Lus10011182 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10011182 pacid=23147162 polypeptide=Lus10011182 locus=Lus10011182.g ID=Lus10011182.BGIv1.0 annot-version=v1.0
ATGATGATGGACGAGGATCAGGTGTACGATACGGAGATGATGGAGGATGTGGAAGGAGATGAGGAAATCACTCAGGAGGACGCGTGGGCGGTAATCTCAG
CTTACTTCGAAGAGAAAGGTCTTGTGCGACAGCAGCTTGATTCGTTCGACGAGTTCATCCAGAACACAATGCAGGAGATTGTGGACGAGTCGGCTGACAT
CGAGATTCGGCCGGAGAGCCAGCATAACCCCGGGCATCAGTCCGATTCCCTTGAGACCATTTACAAAATCAGCTTCGGTCAAATATATTTGAGTAAGCCT
ATGATGACAGAGTCTGATGGAGAGACAGCAACACTGTTCCCTAAAGCCGCCAGGTTGAGGAACTTGACATATTCAGCTCCATTGTATGTGGACGTCTCAA
AAAGGGTGATAAAGAAGGGCCATGATGGTGAAGAAGTAACTGAATCTCAAGATTTCACCAAAGTTTTTATTGGCAAGGTGCCAATCATGCTCCGGTCAAG
TTATTGCACTTTGTACCAGAATTCCGAGACAGATTTGACAGAGCTTGGAGAGTGTCCCTTTGATCAAGGTGGGTATTTCATCATCAATGGGAGTGAAAAG
GTGCTCATTGCACAGGAGAAGATGAGCACAAATCATGTCTATGTTTTCAAGAAACGACAGCCGAACAAGTATTCCCATGTGGCTGAAGTGCGATCCATGG
CAGAGTCTCAGAACAGGCCTCCAAGTACCATGTTCGTTCGGATGCTTTCTCGTGCTGGTGCTAAAGGGGGCTCAACAGGACAATACATTAAGGCTACTCT
ACCATACATCAGGACTGAAATTCCTATCATTATTGTGTTTCGAGCTCTTGGGTTTGTTGCTGACAAAGATATATTAGAGCACATTTGTTATGATTTCGCT
GATACTCAAATGATGGAACTGTTACGTCCCTCGTTGGAAGAGGCATTCGTGATTCAAAATCAACAGGTTGCTCTCGACTATATCGGGAAACGTGGGGCAA
CTGTTGGTGTTACGAGGGAAAAAAGGATCAAGTATGCTAGAGATATTCTGCAAAAGGAAATGCTCCCTCATGTTGGCGTTGGAGAATTTTGCGAGACCAA
AAAAGCTTACTATTTTGGGTATATCATTCACCGGCTTCTACTTTGTGCACTTGGACGAAGGCCCGAGGATGATAGGGATCATTATGGAAACAAGAGGCTC
GATCTTGCTGGTCCTTTGCTCGGTGGCCTTTTCAGAATGTTATTCAGAAAGTTGACTAGGGATGTCAGGTCATATGTGCAGAAGTGTGTTGATAATGGTA
AAGATGTTAATCTGCAATTTGCTATCAAAGCAAAAACCATAACCAGTGGTCTTAAATACTCCCTTGCGACTGGAAATTGGGGGCAAGCAAATGCGGCTGG
CACTAGAGCTGGAGTGTCCCAAGTGTTAAATAGGTTGACCTTTGCCTCTACCCTATCACATCTCCGGCGGCTGAACTCCCCAATTGGGCGTGAAGGAAAA
TTGGCAAAACCAAGGCAGCTTCATAATTCCCAGTGGGGAATGATGTGTCCAGCCGAGACACCTGAAGGACAGGCCTGCGGACTAGTGAAGAATCTTGCTT
