 345 / 5e-120
Oxoglutarate/iron-dependent oxygenase (.1.2)
Potri.017G075100
89 / 3e-20
AT3G28480 436 / 3e-155
Oxoglutarate/iron-dependent oxygenase (.1.2)
PFAM info
Representative CDS sequence
>Lus10011085 pacid=23165720 polypeptide=Lus10011085 locus=Lus10011085.g ID=Lus10011085.BGIv1.0 annot-version=v1.0
ATGAAACCCGTCAAACCTTCCAGGGCCCACTTCTTCCTCAAGCTTAGAGGGAAGTTAGGGTTACCTGCCATCTTCATTTGCTTCTTGTTCTTCTTCCTCG
CCGGCCTTTTTGCCTCCAATCTTATCTCTCAGGAGGATCCTGGTCTGACGTCCAACTCCCGTAAGCTGCAGTCTTCGTTGCTGGAGACGAAGAAATTCGA
GTACAATTTACTCCCTCCGGGACAGTCTGGCGATGATTCCATCACGCTTATCCCTTTCCAGGTCCTGAGCTGGAAACCTCGTGCTTTGTACTTTCCCAAC
TTTGCCTCTGCAGAGCAATGTGAAAGTATAATTAGTATGGCAAAGCCACATCTAAAGCCGTCAACTTTGGCTTTACGCAAGGGAGAAACAGAAGAGAGTA
CGGAGGGAGTAAGAACAAGCTCTGGTGTGTTCCTCAGTTCTTCTGAAGATGACACAGGCGTCTTGGATGTGATCGAAGAAAAAATTGCTAGAGCGAGTAT
GATTCCAAGGGCCCATGGAGAGGCATTCAATATTCTTCGTTATGAGGTTGGCCAAAAGTATGATTCTCACTATGATGCGTTCAATCCTTCAGAATATGGT
CCACAAAAGAGTCAACGGATAACTTATATTCCATACATGAATTTGCTGGTCCACAACAACATGAATACCTGTAAAATCAGCCTTATGGAAGGCGCGATTA
GGCGTCCGCCTAGACGAGAAAAACCCCAAGGCACCGGCAAAACGCCTCGCCTCGGGCAAGGCGATGAAATAAGGATGTCGACCGAAGAATCACTAAGGAA
GAGCGAAATCGATGAGGGATCGATTTCCTGCGACGCCGAGGCGGCGAAATCTGCTATTAGGCGTTAA
AA sequence
>Lus10011085 pacid=23165720 polypeptide=Lus10011085 locus=Lus10011085.g ID=Lus10011085.BGIv1.0 annot-version=v1.0
MKPVKPSRAHFFLKLRGKLGLPAIFICFLFFFLAGLFASNLISQEDPGLTSNSRKLQSSLLETKKFEYNLLPPGQSGDDSITLIPFQVLSWKPRALYFPN
FASAEQCESIISMAKPHLKPSTLALRKGETEESTEGVRTSSGVFLSSSEDDTGVLDVIEEKIARASMIPRAHGEAFNILRYEVGQKYDSHYDAFNPSEYG
PQKSQRITYIPYMNLLVHNNMNTCKISLMEGAIRRPPRREKPQGTGKTPRLGQGDEIRMSTEESLRKSEIDEGSISCDAEAAKSAIRR
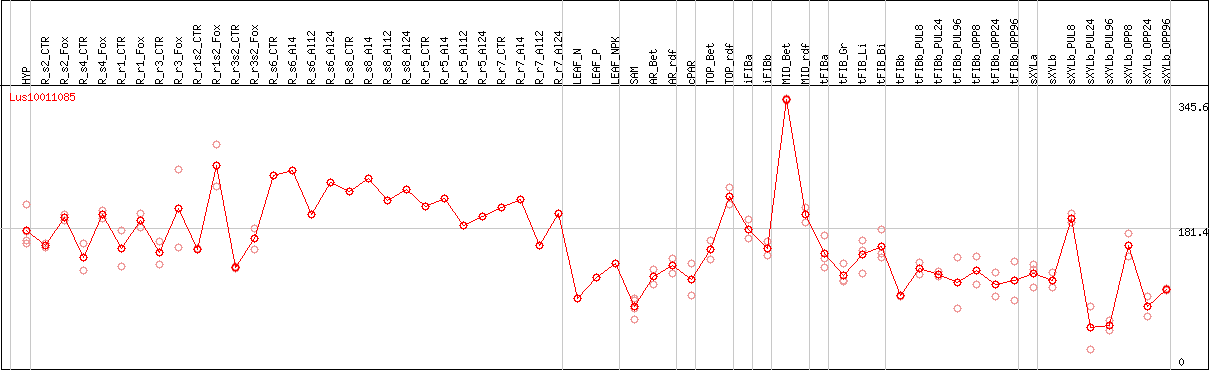
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10011085 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.