TGATGGTATACATTACTGTCGGATCGGCAGCCTACCCCATATTGGAGTTTTTAGAAGAGTGGGGGACTGAAAACTTCGAGGAAATTTCTCCCGCAGTAAT
ACCTCAAGCCACCAAGATTTTCGTTAATGGTTGCTGGGTTGGTATCCATCGAGACCCTGACATGTTGGTAAAGACATTGCGCCGGCTGCGGCGGAGGGTT
GATGTTAACACTGAAGTCGGAGTTGTGAGAGACATTCATTTAAAGGAGCTGCGTATATACACAGATTATGGCCGTTGCAGTCGTCCTTTATTTATTGTTG
AGAGCCAAAGACTTCTTATAAGAAAGAAGGACATCACCGCATTGCAGCAGAGGGAATCAACTGAAGAAGGAGGCTGGCACGATCTGGTGGCCAAAGGGTT
TATCGAGTATATCGATACCGAGGAAGAGGAGACAACTATGATATCAATGACCATACATGATCTTGTACAAGCCAGAACTTACCCGGAGGAGGCTTATTCG
GGCACTTACACTCACTGTGAGATTCATCCCTCGTTGATTCTTGGTGTTTGTGCCTCGATCATTCCGTTTCCTGATCATAACCAGTCTCCACGTAACACTT
ACCAATCTGCTATGGGGAAGCAAGCCATGGGAATTTACGTCACTAACTATCAATTCCGTATGGACACATTGGCGTATGTTCTGTACTACCCGCAAAAACC
ACTCGTTACGACAAGAGCTATGGAGCATCTCCACTTCCGGCAGCTTCCAGCTGGCATCAATGCCATTGTGGCCATCTCTTGCTATTCTGGTTACAATCAA
GAAGATTCTGTCATCATGAATCAATCCTCGATTGATCGTGGATTCTTCAGATCGCTTTTCTTCCGATCTTACAGGGACGAGGAGAAAAAGATGGGCACGC
TCGTTAAAGAAGATTTCGGCCGTCCAGACAGAGCTAACACCATGGGAATGCGACATGGCTCGTATGAGAAATTGGACGATGATGGTCTTGCTCCGCCTGG
TACGAGATGTGCTGGTGAGGATGTTATCATCGGAAAGACCACCCCAATCTCGCAGGAAGACGCCCAGGGTCAAAATGCACGATACACGAGGCGTGACCAC
AGTATAAGCTTACGTCACAGTGAAACCGGGATGGTTGATCAAGTTTTGCTGACGACGAATGCTGATGGGTTGAGGTTTGTCAAAGTGAGGGTAAGATCCG
TTCGGATTCCTCAGATCGGAGACAAGTTCAGTAGTAGACACGGTCAGAAGGGAACCGTTGGTATGACTTATACTCAAGAGGACATGCCTTGGACTATTGA
AGGCGTAACGCCTGATATCATCGTGAATCCGCATGCTATTCCCTCTCGTATGACGATCGGTCAGCTCATTGAGTGTATCATGGGGAAAGTTGCTGCTCAC
ATGGGCAAGGAAGGCGACGCTACTCCGTTCACAGATGTTACTGTGGACAATATAAGCAAAGCCTTGCACAAGTGCGGGTACCAAATGCGTGGTTTCGAGA
CCATGTACAACGGCCATACAGGCCGACGCCTCACAGCCATGATCTTCCTCGGCCCTACTTACTACCAGAGGTTGAAGCATATGGTCGACGACAAGATCCA
CTCTCGAGGAAGGGGACCTGTTCAGATCCTCACAAGGCAGCCGGCTGAAGGACGGTCCCGAGACGGCGGTCTCCGTTTCGGCGAGATGGAGCGTGACTGC
ATGATTGCCCACGGCGCCGCCAACTTCCTCAAGGAGAGGCTGTTCGACCAGAGCGACGCCTACAGAGTTCACGTCTGCGAGCCTTGCGGGCTGATCGCCA
TTGCGAATCTGAAGAAGAGTTCGTTCGAGTGCCGGGGCTGCAAGAACAAGACCGAGATTGTCCAGGTTCGTAATCGTATAAATAACCTTTCTCGGTTAGA
TATCTCGACGATTTTTGCCACTAATCCATTCGTCTCGCAGGTTCAAATACCTTACGCGTGCAAGCTCCTCTTCCAAGAGCTTATGTCCATGGCCATAGCT
CCGAGAATGCTCACGAAGGATATACTTGTAAATCAGAAGTTGAAAGGGCAAAAGAGGAAATGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10011182 pacid=23147162 polypeptide=Lus10011182 locus=Lus10011182.g ID=Lus10011182.BGIv1.0 annot-version=v1.0
MMMDEDQVYDTEMMEDVEGDEEITQEDAWAVISAYFEEKGLVRQQLDSFDEFIQNTMQEIVDESADIEIRPESQHNPGHQSDSLETIYKISFGQIYLSKP
MMTESDGETATLFPKAARLRNLTYSAPLYVDVSKRVIKKGHDGEEVTESQDFTKVFIGKVPIMLRSSYCTLYQNSETDLTELGECPFDQGGYFIINGSEK
VLIAQEKMSTNHVYVFKKRQPNKYSHVAEVRSMAESQNRPPSTMFVRMLSRAGAKGGSTGQYIKATLPYIRTEIPIIIVFRALGFVADKDILEHICYDFA
DTQMMELLRPSLEEAFVIQNQQVALDYIGKRGATVGVTREKRIKYARDILQKEMLPHVGVGEFCETKKAYYFGYIIHRLLLCALGRRPEDDRDHYGNKRL
DLAGPLLGGLFRMLFRKLTRDVRSYVQKCVDNGKDVNLQFAIKAKTITSGLKYSLATGNWGQANAAGTRAGVSQVLNRLTFASTLSHLRRLNSPIGREGK
LAKPRQLHNSQWGMMCPAETPEGQACGLVKNLALMVYITVGSAAYPILEFLEEWGTENFEEISPAVIPQATKIFVNGCWVGIHRDPDMLVKTLRRLRRRV
DVNTEVGVVRDIHLKELRIYTDYGRCSRPLFIVESQRLLIRKKDITALQQRESTEEGGWHDLVAKGFIEYIDTEEEETTMISMTIHDLVQARTYPEEAYS
GTYTHCEIHPSLILGVCASIIPFPDHNQSPRNTYQSAMGKQAMGIYVTNYQFRMDTLAYVLYYPQKPLVTTRAMEHLHFRQLPAGINAIVAISCYSGYNQ
EDSVIMNQSSIDRGFFRSLFFRSYRDEEKKMGTLVKEDFGRPDRANTMGMRHGSYEKLDDDGLAPPGTRCAGEDVIIGKTTPISQEDAQGQNARYTRRDH
SISLRHSETGMVDQVLLTTNADGLRFVKVRVRSVRIPQIGDKFSSRHGQKGTVGMTYTQEDMPWTIEGVTPDIIVNPHAIPSRMTIGQLIECIMGKVAAH
MGKEGDATPFTDVTVDNISKALHKCGYQMRGFETMYNGHTGRRLTAMIFLGPTYYQRLKHMVDDKIHSRGRGPVQILTRQPAEGRSRDGGLRFGEMERDC
MIAHGAANFLKERLFDQSDAYRVHVCEPCGLIAIANLKKSSFECRGCKNKTEIVQVRNRINNLSRLDISTIFATNPFVSQVQIPYACKLLFQELMSMAIA
PRMLTKDILVNQKLKGQKRK
|
DESeq2's median of ratios [FLAX]
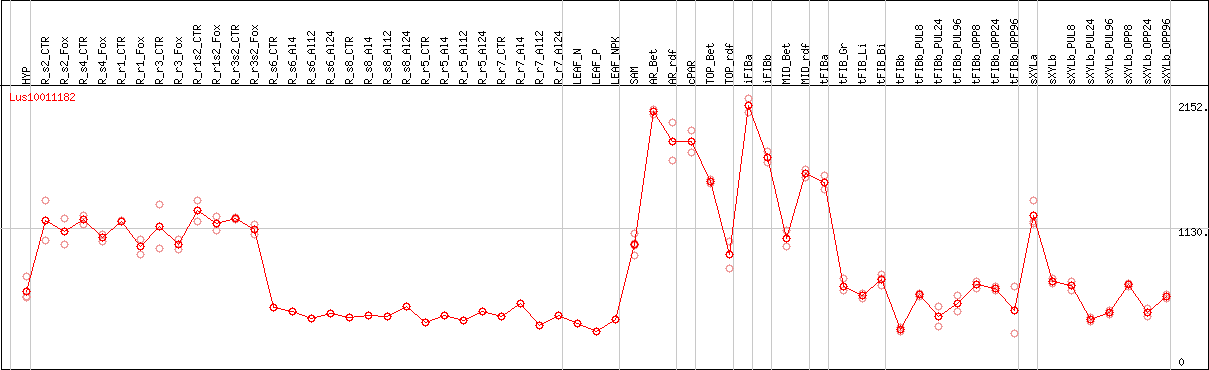
Coexpressed genes
Lus10011182 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
